Abstract
We developed a rapid insulated isothermal PCR (iiPCR) assay for on-site detection of Mannheimia haemolytica using a primer and probe set targeting the superoxide dismutase (sodA) gene. Our iiPCR assay detected M. haemolytica clinical isolates successfully and produced negative results on other bovine or ovine respiratory pathogens, including Histophilus somni, Bibersteinia trehalosi, Trueperella pyogenes, Streptococcus suis, and Mycoplasma spp., indicating that the PCR reactions were specific. Additionally, our iiPCR assay detected as few as 21 copies of genomic DNA and 17.2 cfu/mL of bacterial culture, which was 10 and 100 times more sensitive than conventional PCR, respectively. Our iiPCR assay can be performed on a portable device in a total of 58 min and may be a useful tool for the detection of M. haemolytica in bovine and ovine respiratory disease in the field.
A low number of Mannheimia haemolytica bacteria are frequently isolated from the upper respiratory tract of healthy ruminants. 3 A compromised primary respiratory defense resulting from stress or coinfection with respiratory viral pathogens or Mycoplasma spp. can lead this bacterium to invade the lungs and eventually cause severe pneumonia.11,15 M. haemolytica is believed to be one of the most important pathogens associated with bovine respiratory disease. 3 M. haemolytica can also cause pneumonia in sheep and goats.4,12,14
Molecular assays can provide more rapid, sensitive, and specific detection of pathogens than conventional bacterial culture. To facilitate rapid detection of M. haemolytica, traditional PCR and real-time PCR (rtPCR) assays have been developed and widely utilized in clinical diagnostic and epidemiologic investigations.1,9,13,16 However, post-PCR electrophoresis imaging analysis, or utilizing a costly real-time thermocycler within the laboratory, is usually necessary to obtain the final results. These requirements limit the application of conventional PCR and rtPCR for the on-site detection of pathogens in the field. Insulated isothermal PCR (iiPCR) is a newly developed technology that applies thermal convection to drive PCR cycles in a specially designed portable device and capillary tubes. The Pockit nucleic acid analyzer (GeneReach) is a commercial device designed for the performance of iiPCR assays in the field. The fluorescence signal generated through probe hydrolysis during PCR cycles is detected by the device, converted to S:N ratios (signalafter:signalbefore), and ultimately interpreted as positive or negative results.
The iiPCR assay has been used successfully to detect a variety of veterinary pathogens, including influenza A(H7N9) virus, 6 classical swine fever virus (Pestivirus C), 8 foot-and-mouth disease virus, 2 duck hepatitis A virus (Avihepatovirus A), 10 canine distemper virus (Canine morbillivirus), 18 Salmonella spp., 17 and Mycoplasma synoviae. 7 To develop an iiPCR assay for the detection of M. haemolytica, we initially selected 7 reference genes as targets, including 1) translation initiation factor 2 (infB), 2) β-subunit of RNA polymerase (rpoB), 3) β-subunit of the ATP synthase (atpD), 4) elongation factor Tu (tuf), 5) the gyrase subunit A (gyrA), 6) 16S ribosomal RNA subunit (16S rrnA), and 7) superoxide dismutase (sodA). Next, Beacon Designer v.7.0 software (Premier Biosoft) was utilized for primer and probe design. The resulting sequences with high scores were further applied to in silico assessment using BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to identify the unique sequences of the primers and probe to M. haemolytica. The resulting sequences that were 100% identical to the superoxide dismutase gene sodA of M. haemolytica (GenBank AY702512.1) were employed for iiPCR assay development. The sequences of primers and probe used to target sodA were forward primer sodAF (5′-TTAGTATTAGAAGAGGGTAAATTAG-3′), reverse primer sodAR (5′-AAATCGGATAGCCTGAAA-3′), and probe sodAP (5′-AACCGCTAACCAGGACAACC-3′), with FAM reporter dye and TAMRA quencher affixed on the 5′- and 3′-ends, respectively.
All bacterial isolates used in our study were first identified by species-specific PCR assay and further confirmed by 16S rRNA sequencing (Table 1). The bacterial strains belonging to Pasteurellaceae were also verified by sequencing the coding sequence of sodA. Bacterial strains were grown in trypticase soy broth; strains of Mycoplasma were propagated in Hayflick broth containing 20% horse serum. We also collected 67 clinical samples by swabbing the bronchi of sheep with pneumonic lesions at slaughter examination. Genomic DNA from bacterial cultures or bronchial swabs was extracted (TIANamp genomic DNA kit; Tiangen Biotech), following the manufacturer’s instructions. The DNA extracted from the bacterial cultures and bronchial swabs was eluted in 50 µL and 30 µL of deionized water, respectively.
Results of the insulated isothermal PCR (iiPCR) assay for the bacterial strains tested.
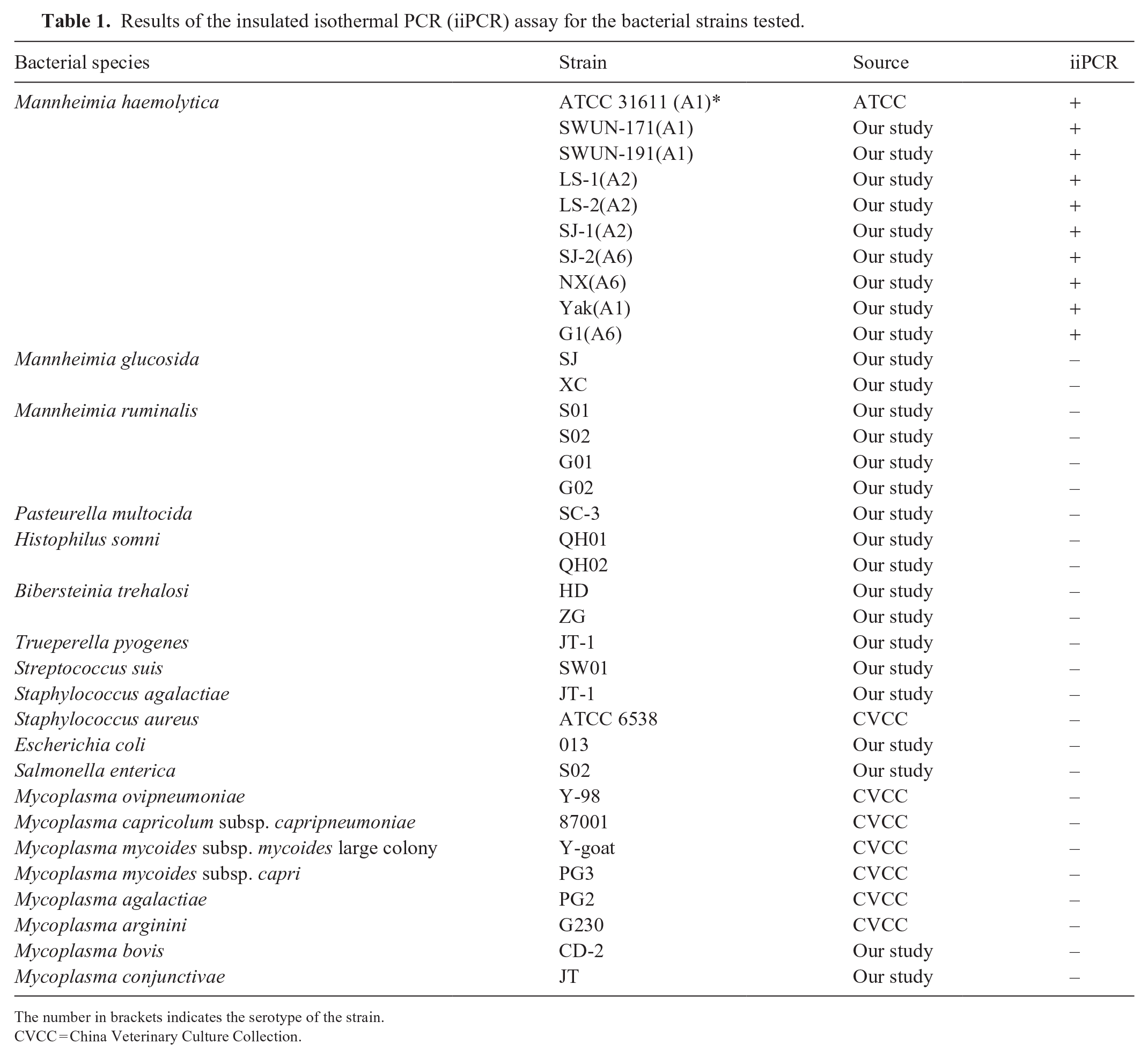
The number in brackets indicates the serotype of the strain.
CVCC = China Veterinary Culture Collection.
All iiPCR assays were performed in a portable Pockit nucleic acid analyzer with manufactured R-tubes compatible with the analyzer. The highest A520:B520 value provided by the Pockit device was also used to optimize the final concentrations of primers and probe. The PCR reaction mixture contained 25 µL of 2 × Premix Ex Taq master mix (TaKaRa Bio), 1.5 µL of 10 µM forward and reverse primers, 0.5 µL of 10 µM probe, 2 µL of DNA extract, and 19.5 µL of nuclease-free water in a total volume of 50 µL. The reaction was completed in 58 min, and the final result was shown on the screen of the device with a symbol of “+”, “−”, or “?”, representing a positive, negative, or undetermined result, respectively.
Our iiPCR assay detected all M. haemolytica clinical isolates used in our study, but not Mycoplasmas spp., other Mannheimia spp., and other common respiratory pathogens in sheep and cattle, including Histophilus somni, Bibersteinia trehalosi, Trueperella pyogenes, and Streptococcus suis (Fig. 1).
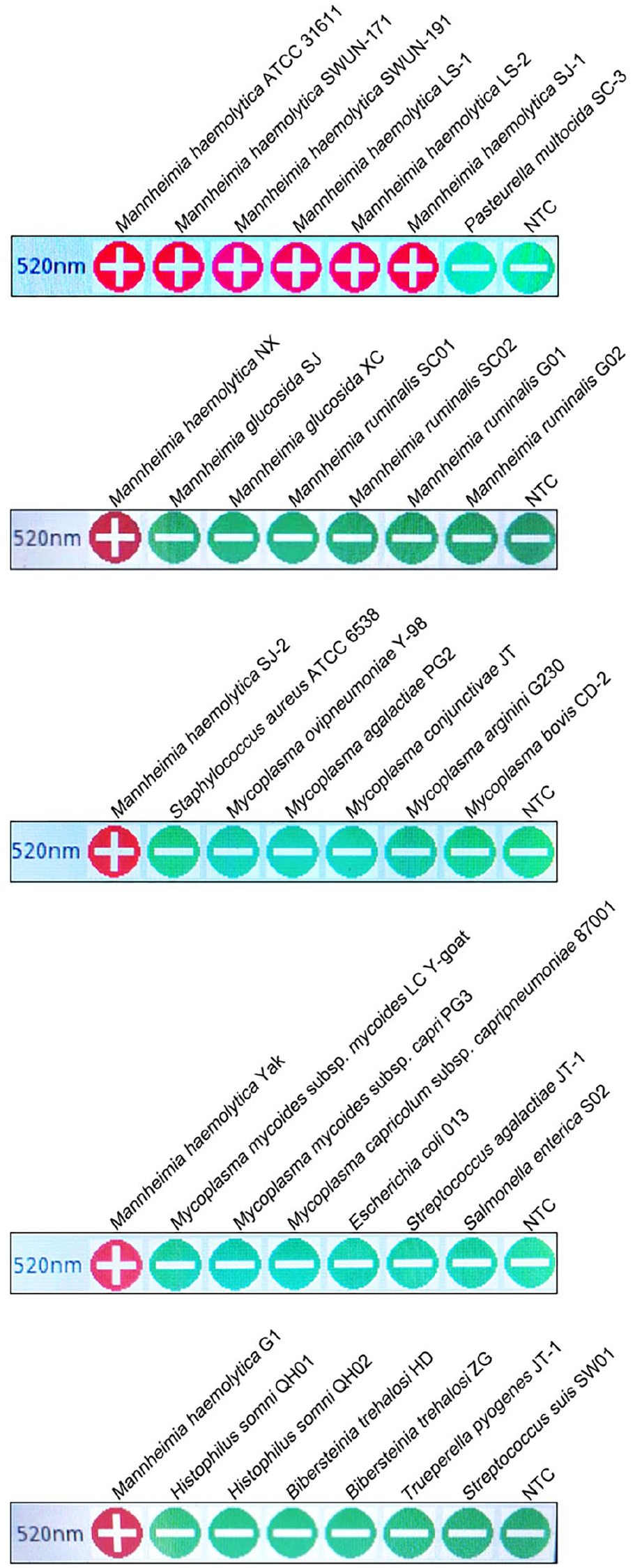
Specificity of the insulated isothermal PCR (iiPCR) assay. The iiPCR assay gave positive results for all Mannheimia haemolytica isolates used in our study and produced negative results for other bovine or ovine respiratory pathogens, including Histophilus somni, Bibersteinia trehalosi, Trueperella pyogenes, Streptococcus suis, and Mycoplasma spp. Given that the portable Pockit nucleic acid analyzer can test 8 samples in a single run, 5 separate runs were performed to test all strains, with a NTC (no template control) in each run.
The detection limits of our iiPCR assay for both M. haemolytica gDNA and bacterial loading were also evaluated. To determine the detection limit of our iiPCR assay for gDNA, the gDNA of strain NX was extracted and further quantified (DU800 spectrophotometer; Beckman Coulter). The copy number was then calculated based on the genome size (2,636,322 bp) of strain NX using an online copy number calculator (https://cels.uri.edu/gsc/cndna.html). Subsequently, 10-fold serial dilutions of the gDNA were prepared and then tested using the iiPCR assay. To assess the sensitivity of our iiPCR assay for bacterial amount, the bacterial number of a log phase M. haemolytica culture was first measured by counting the colony-forming units on the culture plate. A 10-fold serial dilution of a M. haemolytica culture was tested with the iiPCR assay. All dilutions were also tested in parallel with a published conventional gel-based PCR assay protocol 5 that targets the M. haemolytica ssa gene that encodes the serotype 1–specific antigen. The detection limits were 21 copies for gDNA and 17.2 cfu/mL for bacterial loading. These results are 10 and 100 times more sensitive than the conventional PCR assay, respectively (Fig. 2).
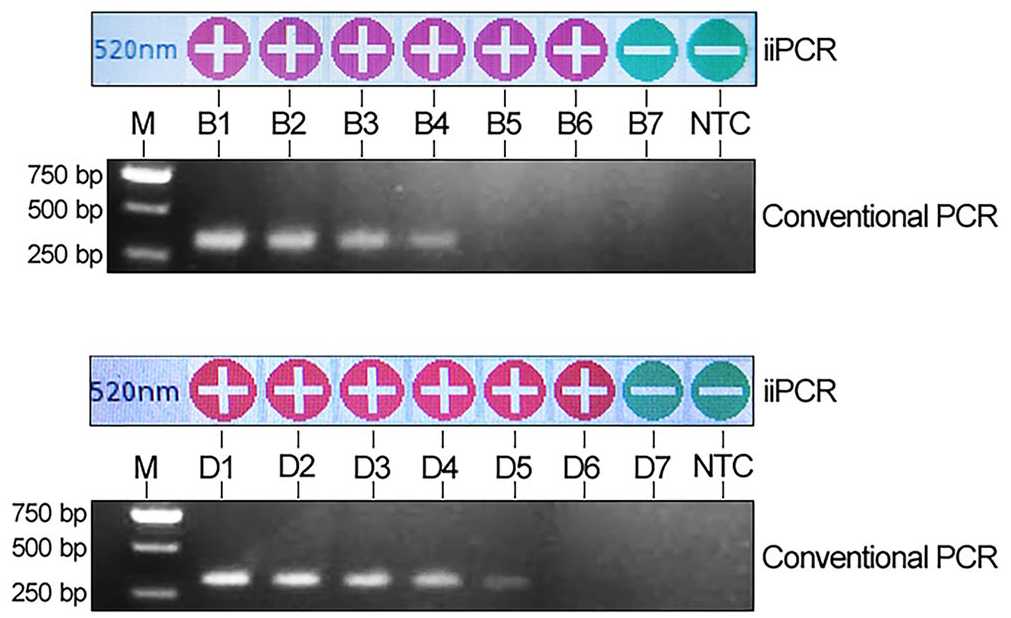
Sensitivity of the insulated isothermal PCR (iiPCR) assay. The iiPCR assay detected as few as 21 copies of genomic DNA and 17.2 cfu/mL of bacterial culture, which was 10 and 100 times more sensitive than conventional PCR, respectively. M = DNA molecular marker; B1–B7 = serial dilution of Mannheimia haemolytica culture (1.72 × 106–1.72 cfu/mL); D1–D7 = serial dilution of M. haemolytica genomic DNA (2.1 × 106–2.1 copies).
To validate the clinical application of our iiPCR assay, the DNA extracted from 67 bronchial swabs was tested using both the iiPCR and conventional PCR assays. 5 Our iiPCR assay was positive for 23 of 67 (34%) swabs; the gel-based PCR assay was positive for 19 of 67 (28%) swabs. All conventional PCR–positive samples were also detected successfully by our iiPCR assay. These results demonstrate that the iiPCR assay provides a better detection capability than the conventional gel-based PCR assay for clinical samples. To further confirm its accuracy, we cloned 8 PCR amplicons (including 4 that tested positive in our iiPCR assay but were negative in the conventional PCR assay, plus 4 randomly selected amplicons) into the pMD-T vector (TaKaRa) for further sequencing. As expected, all amplicons were 100% identical to M. haemolytica sodA, indicating that the results were reliable.
To compare the sensitivity of this iiPCR assay with bacterial culture, all 67 bronchial swabs were plated onto blood agar. Only for 15 of 23 (65%) iiPCR assay–positive samples were M. haemolytica recovered from the culture plates, indicating that our iiPCR assay was more sensitive than bacterial culture. Although our iiPCR assay is a reliable method for quickly screening M. haemolytica, the presence of this organism in bovine and ovine respiratory disease may not be sufficient to make a definitive diagnosis, given that, clinically, a coinfection of M. haemolytica with other respiratory bacterial or viral pathogens (such as H. somni, P. multocida, B. trehalosi, and Mycoplasma spp., bovine respiratory syncytial virus [Bovine orthopneumovirus], bovine herpesvirus 1 [Bovine alphaherpesvirus 1], etc.) is common. Further examinations, including bacterial culture and molecular tests for detecting viral pathogens, should also be considered to obtain additional information for final diagnosis. Performing antimicrobial susceptibility testing on M. haemolytica and other bacterial pathogens isolated from pneumonia cases is necessary to provide treatment options and enhance disease control in the herd.
Footnotes
Acknowledgements
We thank Dr. Quanling Du from Qinghai University, China for kindly providing Histophilus somni strains. We thank Ms. Amy Chen for editing our manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was funded by Sichuan provincial Key R&D projects (2021YFN0008).
