Abstract
Equine infectious anemia (EIA) is a highly infectious disease in members of the Equidae family, caused by equine infectious anemia virus (EIAV). The disease severity ranges from subclinical to acute or chronic, and causes significant economic losses in the equine industry worldwide. Serologic tests for detection of EIAV infection have some concerns given the prolonged seroconversion time. Therefore, molecular methods are needed to improve surveillance programs for this disease. We attempted detection of EIAV in 6 clinical and 42 non-clinical horses in Nuevo Leon State, Mexico, using the agar gel immunodiffusion (AGID) test for antibody detection, and nested and hemi-nested PCR for detection of proviral DNA. We found that 6 of 6, 5 of 6, and 6 of 6 clinical horses were positive by AGID, nested PCR, and hemi-nested PCR, respectively, whereas 0 of 42, 1 of 42, and 9 of 42 non-clinical horses were positive by these tests, respectively. BLAST analysis of the 203-bp 5′-LTR/tat segment of PCR product revealed 83–93% identity with EIAV isolates in GenBank and reference strains from other countries. By phylogenetic analysis, our Mexican samples were grouped in a different clade than other sequences reported worldwide, indicating that the LRT/tat region represents an important target for the detection of non-clinical horses.
Equine infectious anemia (EIA), a disease caused by the macrophage-tropic lentivirus equine infectious anemia virus (EIAV; Retroviridae, Lentivus), affects all members of the Equidae family.1,5,15 EIA is a worldwide animal health problem associated with significant economic losses given its broad distribution and chronic untreatable form of disease. 8 EIA involves periods of anemia, fever, leukopenia, thrombocytopenia, and weight loss.4,9,11,21 EIAV is mechanically transmitted from blood of infected horses to other horses by contaminated syringe needles and by blood-feeding insect vectors (Tabanus spp., Hybomitra spp., and Stomoxys calcitrans).5,13 It is also transmitted in utero or through body secretions (e.g., semen, sweat, urine). 20 Recognizing the global distribution and presence of EIAV in clinical horses, timely diagnosis and elimination of infected individuals are essential in the prevention and control of transmission within the equine population.
The high variation among individual cases, together with the lack of specific disease characteristics, impairs EIA diagnosis based on clinical signs or clinicopathologic findings.2,18 Diagnosis typically depends on serologic detection techniques of limited sensitivity, such as the agar gel immunodiffusion (AGID) test, which produces false-negative results given the seroconversion time that may take up to 157 d. 7 In addition, the scarce information about this disease in Mexico, as well as the economic losses resulting from the elimination of infected horses and the risk of transporting non-clinical individuals inside and outside the country without early diagnosis, prompted us to investigate the prevalence of EIAV in Nuevo Leon, Mexico, by nested and hemi-nested PCR testing.
Sera from 42 non-clinical horses (23 females and 19 males) were obtained from a ranch in the state of Nuevo Leon, during an EIA outbreak. Six clinical horses (4 females and 2 males) from another ranch, served as positive horse samples. Clinical horses were confirmed by an AGID test (VMRD), which was performed at the Centro Nacional de Servicios de Diagnóstico en Salud Animal (CENASA) in Tecamac, Estado de México.
Sera were centrifuged (5,000 × g for 10 min), after which supernatants were refrigerated and submitted to CENASA for processing. For molecular detection of EIAV in clinical and non-clinical horse sera, peripheral blood mononuclear cells (PBMCs) were obtained by gradient centrifugation (Ficoll-Hypaque; GE Healthcare), and genomic DNA was extracted (blood DNA preparation kit; Jena Bioscience), following the supplier’s instructions. For nested PCR, extracted DNA was subjected to a PCR reaction, using 200 nM of the first set of primers [EIAVltr-1F (5′-GACAGTTGGGCACTCAGATT-3′) and EIAVltr-1R (5′-CAGGAACACCTCCAGAAGAC-3′)], 400 ng of genomic DNA, and a Ruby Taq master mix kit (Jena Bioscience) in a volume of 20 µL. Conditions consisted of denaturation at 95°C for 2 min, followed by 30 cycles of 95°C for 20 s, 50°C for 30 s, 72°C for 30 s, and final extension at 72°C for 5 min. The product of the first reaction (2 µL) of all samples was subjected to a second PCR, using 200 nmol of the second set of primers [EIAVltr-2F (5′-ATTCTGCGGTCTGAGTCCCT-3′) and EIAVltr-2R (5′-TAAGTTCTCCTCTGCTGTCC-3′) 9 ] and a Ruby Taq master mix kit (Jena Bioscience) in a volume of 20 µL. Conditions consisted of denaturation at 95°C for 2 min, followed by 30 cycles of 95°C for 20 s, 52°C for 30 s, 72°C for 30 s, and final extension at 72°C for 5 min. The product of the second reaction was subjected to 1.5% agarose gel electrophoresis for 50 min at 100 V, after which it was stained with ethidium bromide, expecting an amplicon of 198–203 bp, depending on the EIAV strain. The identity of 2 amplicons, 1 each from clinical and non-clinical horse serum samples, was then confirmed by nucleic acid sequencing. 9
For the hemi-nested PCR, extracted DNA was subjected to a PCR reaction, using 200 nmol of the first set of primers [EIAVltr-4F (5′-GTCCCTTCTCTGCTGGGCTA-3′) 7 and EIAVltr-1R], 400 ng of genomic DNA, and the Ruby Taq master mix kit (Jena Bioscience) in a volume of 20 µL. Conditions consisted of denaturation at 95°C for 2 min, followed by 30 cycles of 95°C for 20 s, 58°C for 30 s, 72°C for 30 s, and final extension at 72°C for 5 min. The product of the first reaction (2 µL) of all samples was subjected to a second PCR using 200 nmol of the second set of primers (EIAVltr-4F and EIAVltr-2R) and the Ruby Taq master mix kit (Jena Bioscience) in a volume of 20 µL. Conditions consisted of denaturation at 95°C for 2 min, followed by 30 cycles of 95°C for 20 s, 60°C for 30 s, 72°C for 30 s, and final extension at 72°C for 5 min. The product of the second reaction was subjected to 1.5% agarose gel electrophoresis for 50 min at 100 V, after which it was stained with ethidium bromide, expecting an amplicon of 185 bp, depending on the EIAV strain. The identity of 2 amplicons, 1 each from clinical and non-clinical horse serum samples, was then confirmed by nucleic acid sequencing. 7
All clinical horses were positive on AGID testing, and all of the non-clinical horses were negative; 5 of 6 clinical horses were positive on nested PCR, and all 6 clinical horses were positive on hemi-nested PCR. Only 1 non-clinical horse was positive on nested PCR, which was also positive by hemi-nested PCR, and 9 of 42 non-clinical horses were positive on hemi-nested PCR (Table 1). Hence, the hemi-nested PCR was more efficient than the nested PCR for EIAV detection, as reported previously. 7
Serologic and molecular detection of equine infectious anemia virus in clinical and non-clinical horses.

From the PCR products obtained (5 from clinical and 1 from non-clinical samples by nested PCR, and 6 from clinical and 9 from non-clinical samples by hemi-nested PCR), we randomly selected 4 (1 each from clinical and non-clinical horse serum samples from nested and hemi-nested PCR) and purified them using an agarose gel extraction kit (Jena Bioscience). The resulting fragment was sequenced in an automated capillary sequencer (Applied Biosystems) by the Laboratorio de Servicios de Genómica of Laboratorio Nacional de Genómica para la Biodiversidad at CINVESTAV-IPN, México. Long terminal repeat (LTR)/tat segment sequence analysis was performed using MEGA X software with 1,000 bootstrapped data by the neighbor-joining method. 14 PCR product sequences were submitted to GenBank, with accessions pending.
BLAST analysis of the 203-bp 5′-LTR/tat segment from the PCR products of the 4 clinical and non-clinical horses revealed 85–93% identity with EIAV isolates in GenBank. Furthermore, the phylogenetic analysis of this segment, using the sequences from one each of clinical and non-clinical samples by hemi-nested PCR (the most specific method, which identified most samples with EIAV), showed an identity of 84–90% with EIAV isolates reported in Bosnia and Herzegovina, China, England, Hungary, Mongolia, Ireland, Israel, Italy, Japan, and UAE, as well as Liaoning and Miyazaki reference strains. 3 This analysis grouped sequences in a different clade compared with other isolates reported worldwide (Fig. 1), 7 demonstrating that the LRT/tat region represents an important target for the preventive detection of non-clinical horses. 6
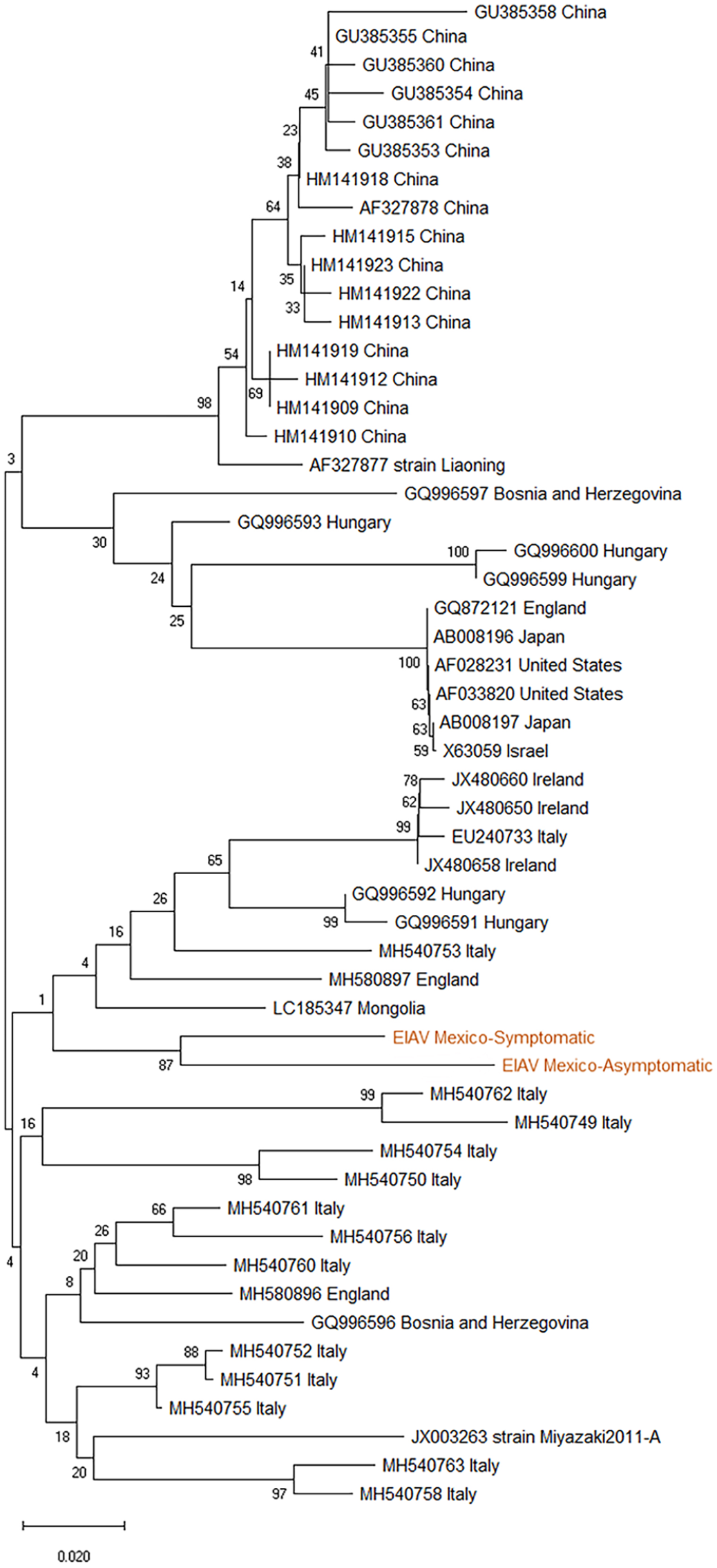
Phylogenetic tree constructed from equine infectious anemia virus (EIAV) LTR/tat nucleotide sequences, using the neighbor-joining method with bootstrap analysis of 1,000 replicates.
As expected, the AGID test was positive only in samples from clinical horses.10,12,16 However, we found EIAV-positive samples from PBMC DNA in both clinical and non-clinical equine samples. Geographic regions have different circulating strains and, given the great variability in EIAV sequences as shown in our results, regionally developed detection techniques should be modified according to the genetic or antigenic variability of the strains that are in circulation.17,19,22 We found that nested and hemi-nested PCR assays were more effective than AGID testing for identification of EIAV in non-clinical horses in the early stages of the disease.
Footnotes
Acknowledgements
We thank Laboratorio de Inmunología y Virología of Facultad de Ciencias Biológicas at Universidad Autónoma de Nuevo León (FCB-UANL) for supporting the development of this study.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by a grant from Programa de Apoyo a la Investigación Científica y Tecnológica-UANL 2019 (grant CN-933) to Ricardo Gomez-Flores and by internal research support of Universidad de Sonora grant USO513006999 to Guadalupe González-Ochoa.
