Abstract
Prudent use of antimicrobials is paramount to slow the development of resistance and for successful treatment. The use of cumulative antibiograms will allow evidence-based antimicrobial selection with consideration of local resistance patterns. We generated a “first-isolate-per-patient” cumulative antibiogram for a regional equine referral hospital. Bacterial organisms cultured from horses between 2011 and 2018, sample origin, antimicrobial susceptibilities, and multidrug-resistant (MDR) status were tabulated. Of 1,176 samples, 50% were culture positive. Overall, 93 of 374 (25%) were MDR. Of the MDR isolates, 11 (12%) were susceptible to high-importance antimicrobials only (as defined by the Australian Strategic and Technical Advisory Group on antimicrobial resistance). β-hemolytic streptococci were uniformly susceptible to penicillin (76 of 76); 17 of 20 (85%) non–β-hemolytic Streptococcus spp. were susceptible to penicillin. Despite veterinary-specific challenges in constructing an antibiogram, our study provides an exemplar of the clinical utility of regional-, farm-, or hospital-specific cumulative antibiograms for evidence-based empirical antimicrobial selection by veterinarians prior to susceptibility result availability.
The development of antimicrobial resistance is a matter of increasing concern in the One Health approach to medicine. 19 Unresolved bacterial infections within wounds, joints, lungs, and other organs can be career-ending and life-threatening for horses.14,15 Prudent use of antimicrobials is important to slow the development of resistance and to increase the likelihood of treatment success. A cumulative antibiogram is a report of microorganism susceptibility to commonly used antimicrobials within a specific time frame (commonly 1 y) at a particular institution, typically for a single species. 3 Antibiograms are used in human hospitals to monitor antimicrobial resistance trends, guide empirical antimicrobial selection prior to the return of culture and susceptibility results, and within antimicrobial stewardship programs. 3 Susceptibility studies are ideally region- or hospital-specific, and there are few susceptibility studies available for equine patients to guide antimicrobial selection.14,16,17 Although multiple calculation methods exist and there are no veterinary-specific guidelines, the “first-isolate-per-patient” strategy appears appropriate for generation of a veterinary hospital antibiogram. 3
Although various studies describe changes in antimicrobial susceptibility patterns in bacterial isolates from horses over time,9,14,16 no veterinary antibiograms appear to have been reported. We aimed to generate an equine cumulative antibiogram for a veterinary referral hospital using a “first-isolate-per-patient” strategy. Antimicrobials were classified as low, medium, and high importance as defined by the Australian Strategic and Technical Advisory Group on antimicrobial resistance (ASTAG). 1 Although cumulative antibiograms are hospital specific, the one presented in our study could serve as a model for veterinary institutions to develop species-specific cumulative antibiograms.
We extracted bacteriologic submissions from an equine hospital between January 2011 and June 2018. All cases had been referred by primary care veterinarians. In vitro antimicrobial susceptibility testing was performed using the disk diffusion method and reported as susceptible, intermediate, or resistant depending on zone of inhibition diameters according to Clinical Laboratory Standards Institute (CLSI) guidelines available at the time of reporting. 4 Although veterinary-specific breakpoints were used preferentially, we used human guidelines 11 if no veterinary breakpoints were available.
Only the first isolate of a bacterial species per patient per year was included in the cumulative antibiogram regardless of the site of sample collection, susceptibility profile, and time interval, according to the CLSI “first-isolate-per-patient” method. 3 We excluded from our study all research, screening, and surveillance cultures, samples with organisms not cultured but identified by direct PCR on the clinical material, and samples from which Streptococcus spp. were cultured but penicillin susceptibility was not available or it was not possible to confirm a resistant phenotype.
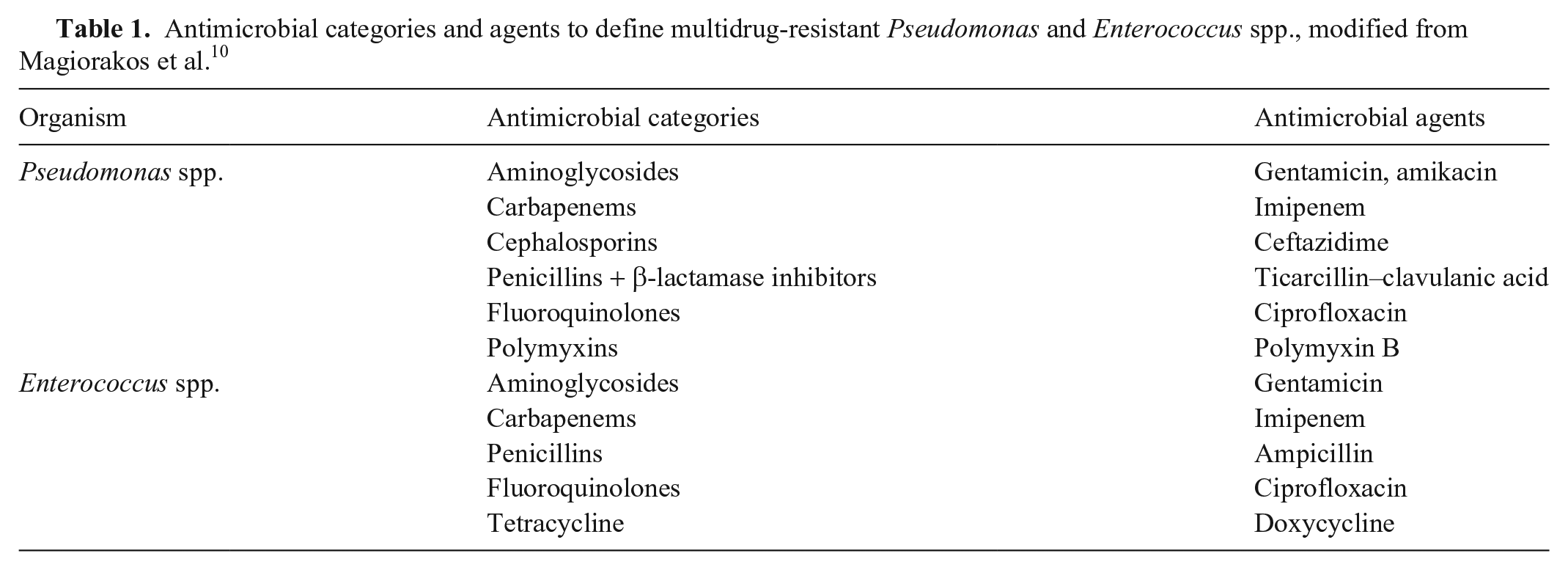
Organisms cultured, antimicrobial susceptibilities, and signalment data were tabulated and categorized by sample site of origin: respiratory, wounds, synovial, reproductive, fecal, abdominal, bone, ocular, and other (skin, urine, blood, subcutis, cerebrospinal fluid, muscle, liver, or unclassified). Antimicrobials tested included: amoxicillin–clavulanic acid, cephalothin and/or cephalexin, ampicillin and/or amoxicillin, penicillin, ceftazidime, ticarcillin–clavulanic acid, ceftiofur, methicillin, oxacillin, gentamicin, amikacin, neomycin, erythromycin, clindamycin and/or lincomycin, chloramphenicol, tetracycline, doxycycline, trimethoprim–sulfamethoxazole (TMS), enrofloxacin, ciprofloxacin, rifampicin, and polymyxin B. However, not all antimicrobials were tested against each isolate. A cumulative antibiogram was generated for the 6 most commonly cultured organisms. Antimicrobials to which a bacterium had intrinsic resistance, according to CLSI guidelines, were omitted from analysis and presented as “R.” 5 The MDR status was defined as resistance to at least 1 agent in 3 or more antimicrobial categories, excluding intrinsic resistance. 10 The MDR status of intrinsically resistant organisms (i.e., Pseudomonas spp. and Enterococcus spp.) was classified as described previously 10 (Table 1). Streptococcus spp. were classified into non–β-hemolytic and β-hemolytic (S. equi subsp. equi or zooepidemicus, S. dysgalactiae subsp. equisimilis, and unclassified). 7 Antimicrobial susceptibility for unclassified gram-positive organisms was evaluated as per Staphylococcus spp., and for unclassified gram-negative organisms as per Enterobacteriaceae. Percentage susceptibility to each antimicrobial for the 6 most common bacterial organisms and the proportion of MDR organisms were calculated. Following CLSI guidelines, Agresti–Coull 95% confidence intervals were calculated for organisms with < 30 isolates. 3
Antimicrobial categories and agents to define multidrug-resistant Pseudomonas and Enterococcus spp., modified from Magiorakos et al. 10

Proportions of organisms susceptible to gentamicin and amikacin, and tetracycline and doxycycline, were compared for the 6 most commonly cultured organisms. Temporal analysis was performed for the 3 most commonly cultured organisms. The database was divided into 2 groups: early (2011–2014) and late (2015–2018); susceptibility results were categorized into resistant and non-resistant. Categorical variables were compared using a chi-square test or a Fisher exact test, and statistical significance (p ≤ 0.05) determined.
In total 1,176 culture and susceptibility reports were included; 565 (49.7%) were culture positive, with susceptibility reports available for 409 (69.9%), after excluding 76 duplicate samples. Most samples were from live patients; 14 isolates were recovered from postmortem samples. Population demographics consisted of 197 female and 214 male horses, 74 of which were foals (< 1 mo). The median age was 5 y (range: 1 d to 30 y). Thoroughbreds were the most common breed (45.7%), followed by Quarter Horses (13.0%) and Standardbreds (9.0%). Overall, the most commonly isolated organisms were Streptococcus spp. (116 of 565; 20.5%), followed by Staphylococcus spp. (81 of 565; 14.3%), Escherichia coli (69 of 565; 12.2%), Pseudomonas spp. (51 of 565; 9.0%), and Actinobacillus spp. (34 of 565; 6.0%; Table 2).
Bacterial organisms isolated from equine samples between 2011 and 2018.
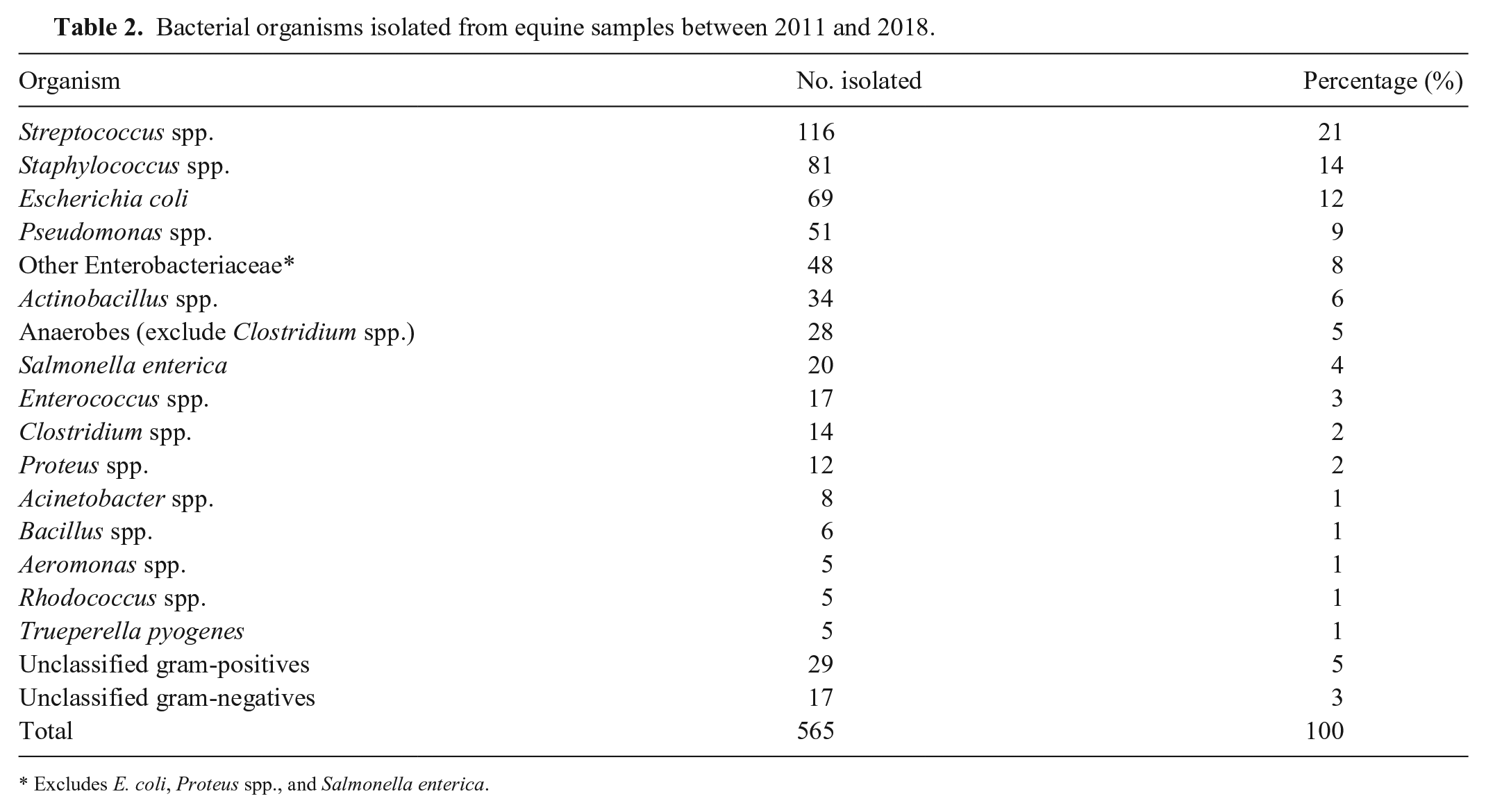
Excludes E. coli, Proteus spp., and Salmonella enterica.
Susceptibility testing was performed on 409 bacterial isolates, of which 374 reports met the criteria of the “first-isolate/patient/year” algorithm. Of these, 192 of 374 (51.3%) were gram-positive and 182 of 374 (48.7%) were gram-negative. Overall, 93 of 374 (24.8%) were MDR, with 42 of 93 (45%) gram-positive and 51 of 93 (55%) gram-negative isolates. Based on morphometric cytologic appearance, 148 of 155 (95.5%) of bacilli were gram-negative, 183 of 183 (100%) of cocci were gram-positive, and 28 of 36 (78%) of coccobacilli were Actinobacillus spp. A higher antimicrobial resistance rate was identified in gram-negative compared to gram-positive organisms (Tables 3–5). Temporal analysis showed no significant differences (p > 0.05) in susceptibility to routinely reported antimicrobials between the early and late time periods for Streptococcus spp., Staphylococcus spp., or E. coli.
Proportion of gram-positive versus gram-negative organisms susceptible to antimicrobials (number shown as % susceptible).
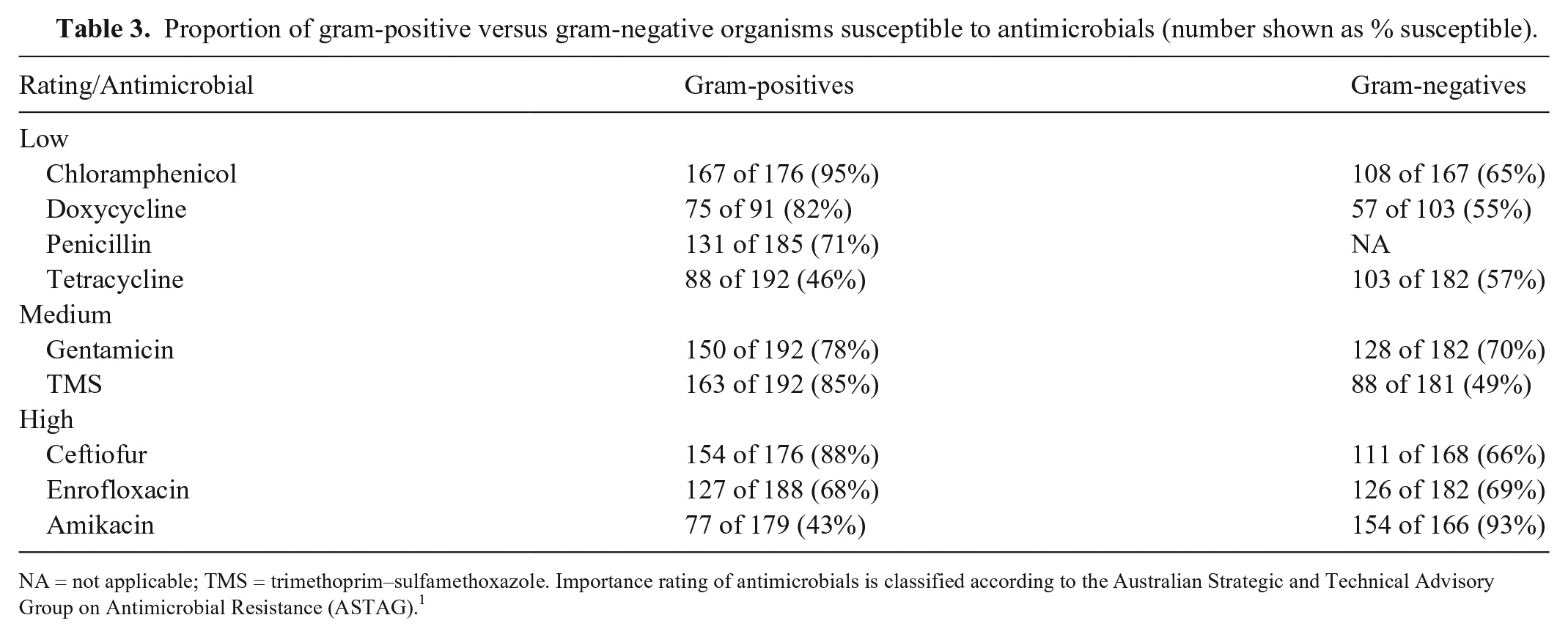

NA = not applicable; TMS = trimethoprim–sulfamethoxazole. Importance rating of antimicrobials is classified according to the Australian Strategic and Technical Advisory Group on Antimicrobial Resistance (ASTAG). 1
Cumulative antibiogram, with low- and medium-importance antimicrobials for clinical use (numbers shown are % sensitive). 95% confidence intervals (CI) are included for antimicrobials tested with < 30 isolates.
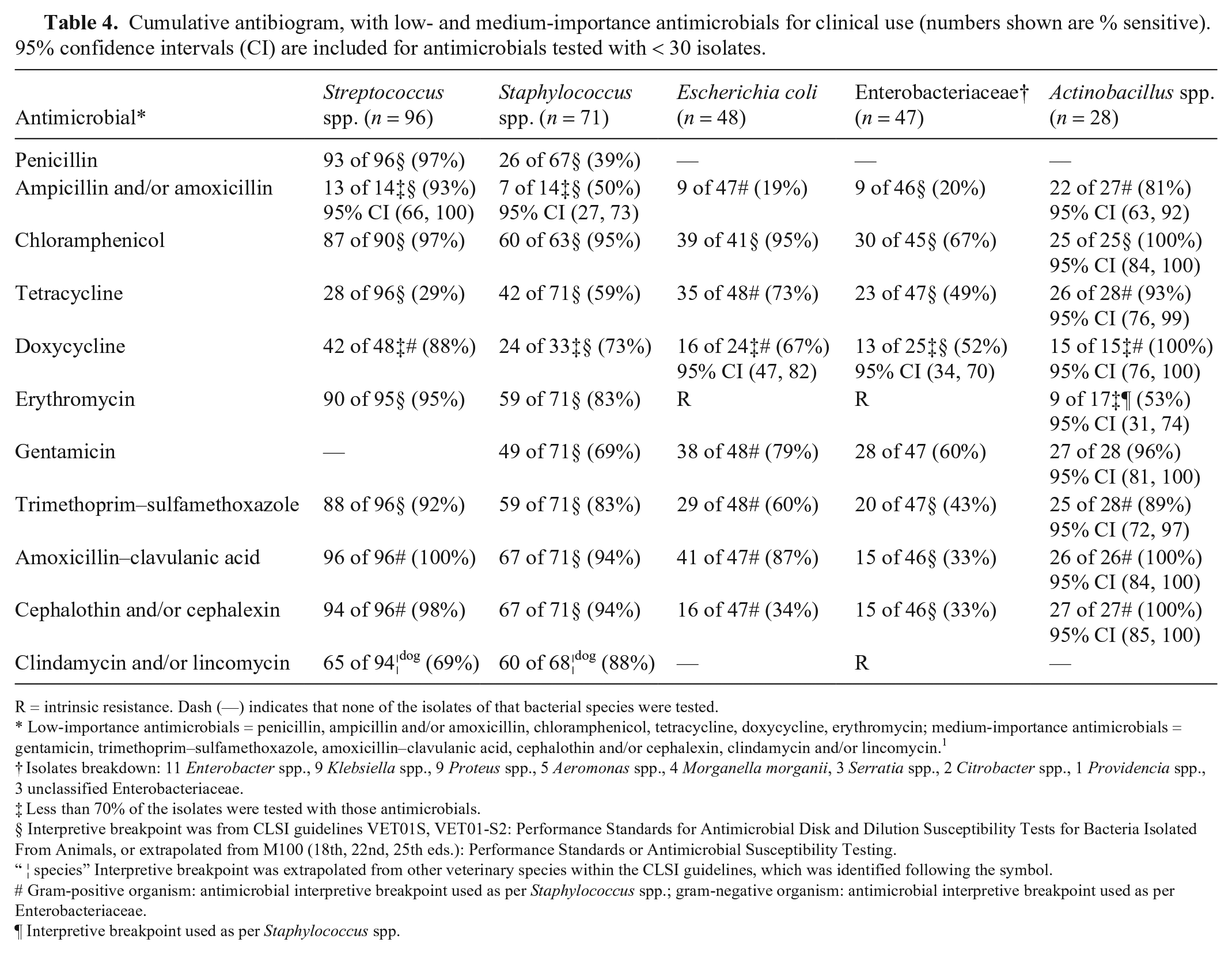
R = intrinsic resistance. Dash (—) indicates that none of the isolates of that bacterial species were tested.
Low-importance antimicrobials = penicillin, ampicillin and/or amoxicillin, chloramphenicol, tetracycline, doxycycline, erythromycin; medium-importance antimicrobials = gentamicin, trimethoprim–sulfamethoxazole, amoxicillin–clavulanic acid, cephalothin and/or cephalexin, clindamycin and/or lincomycin. 1
Isolates breakdown: 11 Enterobacter spp., 9 Klebsiella spp., 9 Proteus spp., 5 Aeromonas spp., 4 Morganella morganii, 3 Serratia spp., 2 Citrobacter spp., 1 Providencia spp., 3 unclassified Enterobacteriaceae.
Less than 70% of the isolates were tested with those antimicrobials.
Interpretive breakpoint was from CLSI guidelines VET01S, VET01-S2: Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals, or extrapolated from M100 (18th, 22nd, 25th eds.): Performance Standards or Antimicrobial Susceptibility Testing.
“ ¦ species” Interpretive breakpoint was extrapolated from other veterinary species within the CLSI guidelines, which was identified following the symbol.
Gram-positive organism: antimicrobial interpretive breakpoint used as per Staphylococcus spp.; gram-negative organism: antimicrobial interpretive breakpoint used as per Enterobacteriaceae.
Interpretive breakpoint used as per Staphylococcus spp.
Cumulative antibiogram, with-high importance antimicrobials for clinical use (numbers shown are % sensitive). 95% confidence intervals (CI) are included for antimicrobials tested with <30 isolates.
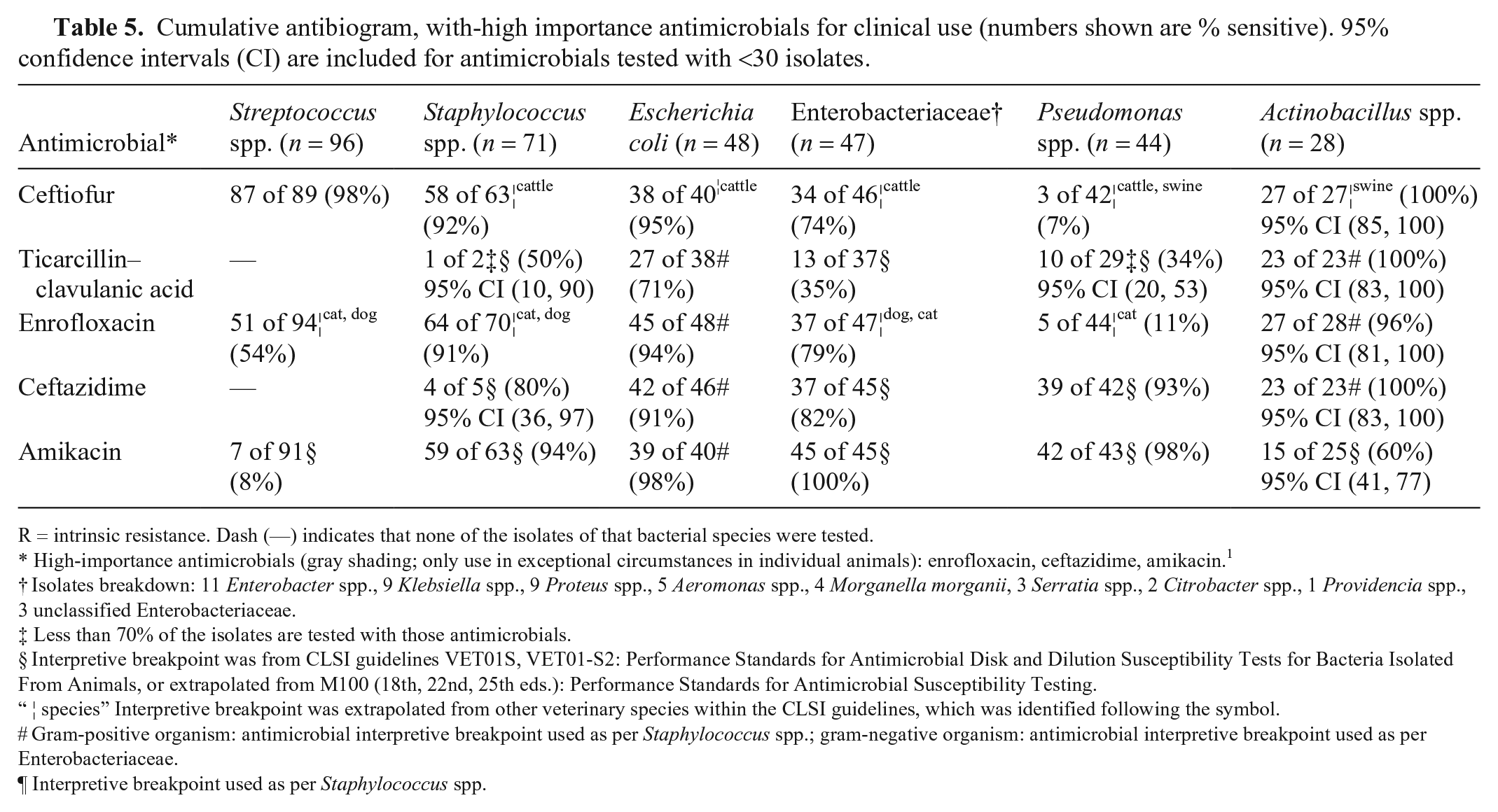
R = intrinsic resistance. Dash (—) indicates that none of the isolates of that bacterial species were tested.
High-importance antimicrobials (gray shading; only use in exceptional circumstances in individual animals): enrofloxacin, ceftazidime, amikacin. 1
Isolates breakdown: 11 Enterobacter spp., 9 Klebsiella spp., 9 Proteus spp., 5 Aeromonas spp., 4 Morganella morganii, 3 Serratia spp., 2 Citrobacter spp., 1 Providencia spp., 3 unclassified Enterobacteriaceae.
Less than 70% of the isolates are tested with those antimicrobials.
Interpretive breakpoint was from CLSI guidelines VET01S, VET01-S2: Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals, or extrapolated from M100 (18th, 22nd, 25th eds.): Performance Standards for Antimicrobial Susceptibility Testing.
“ ¦ species” Interpretive breakpoint was extrapolated from other veterinary species within the CLSI guidelines, which was identified following the symbol.
Gram-positive organism: antimicrobial interpretive breakpoint used as per Staphylococcus spp.; gram-negative organism: antimicrobial interpretive breakpoint used as per Enterobacteriaceae.
Interpretive breakpoint used as per Staphylococcus spp.
Of all MDR pathogens (Table 6), 11 (12%) were susceptible to high-importance antimicrobials only: 10 of 10 (100%) were susceptible to amikacin, 6 of 10 (60%) to ceftazidime, 5 of 11 (46%) to ceftiofur, and 1 of 10 (10%) to rifampicin. All MDR S. equi subsp. zooepidemicus (7 of 7; 100%) were resistant to amikacin; however, only 1 of 7 (14%) was resistant to gentamicin. All 7 (100%) were resistant to tetracycline but only 1 of 3 (33%) was resistant to doxycycline, and 4 of 7 (57%) to clindamycin and/or lincosamide. Nevertheless, 7 of 7 (100%) were susceptible to penicillin.
Proportion of multidrug-resistant (MDR) organisms.
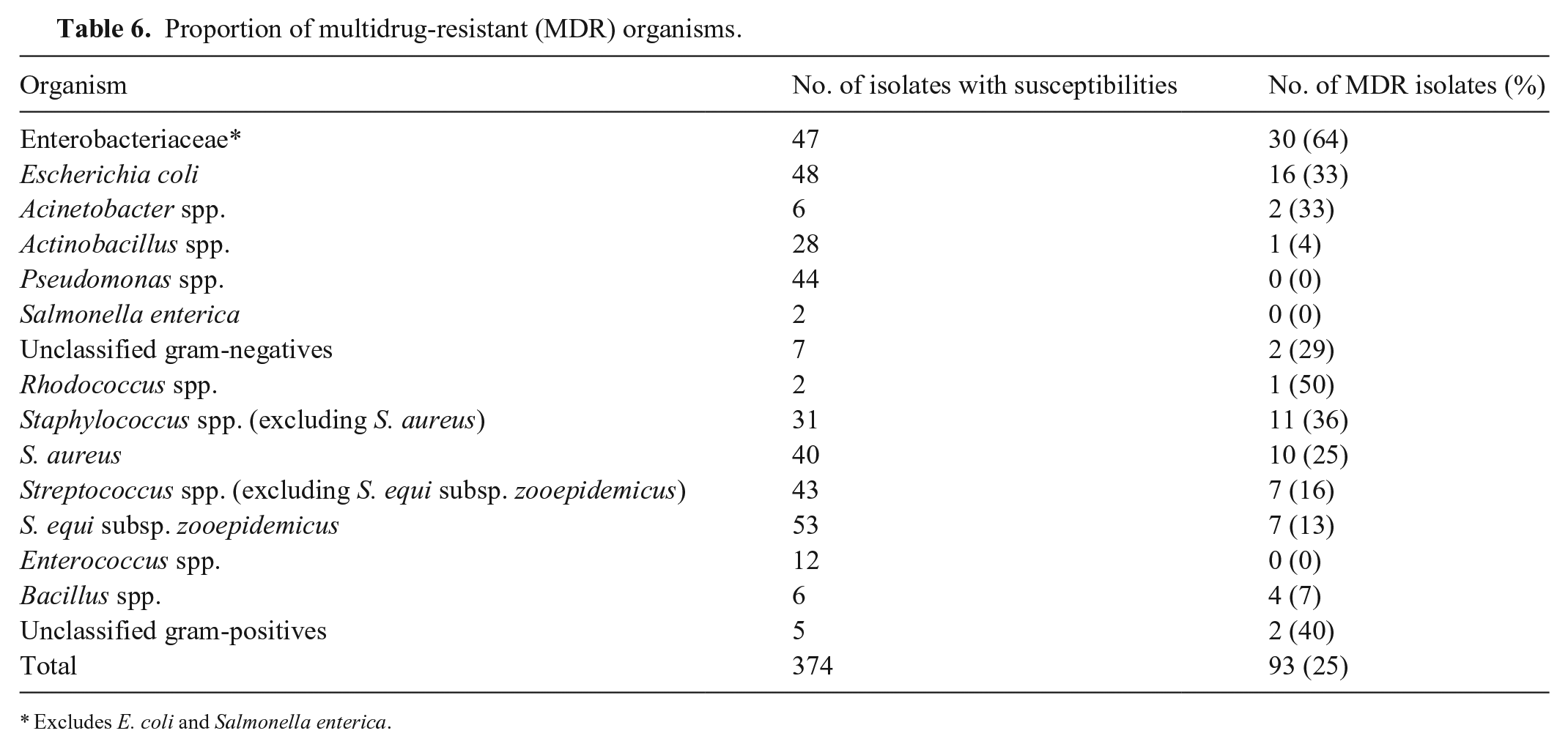

Excludes E. coli and Salmonella enterica.
Of 21 MDR Staphylococcus spp., 10 were S. aureus and all were susceptible to at least 1 β-lactam antimicrobial. However, 20 of 20 (100%) were resistant to penicillin, 19 of 21 (91%) to gentamicin, and 17 of 21 (81%) to tetracycline. Only 3 of 19 (16%) were resistant to amikacin and 1 of 12 (8%) to doxycycline.
Of the MDR E. coli, the resistance level to ampicillin and/or amoxicillin was 14 of 15 (93%), to TMS 14 of 16 (88%), doxycycline 7 of 11 (64%), tetracycline 12 of 16 (75%), and gentamicin 9 of 16 (56%); only 1 of 14 (7%) of the isolates was resistant to amikacin.
The respiratory system had the most isolates cultured (124 of 565), with an 83.2% culture-positive rate. Streptococcus equi subsp. zooepidemicus was the most commonly cultured organism (27 of 124), followed by Pseudomonas spp. (22 of 124) and other Streptococcus spp. (18 of 124). Previous studies have also reported that β-hemolytic streptococci (especially S. zooepidemicus) are more commonly identified than non–β-hemolytic streptococci.2,17 To date, there is no report of penicillin-resistant β-hemolytic streptococci in the equine literature, to our knowledge. According to our cumulative antibiogram, all β-hemolytic, and most (17 of 20) non–β-hemolytic, streptococci were susceptible to penicillin. Therefore, penicillin remains the most effective antimicrobial in vitro and is preferred given that it is a low-importance antimicrobial.
Wounds were the second most common site of origin (71 of 565; 12.5%), with an 88.8% culture-positive rate, from which the most frequently cultured organisms were E. coli (12 of 71; 17%) and S. aureus (11 of 71; 16%). Using the cumulative antibiogram generated, topical chloramphenicol could be considered. If systemic infection is suspected, intravenous gentamicin should be initiated. Alternatively, oral chloramphenicol (if not prohibited depending on regional legislative controls) could be recommended. Although penicillin is a low-importance antimicrobial, it may often be ineffective against S. aureus and E. coli (acquired and intrinsic resistance, respectively).
Synovial fluid accounted for 56 of 565 (9.9%) of all cultures, with a 29.3% culture-positive rate. Actinobacillus spp. were the most frequently cultured organisms (8 of 56; 14%) followed by Staphylococcus spp. (7 of 56; 13%), Streptococcus spp. (5 of 56; 9%), and E. coli (5 of 56; 9%). Although our cumulative antibiogram suggests that penicillin and amikacin would provide the most appropriate empirical choice for synovial infections, systemic amikacin is cost-prohibitive in most adult horses. Administration of amikacin by intra-articular, intra-osseous, or regional limb perfusion is recommended for synovial infections. 18 Given that amikacin is classified as a high-importance antimicrobial,1,20 culture and susceptibility testing should be performed before administration to comply with antimicrobial stewardship standards. Alternatively, gentamicin (a medium-importance antimicrobial) could be used while awaiting culture and susceptibility results.
Bacterial organisms that are resistant to one antimicrobial are likely to be resistant to other antimicrobials of the same class. 21 Interestingly, Streptococcus and Actinobacillus spp. were more susceptible to gentamicin than amikacin (p < 0.0001; p = 0.0015), whereas staphylococci, E. coli, and Pseudomonas spp. were more susceptible to amikacin than gentamicin (p = 0.0003; p = 0.0102; p < 0.0001, respectively). Enterobacteriaceae had a moderately higher susceptibility to amikacin than gentamicin (p < 0.0001), which might be the result of differences in resistance to aminoglycoside-modifying enzymes, such as acetyltransferase(6’). 13 However, the high proportion of resistance to gentamicin identified in Enterobacteriaceae and Pseudomonas spp. is concerning because the use of amikacin is expensive and is considered an antimicrobial of high importance by ASTAG 1 and the World Health Organization. 20
Penicillin, gentamicin, and TMS are some of the most frequently used antimicrobials in equine practice worldwide. In our study, of all organisms tested against penicillin and/or gentamicin, 131 of 187 (70.1%) were susceptible to penicillin, 278 of 374 (74.3%) were susceptible to gentamicin, and 336 of 374 (89.8%) were susceptible to either penicillin or gentamicin. Based on their mechanism of action and synergy in vivo, the clinical benefit of the broad-spectrum antimicrobial combination of penicillin and gentamicin is likely to be even higher. 8 Penicillin should be used to treat streptococcal infections; TMS provided moderate effectiveness against Staphylococcus spp. and Actinobacillus spp. However, TMS is inactivated in purulent environments. 6
There are a few limitations to our study. First, an epidemiologic bias exists. Given that our institution serves as a referral-only practice, many horses would have been treated with various antimicrobials before hospital admission. Another limitation is that a cumulative antibiogram is species-, farm-, hospital-, or clinic-specific. Third, the interpretive breakpoints used were based on the time of reporting; therefore, temporal changes of resistance may not be directly comparable for antimicrobials for which breakpoints changed over the years. Thus, there is a distinct advantage for use of minimum inhibitory concentrations (MIC) because the MIC value can be reinterpreted should breakpoints have changed over time. In addition, given the retrospective nature of our study, data analysis was limited to prior entries into microbiologic reports and hospital records.
There are veterinary-specific challenges in the construction of a cumulative antibiogram. Specifically, the low number of culture and susceptibility test submissions would prevent yearly antibiogram updates as recommended. 3 Bacterial culture and susceptibility testing are often more commonly performed in referral hospitals. On average, only 50–100 organisms (bacterial or fungal) were isolated from horses each year at our institution, making it almost impossible to generate a cumulative antibiogram annually. However, this can be solved by combining data for several years. 3
Antimicrobial resistance patterns vary geographically. 12 Generation of regional, species-specific, cumulative antibiograms in collaboration with private and government veterinary laboratories across each state or province would improve the veterinary antimicrobial stewardship program in each country. We recommend that veterinary-specific guidelines should be developed to allow comparisons between various cumulative antibiograms. We further suggest that a cumulative antibiogram should be presented as selective and cascade reporting (Tables 4 and 5). Only antimicrobials commonly used in the species of interest should be listed in a cumulative antibiogram, according to their importance rating, as per guidelines from each country.1,20 Furthermore, a cumulative antibiogram should be used in conjunction with other measures described in country-specific antimicrobial stewardship programs. Prospective veterinary studies are required to provide evidence that the use of cumulative antibiograms results in the improvement of patient outcomes and reduction of resistance development.
Footnotes
Acknowledgements
We thank Ms. Rochelle Price from the Veterinary Laboratory Services at the School of Veterinary Science, The University of Queensland, for her assistance with database technical support. Part of this paper was presented at Australian and New Zealand College of Veterinary Scientists Science Week 2019 as an oral presentation and the 41st Bain Fallon Memorial Lecture as a poster presentation.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was funded by a Morris Animal Foundation Veterinary Student Scholarship (grant D18EQ-603), The University of Queensland Summer Research Program, and the School of Veterinary Science, The University of Queensland (VETS5017 Veterinary Research Elective).
