Abstract
The origin of chronic wasting disease (CWD) in cervids is unclear. One hypothesis suggests that CWD originated from scrapie in sheep. We compared the disease phenotype of sheep-adapted CWD to classical scrapie in sheep. We inoculated sheep intracranially with brain homogenate from first-passage mule deer CWD in sheep (sCWDmd). The attack rate in second-passage sheep was 100% (12 of 12). Sheep had prominent lymphoid accumulations of PrPSc reminiscent of classical scrapie. The pattern and distribution of PrPSc in the brains of sheep with CWDmd was similar to scrapie strain 13-7 but different from scrapie strain x124. The western blot glycoprofiles of sCWDmd were indistinguishable from scrapie strain 13-7; however, independent of sheep genotype, glycoprofiles of sCWDmd were different than x124. When sheep genotypes were evaluated individually, there was considerable overlap in the glycoprofiles that precluded significant discrimination between sheep CWD and scrapie strains. Our data suggest that the phenotype of CWD in sheep is indistinguishable from some strains of scrapie in sheep. Given our results, current detection techniques would be unlikely to distinguish CWD in sheep from scrapie in sheep if cross-species transmission occurred naturally. It is unknown if sheep are naturally vulnerable to CWD; however, the susceptibility of sheep after intracranial inoculation and lymphoid accumulation indicates that the species barrier is not absolute.
Chronic wasting disease (CWD) and scrapie are transmissible spongiform encephalopathies (TSEs) of cervids 43 and sheep, 23 respectively. TSEs are characterized by fatal neurodegeneration of the host.28,30 This is associated with the accumulation of a misfolded protein (PrPSc) derived from a normal variant (PrPC) that is encoded by the prion protein gene (PRNP).8,29 Both scrapie and CWD are considered to be naturally infectious to susceptible sheep or deer, respectively. In the United States, at least 2 distinct strains of classical scrapie have been identified: strains 13-7 and x124. 26
The origins of CWD are unclear, but the disease was first identified in captive mule deer (Odocoileus hemionus). 43 Subsequently, wild and captive white-tailed deer (Odocoileus virginianus) and elk (Cervus elaphus) became infected, and the presence of the disease has spread or become recognized throughout North America,4,7 Norway, 2 Finland, 20 Sweden, 27 and South Korea. 35 Speculation about the origins of CWD has varied, but one theory suggests that sheep scrapie jumped species to mule deer. 42 The theory is loosely supported by in vitro studies that demonstrate intermediate conversion of sheep PrPC to PrPSc by infectious CWD prions 32 and similarities in their western blot glycoform patterns. 31 In vivo studies investigating interspecies transmission of the CWD agent to sheep are incomplete.
The susceptibility of sheep to the scrapie agent is determined by naturally occurring polymorphisms in the PRNP gene. 3 Alanine (A) and arginine (R) amino acid substitutions at codons 136 and 171 are associated with relative resistance to scrapie; whereas, valine (V) and glutamine (Q) are associated with susceptibility.10,41 To a lesser extent, codon 154 also influences the susceptibility to scrapie. These 3 polymorphisms are frequently considered together as a haplotype designated in triplicate by the amino acids in each allele. The degree of relative susceptibility compared to relative resistance is V136R154Q171 > ARQ > ARR.1,38
In a previous study, we demonstrated the susceptibility of sheep to mule deer CWD (CWDmd) after intracranial inoculation. 18 We evaluated and report here the second passage of sheep CWDmd to sheep, comparing it to classical scrapie in sheep. Our work is significant because it describes similarities between scrapie and sheep-passaged CWDmd (sCWDmd) and offers insight into the possible origins of CWD.
Materials and methods
Ethics statement
The animal experiments were conducted in animal biosafety level 2 spaces that were inspected and approved for importing prion agents by the U.S. Department of Agriculture (USDA), Animal and Plant Health Inspection Service, and Veterinary Services. The studies were done in accordance with the Guide for the Care and Use of Laboratory Animals (Institute of Laboratory Animal Resources, National Academy of Sciences, Washington, DC, USA) and the Guide for the Care and Use of Agricultural Animals in Research and Teaching (Federation of Animal Science Societies, Champaign, IL, USA). The protocols were approved by the Institutional Animal Care and Use Committee at the National Animal Disease Center (protocol 3696).
Animal use
Animals for our experiment were sourced from the scrapie-free flock housed at the USDA National Animal Disease Center (NADC) in Ames, IA, USA. The PRNP gene of 12 Suffolk sheep was amplified by PCR and sequenced by the Sanger sequencing method. The predicted PRNP genotype of each sheep was homozygous at the following polymorphic sites: M112, G127, A136, M137, S138, L141, R151, R154, M157, N176, H180, Q189, T195, T196, R211, Q220, and R223. Only codons 136 and 171 contained polymorphisms; we used 3 different PRNP genotypes of sheep in our experiment: VRQ/ARQ, ARQ/ARQ, and ARQ/ARR.
Inoculum preparation and intracranial inoculation
The transmission of CWD from mule deer to sheep was investigated previously by intracranial inoculation. 18 In that experiment, a Suffolk sheep with the VRQ/ARQ genotype developed clinical neurologic disease and had detectable PrPSc in the brain and lymphoid tissue by immunohistochemistry (IHC). That sheep was used as the inoculum for our experiment. The inoculum was made as described previously. 6 Brain was homogenized mechanically and prepared to a 10% w/v solution with phosphate-buffered saline (PBS) and 100 µg/mL of gentamicin.
Intracranial inoculation of 7-mo-old Suffolk sheep was performed as follows. Animals were sedated with xylazine, and the dorsal frontal region of the head was prepared by clipping and surgically scrubbing with chlorohexidine and alcohol. A 1-mm hole was drilled midline between frontal bones near the intersection with the parietal bone. Subsequently, a 22-ga needle was placed through the hole to the ventral floor of the calvarium. As the needle was withdrawn, 1 mL (0.1-g equivalent brain material) of inoculum was injected slowly. Tissue glue was used to close the skin incision. The sheep were housed in an animal biosafety level 2 facility where they were monitored daily. At the onset of clinical signs unequivocally consistent with scrapie or at the experimental endpoint of 60 mo post-inoculation, the animals were euthanized and autopsied.
Sample collection
During postmortem examination, a standard tissue set was collected for microscopic evaluation of each animal. From the central nervous system (CNS), brain, spinal cord, and retina were collected. Lymphoreticular tissues were collected and included spleen, tonsils (palatine and pharyngeal), lymph nodes (retropharyngeal [RPLN], mesenteric, popliteal, prescapular), gastrointestinal-associated lymphoid tissues (recto-anal and ileum), and third eyelid mucosa. Other collected tissues were esophagus, forestomaches, jejunum, pancreas, salivary gland, adrenal gland, pituitary, thyroid, liver, lung, kidney, urinary bladder, nose skin, trigeminal ganglion, sciatic nerve, nasal turbinate, and muscles (heart, tongue, diaphragm, biceps femoris, triceps, masseter, and psoas major). Tissues were placed and stored in 10% neutral-buffered formalin (globes in Bouin fixative). Portions of the RPLN, tonsils, and brain were frozen for western blot and enzyme immunoassay (EIA).
Microscopic and immunohistochemical evaluation
Fixed tissue samples were processed routinely and stained with hematoxylin and eosin for microscopic examination. 6 Methods for animal procedures and immunohistochemical detection of PrPSc have been described in detail previously. 26 Briefly, all tissues were also evaluated for the presence of PrPSc by applying an anti-PrPSc antibody cocktail that contained F89/160.1.5 and F99/97.6.1 monoclonal antibodies. The antibodies were applied with an automated processor at a concentration of 5 µg/mL each.
We compared the PrPSc distribution in the brains of CWDmd-infected sheep with scrapie-infected sheep. Immunolabeling patterns have been described in detail previously.12,33 For comparison to scrapie in sheep, we used archived tissue samples from sheep that had been inoculated intracranially with either the x124 or 13-7 U.S.-derived classical scrapie strains. 26 The animals and genotypes selected for comparison were chosen to match the inoculation route and amount used in our experiment.
Enzyme immunoassay
For rapid discrimination of positive samples, we used a commercial EIA kit (HerdChek; Idexx). Frozen sections of the palatine tonsil, RPLN, and brainstem at the level of the obex were homogenized and concentrated to a 20% w/v solution with PBS. The assay was conducted according to the kit instructions, and cutoff numbers were determined as values equal to the mean optical densities of the negative controls +0.180.
Western blot
To evaluate the molecular characteristics of second-passage CWDmd in sheep, western blot detection of PrPSc was performed on frozen sections of brainstem at the level of the obex. Samples were prepared as a 20% w/v tissue homogenate using PBS; then the samples were treated with proteinase K, and acetone precipitated as described previously. 5 For each sample, the brain weight equivalent, 0.5–1.5 mg Eq, was loaded into individual wells of a 12% Bis-Tris polyacrylamide protein gel (NuPAGE; Thermo Fisher) and separated by electrophoresis. For immunoblotting of the electrophoretically separated PrPSc, we used the murine monoclonal antibody P4 (diluted to a final concentration of 0.1 µg/mL; R-Biopharm), a biotinylated anti-mouse secondary antibody (diluted to a final concentration of 0.1 µg/mL; Biotinylated anti-mouse IgG, Amersham), and streptavidin conjugated to horseradish peroxidase (diluted to a final concentration of 0.1 µg/mL; streptavidin–horseradish-peroxidase (HRP) conjugate, Amersham). Detectable chemiluminescent signal was created with a commercial HRP substrate (Pierce ECL-Plus; Thermo Fisher), and images were acquired with a digital imaging system (G:BOX Chemi-XT4; Syngene). Two sheep from each TSE group and genotype were blotted with 3 technical replicates. Only one animal/sample from each inoculum source (pooled mule deer and first-passage sheep CWD) was available for assessment. There were no available samples for ARQ/ARR sheep with x124 scrapie.
Resultant immunoblots were analyzed in parallel adhering to identical procedures for band identification and background subtraction. Western blots of TSEs display 3 distinct bands that correspond to unglycosylated, monoglycosylated, and diglycosylated PrPSc. The molecular characteristics of TSEs are defined by the molecular masses (migration distances) and the relative density corresponding to each band (glycoprofile). We measured the relative proportion of diglycosylated, monoglycosylated, and unglycosylated banding for each sample (Image Lab v.6.0.1 for macOS; Bio-Rad). The molecular weights of each fragment were determined by using a protein molecular weight standard (Low range biotinylated SDS-PAGE standard; Bio-Rad). Comparisons and statistical analysis of the resulting data were evaluated (Prism v.8.1.2 for macOS; GraphPad). A one-way ANOVA using the Tukey method for multiple comparisons (α = 0.05) was used to evaluate diglycosylated band densities and the molecular weights of all glycotypes of PrPSc.
Results
Attack rate and incubation period
Every sheep (12 of 12) had PrPSc immunoreactivity in the brain, hence, a 100% attack rate. The mean incubation periods for VRQ/ARQ and ARQ/ARQ sheep were 14.6 and 11.4 mo, respectively (Fig. 1). All of the ARQ/ARR sheep survived to the end of the experiment, but they had detectable PrPSc in the brain (Fig. 2).
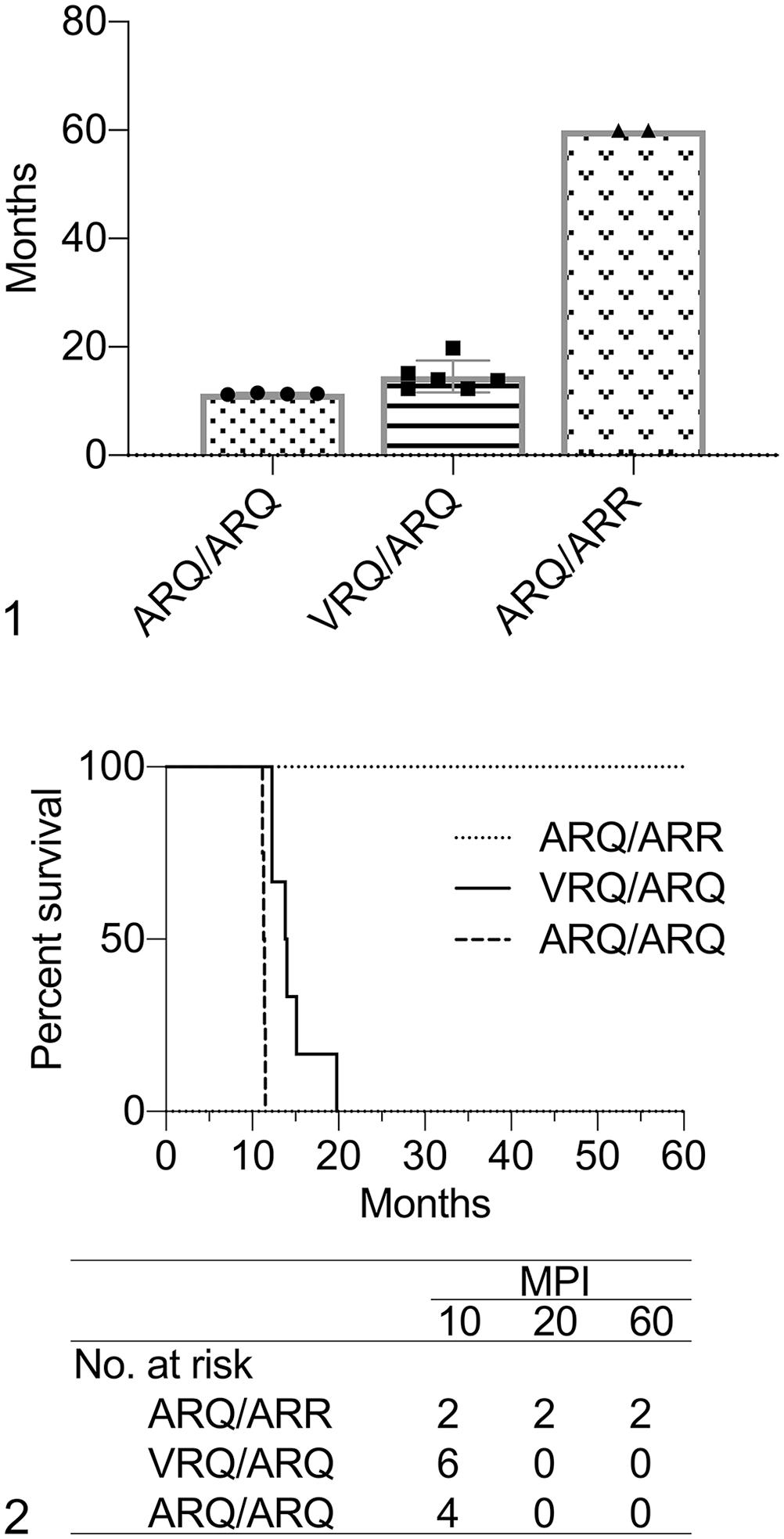
Analysis of survival data in sheep inoculated intracranially with sheep-passaged CWD from mule deer.
Immunoreactivity for PrPSc and tissue distribution
All inoculated sheep had PrPSc in the CNS. In VRQ/ARQ and ARQ/ARQ sheep, there was widespread distribution of PrPSc throughout the lymphoreticular system. These 2 genotypes of sheep also had detectable PrPSc in muscle and other peripheral nervous system tissues.
All 3 genotypes of sheep had positive immunolabeling for PrPSc within the brainstem at the level of the obex that was restricted mostly to gray matter and scant within the intersecting white matter tracts. Common patterns of PrPSc labeling in sheep inoculated intracranially with CWDmd included intraneuronal, perineuronal, intraglial, coarse particulate, and less frequently, stellate patterns. VRQ/ARQ sheep had the densest accumulation of PrPSc, which corresponded with the appearance of more prominent linear-type immunolabeling. A similar distribution of PrPSc types was observed in the rostral brainstem that was largely limited to gray matter tracts with an increasing prevalence of stellate and linear patterns.
In the cerebral cortices, PrPSc accumulation varied between genotypes. VRQ/ARQ and ARQ/ARQ sheep had some white matter accumulation (Figs. 3, 4), but most PrPSc was in perineuronal, linear, stellate, intraglial, and coarse particulate patterns in layers IV–VI of the neocortices. Linear immunolabeling was present in cerebral cortical layers II and III. The PrPSc accumulation differed in ARQ/ARR sheep (Fig. 5). These sheep had mostly white matter accumulation that appeared as stellate, coarse particulate, intraglial, and perivascular glial patterns. There was minimal PrPSc in the gray matter of the neocortex; the immunolabeling observed in this location included fine particulate in the neuropil and scattered intraglial patterns. In the cerebral cortex, there was intraneural PrPSc infrequently in the VRQ/ARQ and ARQ/ARQ sheep, and none in the ARQ/ARR sheep.
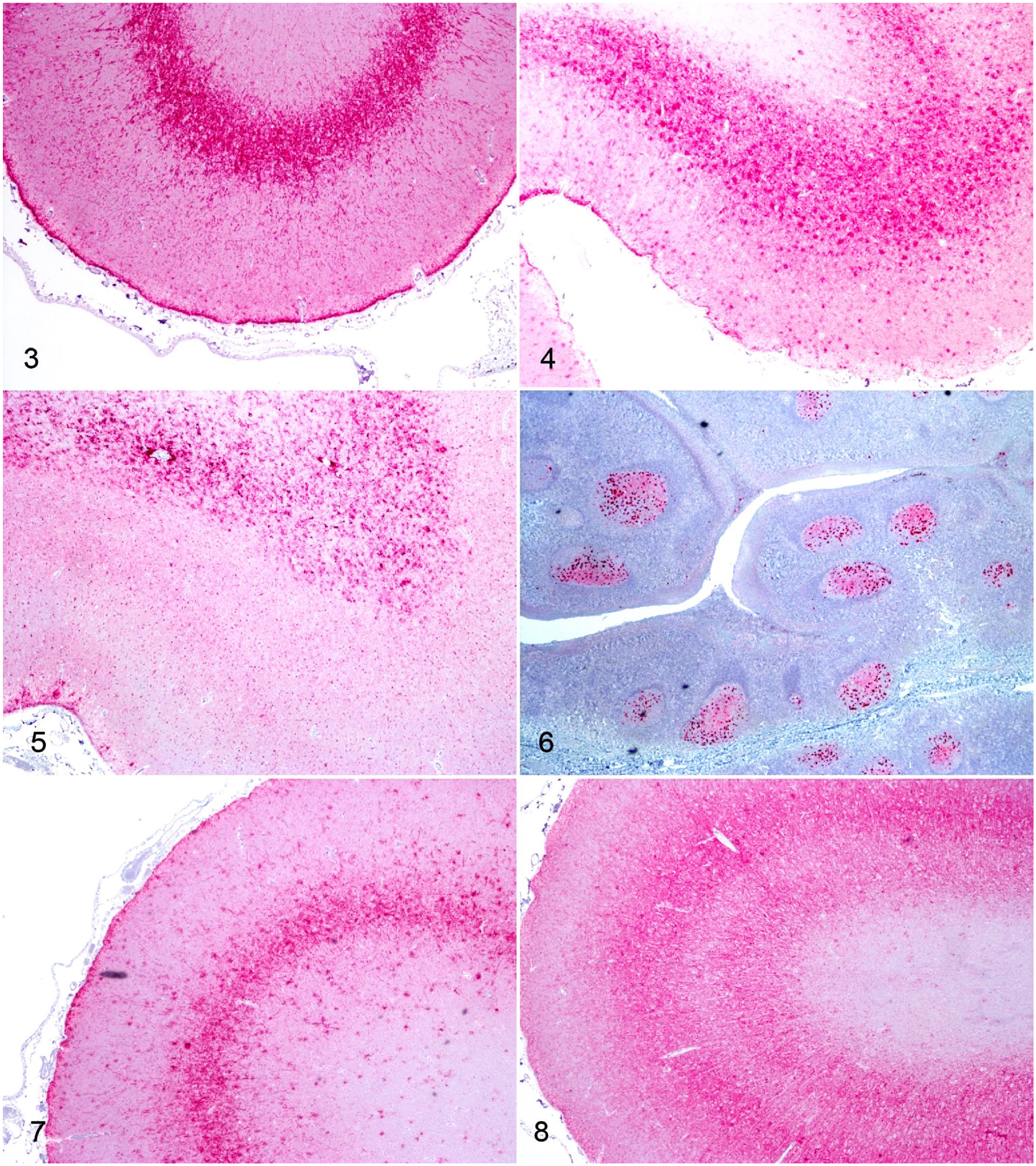
Immunohistochemistry for PrPSc (red) in sheep inoculated intracranially with either sheep-passaged CWD from mule deer (CWDmd) or classical scrapie (strain 13-7 or x124). Original objective 4×.
The distribution of PrPSc in the cerebellum was similar among all genotypes. PrPSc occurred mostly in the granular cell layer and around Purkinje cells. Stellate-type PrPSc was present in the molecular and white matter; although immunolabeling in the white matter was mostly absent in the ARQ/ARQ sheep. Overall, the VRQ/ARQ sheep had denser immunolabeling of PrPSc in the cerebellum than the other genotypes. PrPSc was present in the spinal cords of all genotypes of sheep at the cervical, thoracic, and lumbar levels. The ARQ/ARR sheep had less PrPSc than the other 2 genotypes.
In both the VRQ/ARQ and ARQ/ARQ sheep, there was widespread distribution of PrPSc detectable in lymphoid tissue with IHC (Table 1). We observed PrPSc in the palatine (Fig. 6) and pharyngeal tonsils, RPLN, spleen, mesenteric lymph node, gastrointestinal mucosa-associated lymphoid tissue of the small intestine, prescapular and popliteal lymph nodes, and recto-anal mucosa-associated lymphoid tissue. Immunolabeling of PrPSc was also present in the enteric nervous system, observed most frequently in the ileum. In 3 of 5 VRQ/ARQ sheep, 4 of 4 ARQ/ARQ sheep, and 1 of 2 ARQ/ARR sheep, there was at least 1 lymphoid follicle in the nictitating membrane. Of these samples with representative follicles, 4 of 8 sheep were immunoreactive for PrPSc.
Immunologic testing of lymphoid tissues and enteric nervous system in sheep inoculated with sheep-adapted CWD from mule deer.
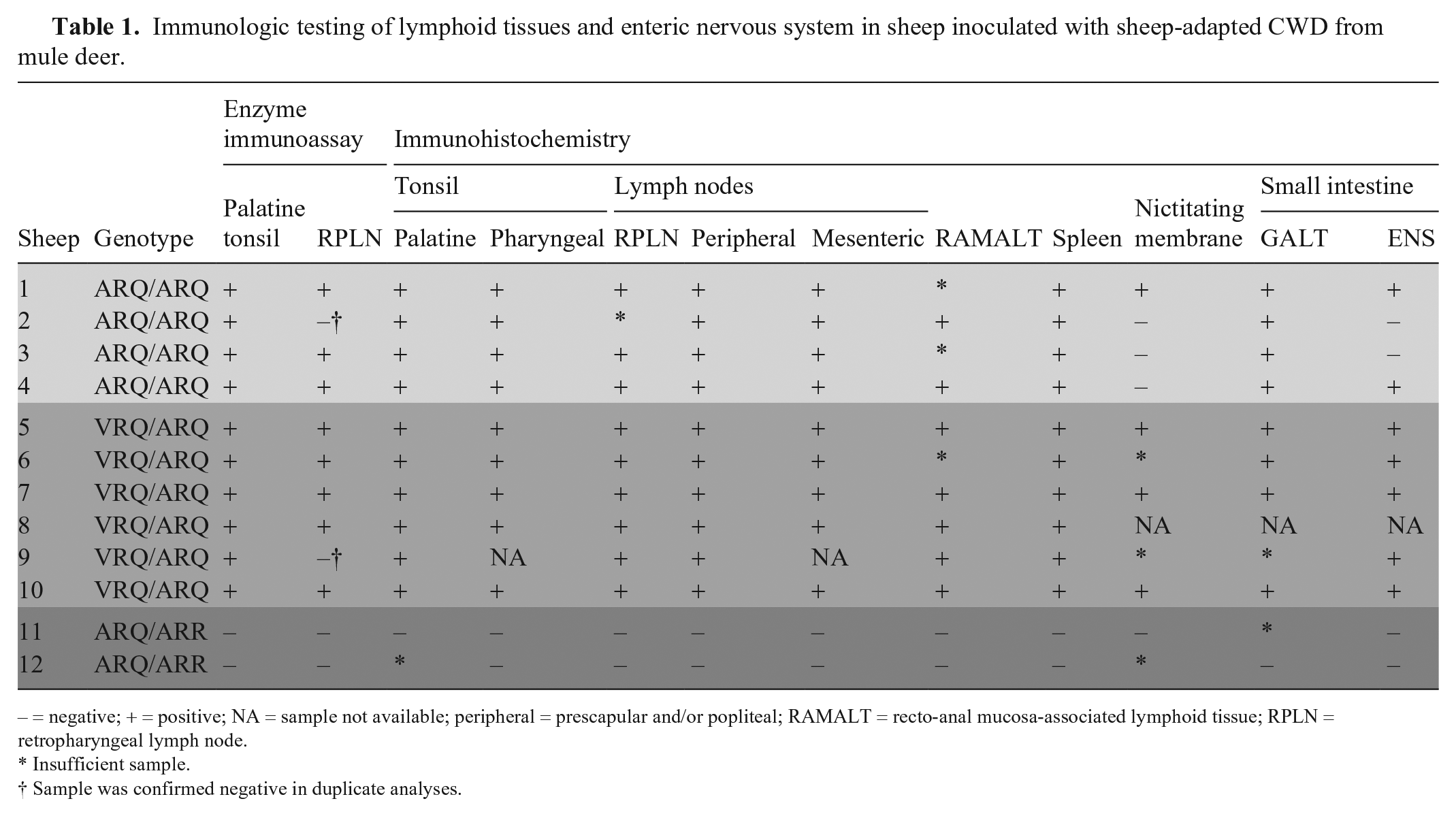
– = negative; + = positive; NA = sample not available; peripheral = prescapular and/or popliteal; RAMALT = recto-anal mucosa-associated lymphoid tissue; RPLN = retropharyngeal lymph node.
Insufficient sample.
Sample was confirmed negative in duplicate analyses.
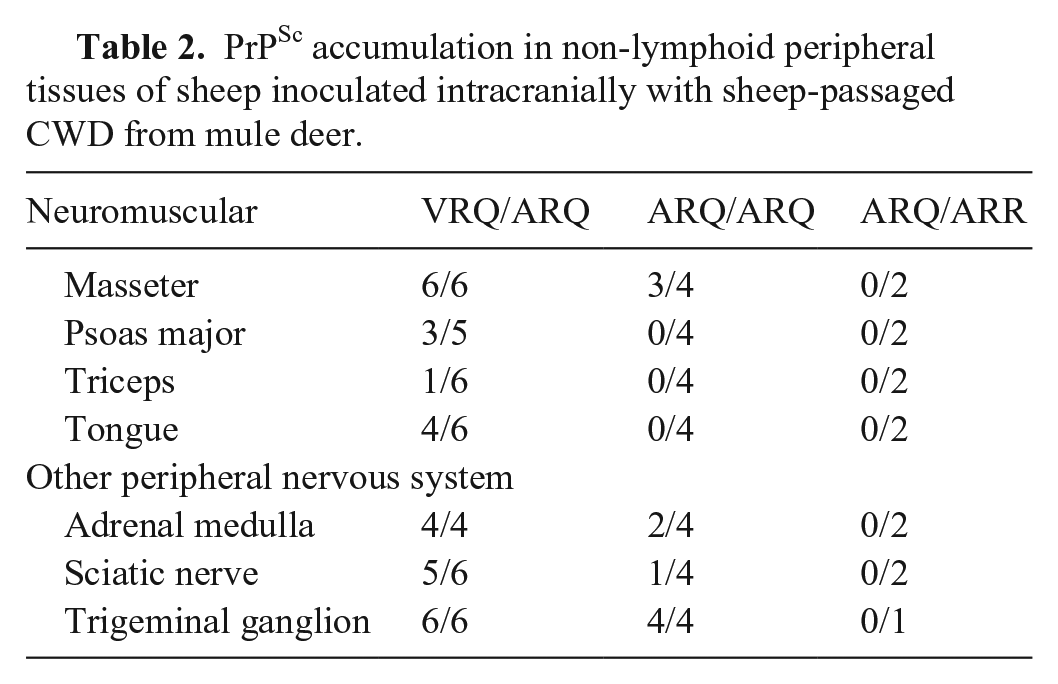
We observed PrPSc in the neuromuscular spindles of the masseter, psoas major, and triceps muscles. VRQ/ARQ sheep had the most prevalent muscle accumulation (Table 2); whereas, ARQ/ARQ sheep only had immunoreactive PrPSc detectable in the masseter muscles. ARQ/ARR sheep did not have any immunoreactive PrPSc in muscle. Several of the VRQ/ARQ sheep had PrPSc in nerves of the tongue. VRQ/ARQ and ARQ/ARQ sheep also had observable PrPSc in the adrenal medulla, sciatic nerve, trigeminal ganglion, and pituitary gland. One ARQ/ARR sheep had PrPSc in the pars nervosa of the pituitary gland.
PrPSc accumulation in non-lymphoid peripheral tissues of sheep inoculated intracranially with sheep-passaged CWD from mule deer.
PrPSc immunoreactivity patterns in brains of sheep with sCWDmd compared to sheep with U.S. classical scrapie strains
The PrPSc immunolabeling in ARQ/ARQ sheep was broadly similar between CWDmd and scrapie 13-7 in the frontal and parietal cerebral cortices, and they were distinct compared to scrapie x124. Sheep inoculated with sCWDmd and classical scrapie 13-7 both had abundant stellate immunolabeling in the cerebral cortex. This was most evident in cortical layers IV–VI for sCWDmd (Fig. 4), but in 13-7–inoculated sheep, the densest accumulation of PrPSc was centered on layer IV (Fig. 7). The pattern of stellate immunolabeling seen with 13-7 and sCWDmd was mostly absent in sheep inoculated with classical scrapie x124 (Fig. 8). Instead, these sheep had prominent coarse particulate immunolabeling in the neuropil.
Similarly, in the cerebellum, PrPSc immunolabeling in sheep with scrapie x124 was different than scrapie 13-7 and sCWDmd. Most labeling occurred in the molecular layer and consisted of extracellular particulate and stellate patterns with some scattered intraglial PrPSc. The labeling in sheep inoculated with scrapie 13-7 included dense accumulations around Purkinje cells. Both the molecular and granular cell layers contained abundant particulate, coalescing particulate, and stellate PrPSc. In sheep inoculated with sCWDmd, cerebellar immunolabeling was less dense and composed predominantly of the particulate pattern that centered on Purkinje cells and extended widely into the molecular and granular cell layers. There was mostly stellate immunolabeling in the molecular layers.
Enzyme immunoassay
Every animal in the study had positive EIA results from their obex sample. The RPLN was EIA-positive in 5 of 6, 3 of 4, and 0 of 2 sheep from the VRQ/ARQ, ARQ/ARQ, and ARQ/ARR genotypes, respectively. Sheep 2 (ARQ/ARQ) and sheep 9 (VRQ/ARQ) had negative RPLNs by EIA that were confirmed on duplicate tests. The RPLN from sheep 9 was positive by IHC; however, there was insufficient sample to confirm the status of sheep 2 by IHC. The palatine tonsil was positive in each of the VRQ/ARQ and ARQ/ARQ sheep. All lymphoid tissue was negative by EIA in ARQ/ARR sheep.
Western blot
The molecular masses of PrPSc western blot bands were similar among sCWDmd, scrapie 13-7, scrapie x124, and mule deer CWD groups; the diglycosylated, monoglycosylated, and unglycosylated bands approximated 28, 24, and 20 kDa, respectively (Fig. 9). There were no detectable differences in the molecular weights of PrPSc between the mule deer inoculum pool, first-passage sCWDmd, and second-passage sCWDmd. Furthermore, no significant differences were present between scrapie and sCWDmd. There were rare differences between sheep scrapie strains x124 and 13-7. The average weight of unglycosylated PrPSc in VRQ/ARQ sheep with scrapie x124 was 21.5 kDa. This resulted in a statistical difference between VRQ/ARQ sheep with scrapie x124 and ARQ/ARQ sheep with scrapie x124 (p = 0.0028) and ARQ/ARR sheep with scrapie 13-7 (p = 0.0037).
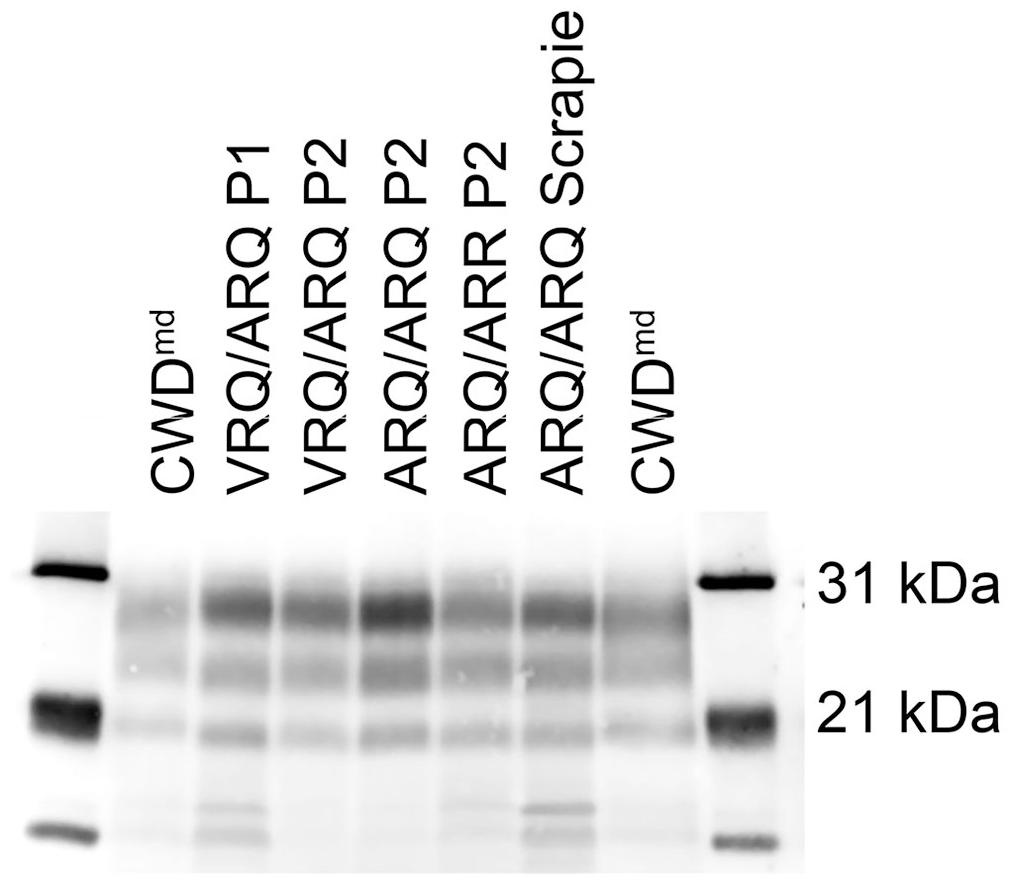
Representative western blot for PrPSc using mouse monoclonal antibody P4. Left to right lanes include mule deer CWD inoculum (CWDmd), first-passage (P1) sheep CWDmd in a VRQ/ARQ sheep, second-passage (P2) sheep CWDmd in different genotypes, and classical scrapie strain 13-7 in a sheep with the ARQ/ARQ genotype.
The relative proportions of diglycosylated and monoglycosylated bands were plotted to visualize differences between host genotype and TSE inoculum type (Fig. 10). Passage of mule deer CWD to sheep (P1) followed by a second passage in sheep (P2) had no observable effect on the glycoprofile. Furthermore, there were no significant differences in the glycoprofile of sCWDmd and scrapie when analyzing each sheep genotype and TSE inoculum group. Independent of host genotype (Fig. 11), there was a difference between scrapie x124 and scrapie 13-7 (p = 0.0013), and between scrapie x124 and sCWDmd (p = 0.0087). No significant difference was detectable between scrapie 13-7 and sCWDmd.
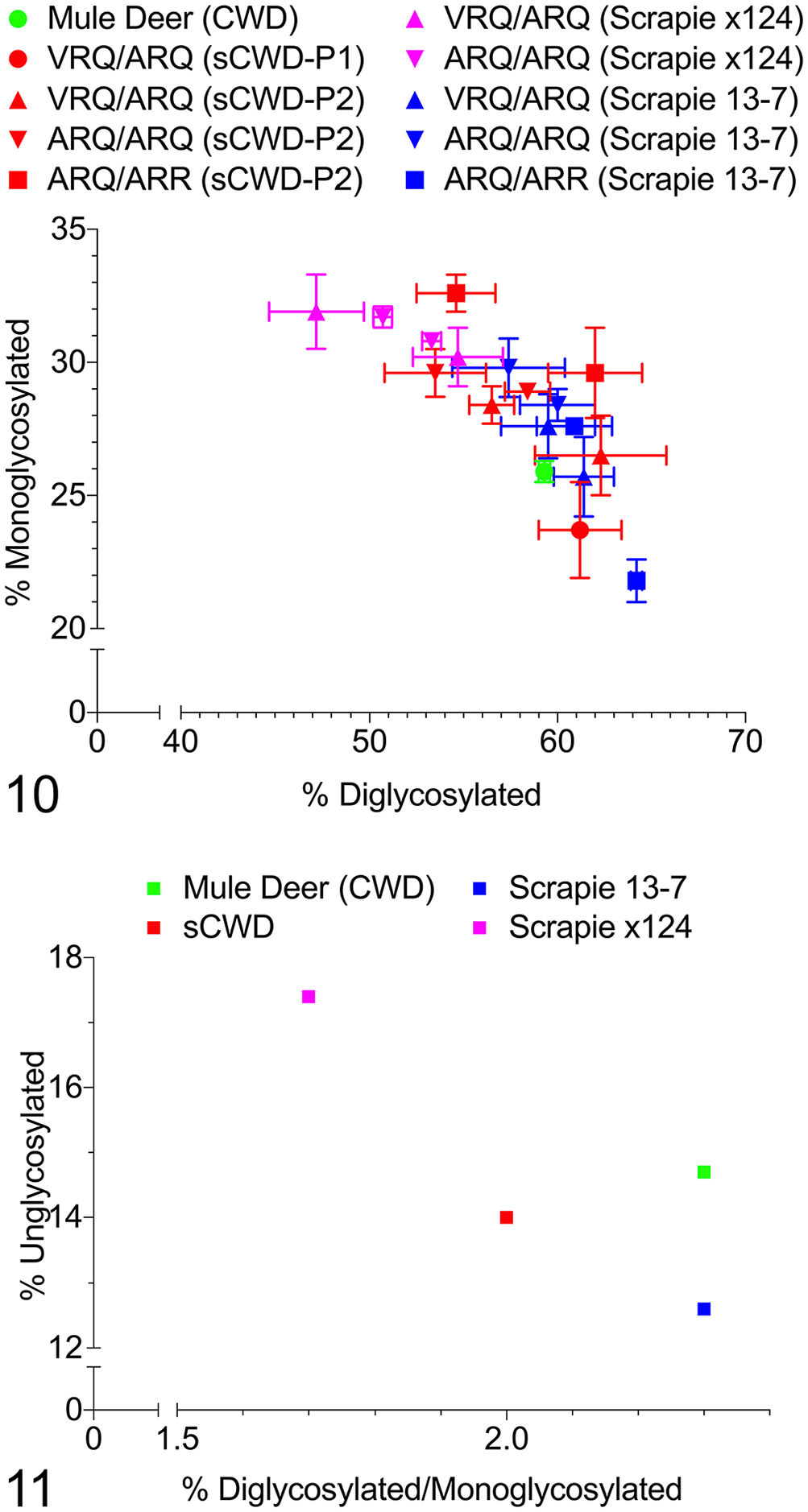
Scatterplots of glycoprofiles.
Discussion
We demonstrated similarities between sheep-passaged CWD and classical scrapie in sheep. Transmission of CWD from white-tailed deer or mule deer has been investigated in several livestock species including cattle, sheep, and swine.17–19 Limited information is available on the transmission of CWD to sheep. Mule deer–derived CWD was initially studied in Suffolk sheep. 18 In that experiment, 1 of 1 VRQ/ARQ sheep developed clinical disease and had detectable PrPSc in the brain and lymphoid tissues after intracranial inoculation; only 1 of 3 ARQ/ARQ sheep and 0 of 4 ARQ/ARR sheep had IHC-positive brain tissue. In our second-passage experiment, there was a higher attack rate and more prevalent lymphoid accumulation. This observation is typical of interspecies transmission studies in which a species barrier is present initially. 9
Brain material from a VRQ/ARQ Suffolk sheep intracranially inoculated with CWDmd was used as second-passage inoculum. After intracranial inoculation, VRQ/ARQ and ARQ/ARQ sheep had widespread lymphoid distribution of PrPSc that was reminiscent of PrPSc distribution in natural scrapie infection.6,14 However, ARQ/ARR sheep did not have detectable lymphoid PrPSc with IHC. The overall absence of lymphoid accumulation in the ARQ/ARR sheep is similar to ARQ/ARR sheep inoculated intracranially with the scrapie agent. 15 In scrapie-infected sheep, the genotype-associated absence of PrPSc in lymphoid tissues of sheep inoculated intracranially corresponds with resistance to oral challenge. 16 Therefore, lymphoid distribution in VRQ/ARQ and ARQ/ARQ sheep but not ARQ/ARR sheep suggests that susceptibility of sheep to CWDmd should be further investigated using the oral route of exposure. It is likely that ARQ/ARR sheep would be resistant to oral inoculation; however, the oral susceptibly of sheep to CWDmd remains untested. Lymphoinvasion is a vital step in the early pathogenic phase of scrapie 40 and CWD in mule deer. 34 The presence of widespread lymphoid PrPSc suggests that some genotypes of sheep may be susceptible to oral infection with CWD.
We observed intergenotypic differences in the disease phenotype. The topographic accumulation of PrPSc in the brains of ARQ/ARR sheep was distinct from VRQ/ARQ and ARQ/ARQ sheep. In the cerebral cortex, sheep with the more resistant ARQ/ARR genotype had extracellular PrPSc predominantly in white matter; conversely, the other genotypes had PrPSc predominantly within gray matter. Inverted gray and white matter PrPSc accumulation occurs after intracranial inoculation in sheep with dissimilar genetic susceptibility characteristics. For example, we previously demonstrated that ARK/ARK sheep had abundant white matter PrPSc after intracranial inoculation that was largely absent in the cerebral cortices of ARQ/ARQ sheep. 6 However, after oral inoculation, ARK/ARK sheep were resistant to the scrapie agent, and ARQ/ARQ sheep were susceptible. Therefore, based on this observed pattern, we would not expect ARQ/ARR sheep to be susceptible to the CWDmd agent via the oral route.
PrPSc was present in all sheep in our study, resulting in a 100% (12 of 12) attack rate; however, the incubation periods were different among the 3 PRNP genotypes represented. ARQ/ARQ sheep had the shortest mean incubation period. This was an unexpected finding for 2 reasons. First, ARQ/ARQ sheep are considered to be less susceptible to scrapie than VRQ/ARQ sheep. 38 Second, the donor inoculum was derived from a VRQ/ARQ sheep, and we expected faster incubations in second-passage sheep of the same genotype.11,26 A possible explanation is that the inoculum contained a preponderance of ARQ allelotype PrPSc instead of VRQ. Another possibility: there could be multiple conformers or sub-strains in the VRQ/ARQ inoculum that template better with ARQ allelotype PrPC. Scrapie isolates have been shown to contain multiple prion strains. 37 Furthermore, the original mule deer inoculum was a pooled homogenate from 28 CWD-affected mule deer, thus multiple conformers likely arose in the first-passage sheep brain that was used as the inoculum for the second passage. Other work has demonstrated that strain properties independent of PrPC amino acid sequence (probably conformational-dependent properties) determine the efficiency of transmission. 39 A conformational-dependent sub-strain in our inoculum may have contributed to superior transmission efficiency in ARQ/ARQ genotype recipients.
The pattern, distribution, and type of PrPSc were similar between scrapie strain 13-7 and sCWDmd; however, this was not observed with scrapie strain x124. Similarly, the molecular characteristics of scrapie strain 13-7 and sCWDmd were indistinguishable with western blot, but scrapie strain x124 had a different glycoprofile (glycoform ratio) than strain 13-7 and sCWDmd. The migration distances were similar for all isolates.
We compared sCWDmd to archived brains of sheep intracranially inoculated with either the x124 or 13-7 U.S. scrapie strain. Scrapie strains x124 and 13-7 were previously shown to have different patterns of immunoreactivity in the brain. 26 We found that 13-7 more closely resembled sCWDmd than x124. Because of the existence of different scrapie strains, it would be difficult to demonstrate experimentally if the scrapie agent jumped species from sheep to deer. Ideally, the researcher would use the foundational scrapie isolate; however, such an isolate is unidentified and not available for characterization. The results from our intracranial inoculation experiment indicate that there is a basis for new work to determine the oral susceptibility of sheep to CWD and compare the PrPSc lesion profiles in orally inoculated sheep.
The molecular profiles of second-passage sCWDmd were indistinguishable from first-passage sCWDmd and the original pooled mule deer inoculum. The first passage of CWDmd in sheep resulted in western blot profiles that were similar but not identical to scrapie. 18 In our study, a larger sample of animals from different TSE groups and genotypes was used to compare molecular profiles. Software measurements and statistical analysis were unable to identify differences in the molecular weight of PrPSc between CWDmd, first-passage sCWDmd, and second-passage sCWDmd. However, we did identify a difference in the molecular weight of unglycosylated PrPSc between scrapie x124 in VRQ/ARQ sheep and both ARQ/ARQ x124 and ARQ/ARR 13-7. There was considerable overlap in the glycoprofiles of classical scrapie and sCWDmd. Statistical differences were not detectable when comparing individual sheep genotypes and TSE inoculum groups. Independent of sheep genotype, there were statistical differences between scrapie x124 and both scrapie 13-7 and sCWDmd. The glycoprofile of TSEs can be used to differentiate strains and TSE types.21,22,24,25,36 We used a single monoclonal antibody P4 to label PrPSc, and sCWDmd could not be distinguished from scrapie strain 13-7. Future work will be aimed at exploring differential binding of a panel of anti-PrPSc antibodies and proteinase K sensitivity to examine potential molecular differences.
The comparison of PrPSc lesion profiles has been used previously to distinguish scrapie strains in sheep in the context of concurrent differences between incubation periods and clinical signs.13,26 Comparison of sheep inoculated orally with scrapie and CWD would be ideal for comparing PrPSc profiles. Positive tissues from sheep inoculated orally with the CWD agent are not available. We examined intracranially inoculated sheep. This precluded comparison with orally infected scrapie field isolates because the experimental inoculation route affects the lesion profile. 26
To date, there is no evidence that sheep are susceptible to CWD under natural field conditions; however, given our findings, it would be difficult to discriminate the phenotype of CWD in sheep from scrapie using routine detection methods. Epidemiologic assessment of CWD suggests that it first appeared in captive mule deer before spreading to elk and white-tailed deer, but the origin of CWD is unknown. 43 It has been hypothesized that CWD originated from scrapie; that is supported by intermediate in vitro conversion efficiencies of sheep PrPC induced by CWD prions. 32 Here we demonstrated strong phenotypic similarities between sCWDmd and scrapie strain 13-7. Another question to consider in future studies is whether various CWD strains from white-tailed deer, mule deer, and elk have distinct phenotypes in sheep.
We demonstrated that sheep inoculated with sheep-adapted CWDmd amplify and distribute PrPSc with abundant lymphoid involvement similar to sheep with scrapie. Intracranial inoculation can overcome natural species barriers. Oral susceptibility would be unlikely in some genotypes of sheep; however, the oral susceptibility of sheep to CWD has not been investigated. Such research will help determine if sheep are susceptible to CWD under natural conditions. Furthermore, studies involving mouse models will further characterize similarities between sheep scrapie and sheep CWDmd in hopes of shedding light on the origins of CWD.
Footnotes
Acknowledgements
We thank Martha Church, Kevin Hassall, Joe Lesan, Leisa Mandell, and Trudy Tatum for providing technical support to our project. We also recognize the contributions of Amir Hamir (decd.) towards the original conception and design of our research. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA. The USDA is an equal-opportunity provider and employer.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our research was funded in its entirety by congressionally appropriated funds to the USDA, Agricultural Research Service (ARS). The funders of the work did not influence study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Our research was supported in part by an appointment to the ARS Research Participation Program administered by the Oak Ridge Institute for Science and Education (ORISE) through an interagency agreement between the U.S. Department of Energy (DOE) and USDA. ORISE is managed by Oak Ridge Associated Universities (ORAU) under DOE contract DE-SC0014664. All opinions expressed in this paper are the authors’ and do not necessarily reflect the policies and views of USDA, ARS, DOE, or ORAU/ORISE.
