Abstract
A 4-bp deletion in the ATP-binding cassette subfamily B member 1 (ABCB1) gene, also referred to as the multidrug resistance gene (MDR1), produces stop codons that cause premature termination of P-glycoprotein 1 (P-gp) synthesis. Dogs with the homozygous mutation do not express functional P-gp, which increases their sensitivity markedly to many common veterinary drugs. We detected the nt230 (del4) ABCB1 mutation in Border Collie dogs in western Mexico with a simple and affordable primer-introduced restriction analysis PCR (PIRA-PCR). PIRA-PCR clearly identified all genotypes in our sample of 104 dogs. Genotype frequencies were 0.952 (wild/wild), 0.029 (wild/mut) and 0.019 (mut/mut). Allele frequencies were 0.033 (mutant alleles) and 0.966 (wild-type alleles). In this small subset of the Mexican dog population, we found a higher prevalence of the nt230 (del4) MDR1/ABCB1 gene mutation than reported in other countries.
P-glycoprotein 1 (permeability glycoprotein 1, or P-gp) is encoded by the ATP-binding cassette subfamily B member (ABCB1) gene, also referred to as the multidrug resistance 1 gene (MDR1), located on canine chromosome 14. This membrane glycoprotein acts as an important barrier to the distribution of drug substrates in select tissues, and it restricts xenobiotic access across the blood-brain barrier and the placenta. It also has important excretory functions in enterocytes, in the hepatocyte canalicular membrane, and in renal tubular epithelial cells. 15
A 4-bp deletion in gene ABCB1 changes the gene’s reading frame, producing stop codons that cause premature termination of P-gp synthesis and loss of P-gp function. 15 Dogs that are homozygous to the nt230 (del4) mutation do not express functional P-gp, and therefore have much greater sensitivity to many drugs transported by P-gp. 12 ABCB1 gene polymorphism is associated with increased susceptibility to adverse reactions and hypothalamic–pituitary–adrenal axis suppression for various commonly used veterinary drugs, resulting in buildup of these drugs in the brain and hence severe neurotoxicity.14,16
The primer-introduced restriction analysis PCR (PIRA-PCR) is a reliable and useful method that is widely used in diagnostic laboratories for extensive screening for SNP mutations using DNA polymerase and PCR machines.8,9 The method introduces an artificial restriction site into a PCR amplicon near the mutation site by the use of a primer with a single-base mismatch close to its 3′end, and then uses restriction-fragment length polymorphism (RFLP) analysis.11,13
Many assays are available today to identify this nt230 (del4) ABCB1 mutation.1,7,10,12,17,18 More practical assays have been described in the literature, such as amplified-fragment length polymorphism (AFLP), 17 which do not require an endonuclease digestion step because different length segments are amplified depending on the allele. But differentiating between 2 segments that differ by only 4 bp is at the lower limit of gel electrophoresis visual analysis and can lead to interpretation errors. In the previous study, 17 the amplified bands differing by 4 bp were detectable using polyacrylamide gels with silver stains but were undetectable using agarose gel electrophoresis with ethidium bromide stains. Multiplex PCR assays that amplify fragments of different sizes for each allele have also been used. 1 The use of different primers may necessitate different cycling profiles if a consensus cycling profile cannot be established. Also, another pair of primers is needed to amplify an endogenous region to verify the results and avoid misinterpretation from the lack of amplified products. With PIRA-PCR, it is possible to differentiate between mutant and wild-type ABCB1 genotypes using agarose gel electrophoresis with a nontoxic intercalating agent.
Our objective was to develop a simple PIRA-PCR genotyping assay for the nt230 (del4) polymorphism and use it to determine actual allele frequency in a sample of Border Collie dogs in Mexico. We collected blood samples from 104 Border Collie dogs after obtaining informed consent from the dog owners. Laboratory analysis was performed in the Animal Biotechnology Institute, Department of Animal Production, University of Guadalajara, Mexico. The study was approved by the Internal Bioethics Regulations of the Center for Biological and Agricultural Sciences, University of Guadalajara, Mexico (approval CC/NN11-12/00/2012).
To reduce possible contamination events resulting from the use of an electrophoresis gel detection test, filter tips were used. In addition, the reactions were carried out in a UV PCR cabinet to prevent the inadvertent transfer of nucleic acids. The genotypes of control samples were corroborated through the services of a commercial veterinary genetics laboratory (Helica, Monterrey, Nuevo Leon/Mexico).
We extracted DNA from blood samples (Quick-DNA universal kit; Zymo Research). Sample canine genotypes were corroborated by a previously described real-time PCR (rtPCR) technique. 17 PIRA-PCR tests were carried for samples in a 20-μL reaction mix (with ~100 ng DNA, 2 μL of 1× PCR buffer containing 0.395 μL of 20 mM MgCl2, 1 μL of 10 mM dNTP mix, 0.5 μL of DNA polymerase; DreamTaq PCR kit, Thermo Scientific), 1.25 pmol of both primers, and double-distilled water for the remaining volume.
Primers were designed based on the ABCB1 gene sequence in GenBank (AF045016.1). The PCR protocol consisted of initial denaturation at 94°C for 5 min, 10 cycles of 30 s at 94°C followed by 30 s at 56°C and 50 s at 72°C, subsequently 25 cycles of 30 s at 94°C followed by 30 s at 65°C and 50 s at 72°C, and finally extension at 72°C for 5 min. We amplified a 68-bp fragment by introduction of site-restriction in the forward primer, exchanging a cytosine with an adenine in the first nucleotide of the last codon before deletion of 4 ATAG nucleotides in the ABCB1 gene sequence (Table 1). The deletion present in the PCR-amplified fragment creates a recognition site for HindIII endonuclease (AAGCTT) via the union of the modified codon and the next codon preceding deletion (Fig. 1). Samples were then digested with HindIII restriction endonuclease (New England Biolabs) in a gradient thermocycler (Techne). We subjected the PCR analysis products to 4% agarose gel electrophoresis with 3 μL of intercalating nucleic acid stain (Biotium) at a ratio of 1:100.
Characteristics of primers used in our study of genotyping of the canine ABCB1 gene with a primer-introduced restriction analysis PCR assay.
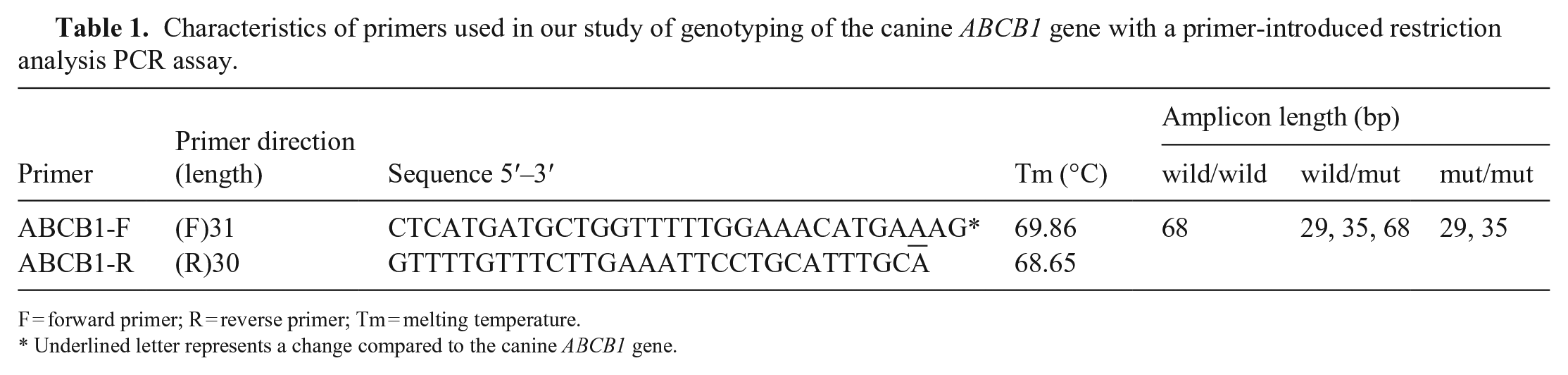

F = forward primer; R = reverse primer; Tm = melting temperature.
Underlined letter represents a change compared to the canine ABCB1 gene.
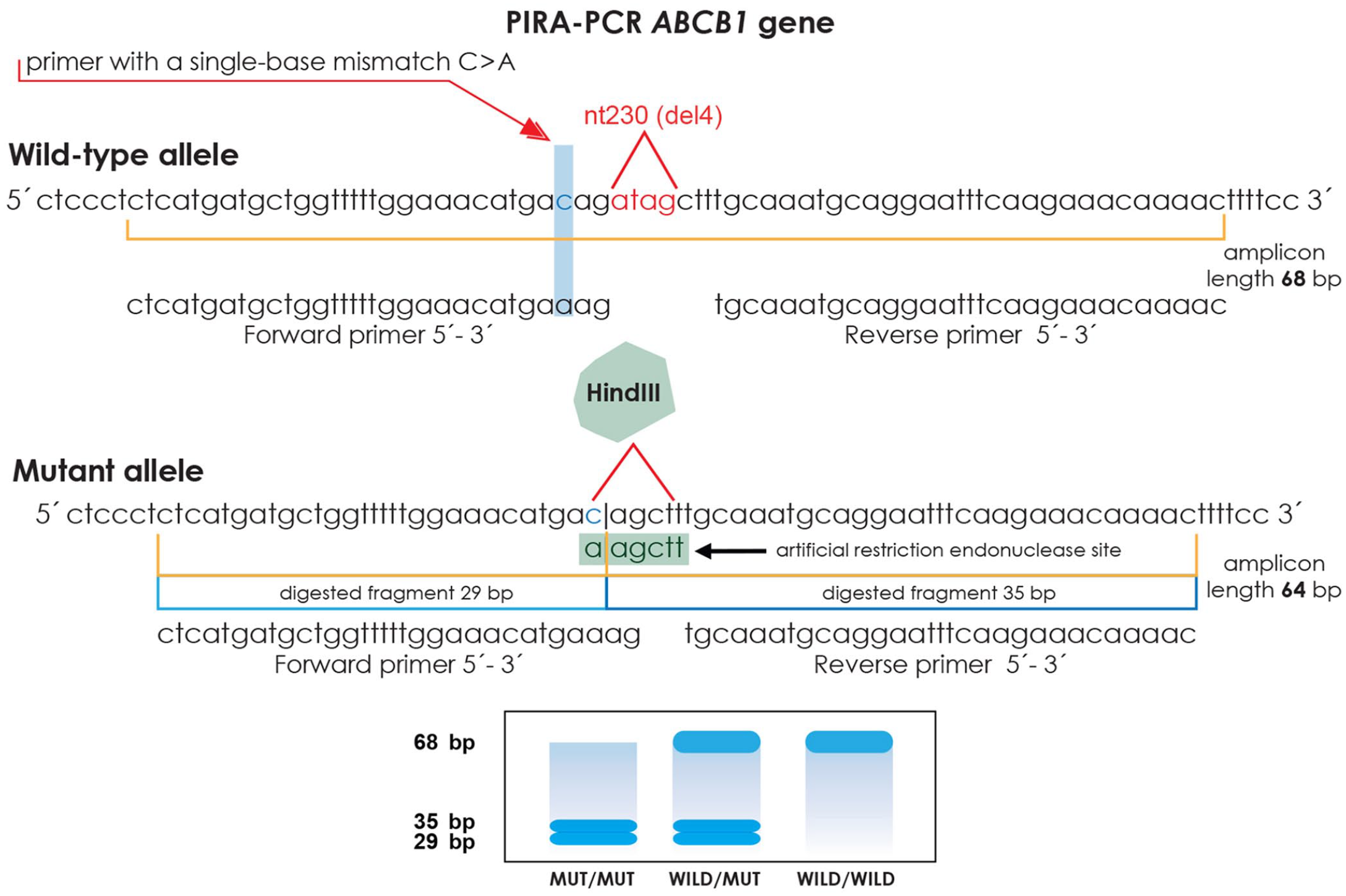
Schematic presentation of the ABCB1 gene, depicted by the sequences closest to the nt230 (del4) mutation site of the wild-type and mutant alleles, and the nucleotide sequences of forward and reverse primers. The nucleotide highlighted in light blue shows the mismatched primer sequence for the introduced restriction site. At the bottom, the expected gel image.
All of the dogs in our study were genotyped correctly with the PIRA-PCR assay, and had the same genotype obtained using a PCR technique. The mutant homozygous genotype (mut/mut) produced a 64-bp product (given the 4-bp deletion) that was digested by the HindIII endonuclease creating bands of 29 bp and 35 bp. For the heterozygous genotype (wild/mut), we observed digested bands of 29 bp and 35 bp, and an undigested band of 68 bp (Fig. 2). An ABCB1-1Δ allele was found in 2 homozygous mut/mut dogs and 3 heterozygous wild/mut dogs; 99 dogs had the wild/wild genotype. Genotype frequencies were as follows: 0.952 (wild/wild), 0.029 (wild/mut), and 0.019 (mut/mut); estimated allele frequencies were 0.033 for mutant allele and 0.966 for the wild-type allele. PIRA-PCR electrophoresis migration patterns displayed wider band separation from enzyme digestion, enabling clear identification of the mutant allele, compared to end-point PCR in which amplicons of the alleles differ in size by only 4 bp, making interpretation more difficult and extending the amplicon migration period in the gene.
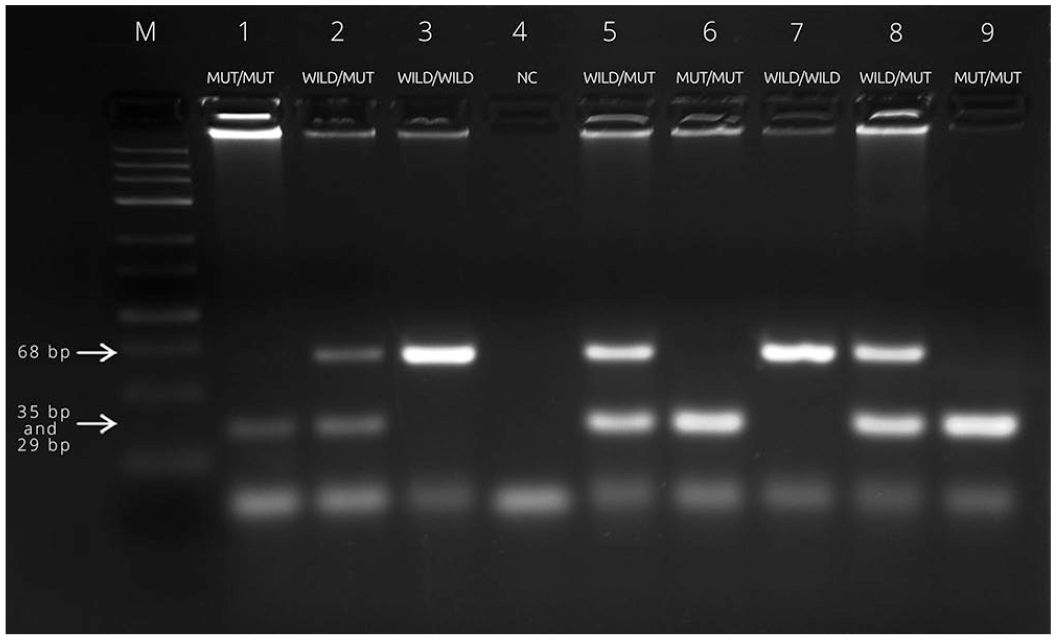
Agarose gel electrophoresis of primer-introduced restriction analysis PCR products to detect the nt230 (del4) ABCB1 mutation associated with P-glycoprotein 1 (P-gp) substrate drug sensitivity. Lanes: M = molecular marker (50 bp; Life Technologies); 1–3 = control samples for dogs with representative patterns for all 3 genotypes (MUT/MUT = affected; WILD/MUT = carrier; WILD/WILD = wild-type); 4 = NC, represents a negative PCR control without a template; 5, 8 = carrier Border Collie dog (29, 35, and 68 bp); 7 = homozygous mutation–free Border Collie with a 68-bp fragment; 6, 9 = homozygous mutation Border Collie dogs (29 and 35 bp); a difference of 6 bp between digested fragments of 29 and 35 bp is seen as a single band.
The presence of the nt230 (del4) ABCB1 gene mutation in Border Collies varies by country. In contrast with other studies, we obtained a considerably higher frequency of 0.033 for this mutation. Only the study from Israel 2 had similar results: a frequency of 0.048 for 261 purebred Border Collies and a frequency of 0.003 for 183 cross-breed dogs. A much lower frequency was reported in Japan, with a mutant allele frequency of 0.0025 for 407 dogs sampled. 17 A study from continental Europe did not find any carriers of the mutation. 5 This absence of mutation is also reported in other studies,1,4,6,10 but these studies genotyped a very small number of Border Collie dogs within the total population, probably because they emphasized animals with a higher historical frequency of the mutation. Thus, it is possible that the mutation was present in other Border Collie populations but went undetected given the small sample size.
By providing a new recognition site for the HindIII enzyme, our PIRA-PCR generated 3 more highly distinguishable digestion fragments of 29, 35, and 68 bp for the different genotypes, substantially improving interpretation. Our assay provides a rapid genotyping test for ABCB1 using a small quantity of blood (100 µL). Designing simpler, more accessible assays with lower setup costs can facilitate diagnosis in breeds where P-gp substrate drug sensitivity is suspected, as is the case with epileptic Border Collies that have the deletion. 3 Correctly identifying the mutation may lead to selecting more suitable drugs in countries such as Mexico where access to more complex and costly methods is still difficult. Given the low prevalence of the mutation in Border Collie dogs, selection assisted by genotyping could eliminate the mutation without affecting the population’s genetic diversity and without increased risk of unintentional selection as a result of possible concomitant negative characteristics. 4
Footnotes
Acknowledgements
We thank Rogelio Granja who designed and edited the figures in this manuscript.
Data sharing statement
All available data can be obtained by contacting the corresponding author.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was funded by the University of Guadalajara through project P3E-255382 with the support of the Institute of Animal Biotechnology in the department of Department of Animal Production, University Center of Biological and Agricultural Sciences, which provided the facilities and equipment to carry out our study.
