Abstract
The exquisite sensitivity of in vitro amplification assays such as real-time polymerase chain reaction (rtPCR) requires the establishment of thorough and robust laboratory practices. To this end, an American Association of Veterinary Laboratory Diagnosticians (AAVLD) committee of subject matter experts was convened to develop a set of best practices for performance of nucleic acid amplification assays. Consensus advice for the performance of preanalytical, analytical, and postanalytical steps is presented here, along with a review of supporting literature.
Introduction
The polymerase chain reaction (PCR) assay, which mimics cellular genome replication, was developed in 1985.50,56,57 Improvements to the in vitro replication process over the subsequent 35 years have contributed to the increased use and utility of nucleic acid amplification approaches, and in particular PCR assays in diagnostic laboratories. Among the improvements, real-time PCR (rtPCR)27,29 has made the greatest impact, to date, for veterinary diagnostic laboratories (VDLs). The sensitivity of nucleic acid amplification–based assays, and in particular rtPCR assays, requires stringent adherence to performance guidelines to minimize contamination and to ensure consistent and reproducible results across different reagent lots, environmental conditions, and personnel within a laboratory. In veterinary laboratory testing, this is further complicated by the diversity of diagnostic materials involved, which includes multiple species, unique sample matrices, and samples arising from complex environments.
Here we describe the consensus document developed by the Laboratory Technology Committee (LTC) of the American Association of Veterinary Laboratory Diagnosticians (AAVLD), which addresses the performance of rtPCR in the unique situations encountered in veterinary diagnostic medicine. These best practices are applicable to rtPCR and to any molecular method that may have an amplification step as part of the protocol, including but not limited to: digital PCR, 78 isothermal PCR, 23 cycle sequencing, 70 and library preparation for whole-genome sequencing.2,5 Although this review uses the abbreviation rtPCR, the principles and practices discussed apply equally to reverse-transcription rtPCR (RT-rtPCR). Preanalytical, analytical, and postanalytical steps in the rtPCR process are all subject to standardization of practices. Preanalytical steps include design and preparation of the molecular testing area, as well as collection, transport, and storage of samples. Analytical steps include sample preparation, assay performance, and result interpretation. Postanalytical steps include reporting and assay-control trend analysis. Our best practice recommendations were harmonized with AAVLD accreditation requirements as discussed in each section. 1
Brief overview of rtPCR and recommended abbreviations
The basic process of amplification—strand separation, primer annealing, and extension—is the same in conventional and rtPCR. As rtPCR started replacing conventional PCR in VDLs, assays using intercalating dyes, such as SYBR Green, were often implemented based on cost-effectiveness (i.e., a costly probe was not required). In these assays, the target is quantified by measuring the increase in fluorescence generated as the dye intercalates into the increasing number of target copies generated during the amplification step. Intercalating dyes, such as SYBR Green, are nonspecific (i.e., bind double-stranded DNA), therefore if primers are not properly designed, undesired products such as primer–dimers can form and may be falsely identified as target.7,38 As fluorescent-labeled probes became increasingly available at lower costs, their use became more routine and provided an associated improvement in the specificity of the assay. The most commonly used probes in VDLs are those labeled with a fluorescent dye (reporter dye) at the 5’-end and with a quencher molecule at the 3’-end. The lower energy state of the quencher and its proximity to the reporter keeps the fluorescence quenched until the probe anneals to its target and the polymerization action commences. When cleaved from the probe, the reporter dye molecule fluoresces under external excitation from the instrument laser beam. The increase of fluorescence in an assay over time is the result of increasing levels of the reporter molecule in solution, indicating successful amplification of the target.
PCR abbreviations.
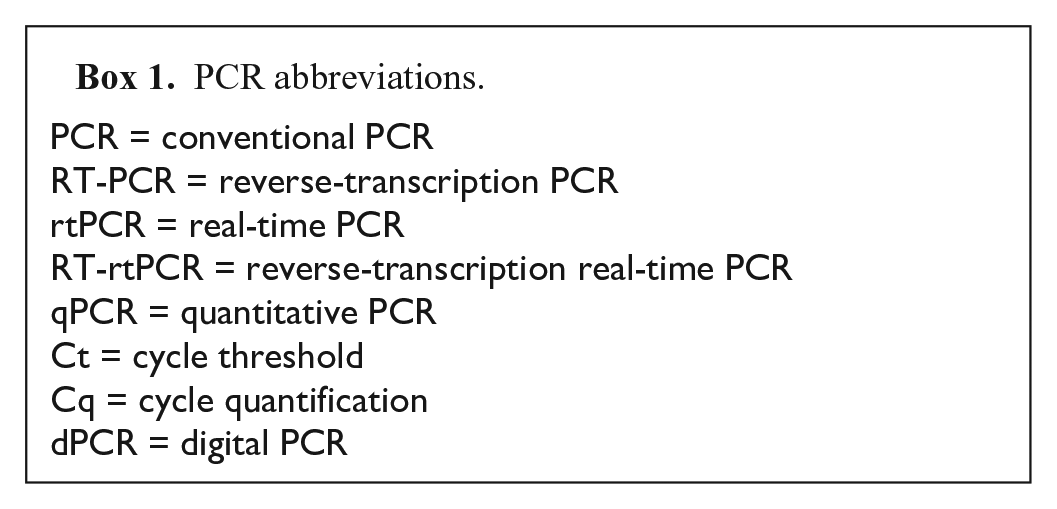

Quantification is possible with rtPCR and RT-rtPCR; however, most VDLs do not routinely perform quantitative rtPCR assays. The absolute quantification of nucleic acid targets requires standard curves, and although necessary during development and validation, 73 standard curves are rarely used in routine veterinary diagnostic testing because of expense. Instead, the rtPCR cycle at which a sample is labeled positive (cycle threshold, Ct) is established based on the relative amount of target nucleic acid in the test sample in relation to that of a thoroughly characterized positive control sample. Highly consistent performance of the positive control, as well as an internal control to confirm the absence of PCR inhibitors, is required for estimates to be meaningful. It is appropriate to use cycle quantification (Cq)6 –8,39 when reporting values from an assay that is performed with a standard curve. Digital PCR (dPCR), which enables quantification without a standard curve, produces a result in the format of copy numbers of nucleic acid target. 78 Box 1 lists recommended abbreviations for commonly used terminology associated with PCR methods. In-depth explanations of rtPCR and the different versions of the method are described elsewhere.23,27–29,37,38,44,78
Preanalytical: facilities and environment
Design of facilities
The potential for contamination of assays as a result of the physical layout of the laboratory was recognized early on, 39 with experts agreeing that separation of functions and establishment of a unidirectional workflow (i.e., pre-PCR to post-PCR) to avoid contamination between workstations was essential.59,82,86,87 The clean reagent preparation room should ideally be under positive pressure airflow, where air flows out of the room. The room with the greatest contamination potential (i.e., post-amplification room) should have the most negative air pressure, where air flows into the room. Ideally, workstations should be located in separate rooms and further equipped with dedicated biological safety cabinets or dead air boxes. When individual rooms are not available because of budget or space constraints, the use of a single room with multiple biological safety cabinets or dead air boxes providing enclosed and dedicated spaces for each task is possible. Listed below are the 5 essential workspaces and the functions associated with each area:
In addition to the 5 areas listed above, a separate area is strongly recommended for the preparation of positive controls containing a very high pathogen or target concentration (e.g., cell culture supernatants, bacterial cultures, allantoic fluids, synthetic DNA or RNA targets, and high copy number plasmid preparations). It is critical to avoid contaminating areas designated for diagnostic sample processing. 28
Other sources of contamination
Analysts should be aware of the potential for the introduction of contamination through their apparel. Personal protection equipment (PPE) should be changed before entering a clean area and, preferably, dedicated PPE should be used for clean tasks. Lab coats worn for production areas where large amounts of nucleic acid targets, such as bacterial or viral stocks, are used should not be worn in any PCR areas.48,81,82 Analysts should also be aware of the contamination risks posed by small lab equipment items such as tube and plate racks, plate sealers, ice buckets, etc.39,48,82 Multiple options are available for limiting the potential for fomite contamination, including use of disposable containers or use of ice beads and ice buckets that can be decontaminated between uses. In all pre-amplification PCR workspaces, the analyst must ensure that 96-well optical plates and strip tubes are in an appropriate support base to prevent nonspecific fluorescence contamination on the exterior of the tubes caused by environmental and bleach residues. The use of personal items (e.g., electronic devices such as cell phones) should be prohibited to prevent contamination of the device and subsequent contamination of the PCR environment, as well as to prevent pathogen contamination outside of the laboratory.10,63,75
Temperature and humidity
Monitoring environmental temperature and humidity helps to ensure that assay instrumentation performs properly. Manuals for amplification and robotic extraction equipment should be consulted for acceptable humidity as well as temperature ranges. Commercial extraction and assay kits may also specify an acceptable temperature range required for proper assay performance.
Disinfection and decontamination
A stringent routine should be in place to keep benchtop surfaces and all safety cabinets and dead air boxes clean and decontaminated. Chemicals appropriate for disinfection may be excellent at rendering pathogens noninfectious but fail in removing contaminating nucleic acid. Household bleach, typically 5–6% sodium hypochlorite as purchased, is diluted 1:10 to make a “10%” solution (i.e., final concentration of 0.55% sodium hypochlorite), which has been used reliably in molecular biology labs for decades20,53,54 based on its ability to degrade nucleic acid.21,53 Minimum contact time for bleach disinfection is documented as 2–10 min, dependent on the reference and the goal, with nucleic acid degradation on heavily used surfaces requiring longer contact times compared to that of visibly clean surfaces. 20 Wiping surfaces with individually packaged bleach wipes or utilizing bleach and water dispensing units that create fresh 10% bleach solution at the point of use avoids the risk of disinfection failures associated with the rapid degradation of diluted bleach solutions. Where local safety regulations or laboratory personnel sensitivity to bleach precludes the use of bleach, substitution with 3% hydrogen peroxide is acceptable and has been well documented to degrade DNA. 41 Hydrogen peroxide disinfecting wipes can be purchased in single-use packets. Copper-bis-(phenanthroline)-sulfate/H2O2 (CoPA) is a peroxide-based decontamination agent equal to bleach in effectiveness.11,81 Chlorine dioxide is a disinfectant widely used in veterinary medicine; however, in one report, disinfection did not degrade all genome fragments of a nonenveloped RNA virus. 64 More studies are needed to determine the efficacy of chlorine dioxide as a nucleic acid decontamination chemical. The use of UV lights contained in biological safety cabinets, when allowed by local regulations, may assist in decontamination 82 provided that the UV output is monitored during annual biological cabinet calibration and the bulbs are cleaned weekly to maintain efficacy. 47 UV light alone is not, however, a reliable way to control contamination given that only the surface contamination accessible to the light is eliminated; additionally, UV light can result in degradation of plastics.11,82 Decontamination requirements have been investigated thoroughly for fields that are especially sensitive to contamination, such as whole-genome sequencing 49 and crime or archeological forensics, 11,76,81 and can serve as good resources should contamination issues arise and persist.
Air and surface contamination monitoring
Schedules should be established for monitoring bench, biological safety cabinet, and dead air box surfaces in the processing areas as well as in autopsy suites. Swabs, sterile gauze pads, or commercial cloth cleaning pads 24 may be used to wipe surfaces at traceable locations using a key or map so that repeat testing can be performed. A best practice for monitoring surfaces is to swab the area and test using rtPCR for an organism that is resistant to degradation, such as a nonenveloped virus (e.g., rotavirus or parvovirus). Another monitoring strategy is to use spare wells in a 96-well plate and test multiple “no template” controls (~5% of wells) to check for aerosol contamination generated during template addition.82,86,87
Preanalytical: equipment
Pipettors
A designated set of pipettors should be available for each PCR workstation.39,76 Positive displacement pipettes were recommended to prevent contamination, 39 but the use of filter tips is now strongly recommended.48,76 All pipettors must be calibrated according to laboratory policy, and where applicable with regard to AAVLD or other applicable laboratory accreditation requirements.1,86,87
Real-time PCR instruments
Selection of rtPCR instruments includes the following considerations: number of reactions typically performed, reporter dye combinations needed, chemistries allowed, run time, and compatibility with automation. The choice may be limited if instruments are a component of laboratory networks (e.g., the U.S. Department of Agriculture, National Animal Health Laboratory Network) that require specific platforms in order to limit variability in assay performance among laboratories. Maintenance and calibrations should be regularly performed according to the manufacturer’s instructions, and where applicable in compliance with AAVLD or other relevant accreditation requirements. 1 Instruments under a service contract still require, at a minimum, monthly well cleaning and background checks performed by the laboratory according to the manufacturer’s recommendations. Also, it is advisable to monitor well performance by recording well positions to ensure that failed or contaminated reactions are not always occurring in the same position.
Automated extraction instruments
As with PCR instruments, different automated extraction instruments may yield different results because of the design and manufacturer. For this reason, laboratory networks, as previously noted, specify platforms for the performance of network-supported assays; review of any specific requirements is recommended before ordering equipment. Maintenance and calibrations are performed as instructed by the manufacturer and following laboratory policy, and where applicable with regard to AAVLD or other relevant accreditation guidelines. It is important to monitor the performance of each well, which can be accomplished by use of an exogenous internal control. 92 When equipment is relocated, automated extraction equipment and liquid handling devices require re-evaluation. For example, the magnet head of a magnetic bead processor extraction instrument must be visually checked to determine if misalignment of magnetic pins occurred during movement, including minor movements such as to a different side of the room. To test functionality, LTC members recommend testing an extraction protocol with an internal control in each of 96 wells in a commonly used matrix, followed by rtPCR testing for the internal control target to check that each well gives the expected value.
Preanalytical: sample acquisition and transport
Sample acquisition in the field or clinic
Proper collection of samples by technicians and veterinarians is critical to obtaining interpretable PCR results. Instructions to submitters regarding sample acquisition should be accessible on the laboratory’s website and include the main points identified in Box 2.
Practitioner communication points.
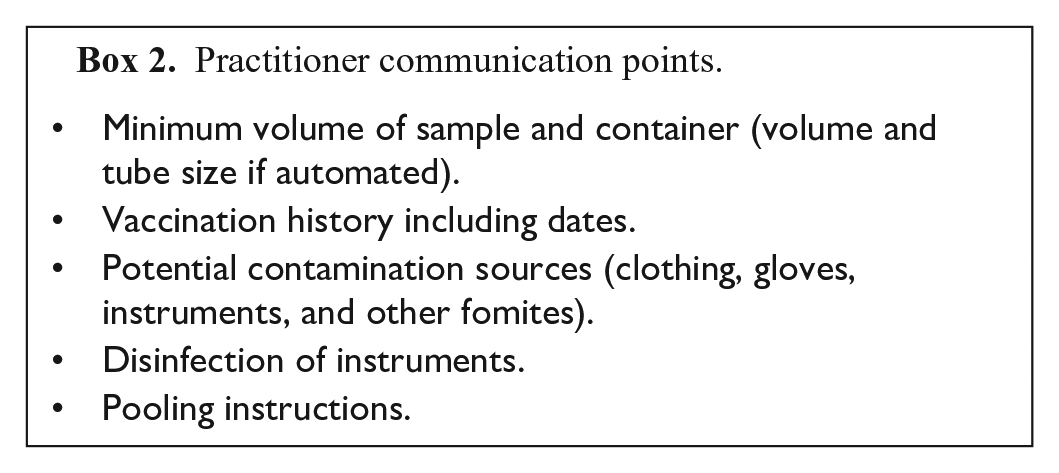

Work processes that are automated may require a certain diameter and depth of tube. Recent vaccination history, including the type of vaccine and route, should be recorded with dates administered because both modified-live vaccine agents 79 and inactivated vaccines 68 can be detected by PCR post-vaccination. Vaccination at the same time as sample collection should be discouraged because of potential contamination concerns. Practitioners should be made aware of the potential for contamination from other environmental sources such as clothing, gloves, and other fomites, as well as cross-contamination from reusable equipment such as ear notchers or knives, which should be disinfected with bleach or peroxide solution before collection of each sample. Additionally, thorough rinsing after bleach or peroxide decontamination is required to prevent false-negative results caused by PCR inhibition from residual bleach or peroxide. 76 Specimen pooling on the farm is often challenging because of the inability to effectively control contamination. Laboratories using pooling protocols should be aware that often the pooling potential is predicted from the Ct level of individual samples and is not empirically derived, and such predictions do not take into account the increased possibility of more inhibitors given the diversity and complexity of biological samples. A method comparability study55,73,89 is appropriate before adopting a pooling strategy.
Fresh fecal samples should be collected, as soon after voiding as possible, into a sterile container that can be sealed and placed inside secondary containment, such as a plastic bag, to prevent leakage and cross-contamination. 84 Blood samples should be collected with a separate sterile needle and syringe for each animal. Collection of sample types critical for the detection of reproductive disease and thus of concern for international trade are well-reviewed in the World Organisation for Animal Health (OIE) manual, and the individual pathogen chapters should be consulted. 83 Respiratory samples are usually collected with swabs. Calcium alginate swabs13,19 and bacterial transport swabs 22 are reportedly unsuitable for direct detection of pathogens by PCR. Swabs made with cotton tips and wooden applicator shafts have also been found to be unsuitable because of the presence of bleach, formaldehyde, or other toxic residues used to treat the cotton and wood, based primarily on studies using influenza virus samples.16,19,35,36 Additional studies using influenza virus indicate that polyethylene terephthalate or polyester tips and a plastic applicator shaft are preferable for the collection of samples, with optimal recovery obtained with nylon-flocked or foam-tipped swabs. 67 The development of universal transport media coupled with flocked or sponge-tipped swabs ushered in a new era in sample collection and transport for bacteria as well as viruses. These systems are reportedly suitable for direct PCR (i.e., no purification step).4,30,45,60,62,77 Stability studies for some of these systems show that nucleic acid integrity is maintained for as long as 14 d at 38°C. 18 Newer transport systems have also been developed for parasites, with collection of eggs subsequently preserved by vacuum packaging of the sample. 61 Although most of these studies were focused on human subjects, some of the target organisms are also found in animals. That said, method comparability studies73,89 are recommended before adopting universal transport systems. Emergency situations often arise and, in the absence of a suitable collection system, a sealed, sterile vial with several drops of saline can be a satisfactory alternative for the collection of viruses, bacteria, and parasites.
Several activated filter paper systems have been evaluated26,34,40 and reviewed 58 for specific agents and species. Activated filter paper systems for the collection of wildlife samples are reviewed in this focus issue. 33 As with universal transport systems, most of the efficacy studies have utilized human samples; comprehensive studies are urgently needed for veterinary specimens.
Preservation of nucleic acid and retrieval from paraffin-embedded tissues (PET)9,12,25,32,46 and evaluation of alternatives to formalin fixation are areas also being evaluated actively. 25 In general, the length of time that tissues are exposed to formalin has a measurable deleterious effect on the retrievability of DNA and an even more profound effect on retrievability of RNA, with an optimal time found to be 12–24 h. 12 In the same study, slower processing times for paraffin embedding resulted in improved RT-rtPCR performance. 12 Another study comparing different storage temperatures of buffered formalin-fixed PET and alcohol-fixed PET found that paraffin blocks held at 4°C or −20°C maintained nucleic acid integrity and rtPCR performance with very little difference in loss of sensitivity over a 9-y span, whereas detection capability was not maintained when blocks were stored at room temperature. 25 Acquiring knowledge of the retrievability of nucleic acids from preserved pathology submissions is important, both for current cases in which only formalin-fixed tissue is available and also for the conduct of retrospective studies using archived paraffin blocks.
Transport of field or clinic samples
Once collected, fresh samples should be transported to the testing laboratory using cold packs, with next-day delivery recommended. 84 Whole blood samples should be protected from freezing to protect red blood cells and peripheral blood monocytes from rupture, which could affect detection of a pathogen. Room temperature transport is acceptable for formalin-fixed tissues, activated filter paper, universal transport systems, and chaotropic salt buffers. Some sample types require special transport conditions (e.g., semen straws destined for bovine viral diarrhea virus testing, including PCR, must be shipped in a liquid nitrogen vapor tank).52,90
Sample acquisition in the autopsy suite
Tissue samples must be collected with decontaminated equipment and tools. Decontamination between tissues within a case is ideal, and between cases is critical given the detection sensitivity of rtPCR and potential for cross-contamination. Some controversy exists regarding retrieval of pathogens using swabbed tissues compared to homogenization of the tissue. Swabs of tissues are considered easier to process; however, method comparability data are needed to support equivalent recovery of the pathogen(s). Tissues can be collected in buffers that preserve nucleic acid (e.g., alcohol or chaotropic salt-based buffers) but only if subsequent culture and isolation attempts are not required. 58
Analytical: sample processing
Sample storage at the laboratory
Stability of the sample and optimal storage conditions are pathogen dependent. The OIE manual has dedicated chapters to major pathogens encountered in VDLs and should be consulted. Some high-consequence pathogens, such as influenza virus and exotic Newcastle disease virus, have sufficient data to support storage at 4°C for up to 96 h. 74 In one meticulous study, sample handling techniques assessed to determine the impact on detection of porcine reproductive and respiratory syndrome virus by rtPCR demonstrated that thawing conditions were critical, with thawing at refrigerator temperatures far superior to thawing at room temperature. 80 In the absence of specific data for a sample type or pathogen, it is recommended that a cold chain be maintained for fresh tissues as well as for swabs, feces, and bodily fluids and that these samples be tested as soon as possible. 84 Sample suitability and quality should be assessed carefully before processing by evaluating the appropriateness of the sample for the requested test, the suitability of transport media and conditions, and sample integrity. An unsuitable submission may be rejected for testing. If only a marginally acceptable sample is available and is tested, the client should be informed by a cautionary statement in the laboratory report.1,84
Pretreatment of sample
Sample pre-enrichment may be necessary before PCR amplification. Notably, PCR for
Processing at the diagnostic laboratory
Homogenization of tissues using ceramic or metal beads before extraction is the “gold standard” tissue preparation method. 58 Detailed drawings and photos of how to do this without contamination are referenced. 58 Disposable instruments such as biopsy punches, scalpels, or microtome blades are the preferred tools. Reusable instruments should be decontaminated with a DNA-degrading chemical (e.g., 10% bleach or 3% hydrogen peroxide) followed by proper rinsing before further sterilization. Molecular biologists with a focus on high-throughput testing have evaluated swabs in place of homogenization. In one such study, swabs of feces gathered with a concurrent scraping of intestinal mucosal cells gave results equivalent to homogenized tissue. 14 Again, comparability studies should be performed to show that no sensitivity is lost in agent detection.
Analytical: reagents and kits
Reagents and commercial kits
Core reagents, kits, and supplies should be assessed for suitability before use and include appropriate documentation of results in harmonization with laboratory policies and relevant accreditation standards (e.g., AAVLD requirements). 1
Confirmation of primer and probe quantification upon receipt from the manufacturer is recommended to ensure that the concentrations are correct, noting that quantification of fluorescent probes may differ from the manufacturer’s reported concentration, depending on the method used. 51 Lyophilization has been identified as a source of contamination, 39 which should be considered when concurrently ordering synthetic target plus the corresponding primers and probes from the same vendor.
Contamination of molecular reagents is well documented, indicating that it is not sufficient to rely solely on the certificate of analysis.11,17,20,31,43,48,76 Although rare, contaminants in commercial products can be problematic, as has been acknowledged by manufacturers; examples include carrier reagent (
In addition to monitoring for contamination, it is recommended that each lot of a commercial PCR enzyme kit be tested before use to ensure that the kit performs as expected. To limit resource expenditure, this assessment can be performed concurrently with the previous lot, using the most frequently performed assay, and monitoring with the positive reference control set at 1 log more concentrated than the limit of detection. Alternatively, new lots can be qualified with a reference panel. Suitability testing is performed in addition to the verification required for introduction of a new assay to the repertoire of tests for a laboratory.55,73,89
Control of nucleic acid cross-contamination during setup and performance of PCR may be assisted by the use of amplification reagents with modified nucleotides and uracil-N- glycosylase.42,48,53 This approach allows for degradation of residual amplicon that may be contaminating the reagents and environment. To facilitate workflow, many laboratories have found it convenient and reliable to create bulk master mixes including all components except amplification controls and nucleic acid target.65,66 As an additional step to reduce errors, some laboratories prepare bulk master mixes using a set amount (e.g., 100 reactions). Adoption of pre-made bulk master mixes should be evaluated by a method comparability study before implementation.55,73,89
Analytical: procedure performance
Extraction performance
Selection of the extraction process is critical in PCR assay development, with the extraction and PCR assay linked through concurrent validation as a system. 73 Recommended controls when performing extractions are listed in Table 1. An internal control is highly recommended and is discussed at length in an accompanying article. 92 Contamination of equipment may occur during the extraction of a strong positive sample or extraction of bulk positive control reagent because of the potential for aerosolized target to contaminate surfaces and then contaminate subsequent extractions. Consensus opinion of the LTC holds that, once extracted, it is unnecessary to quantitate the target nucleic acid before amplification.
Types of controls for extraction and PCR.
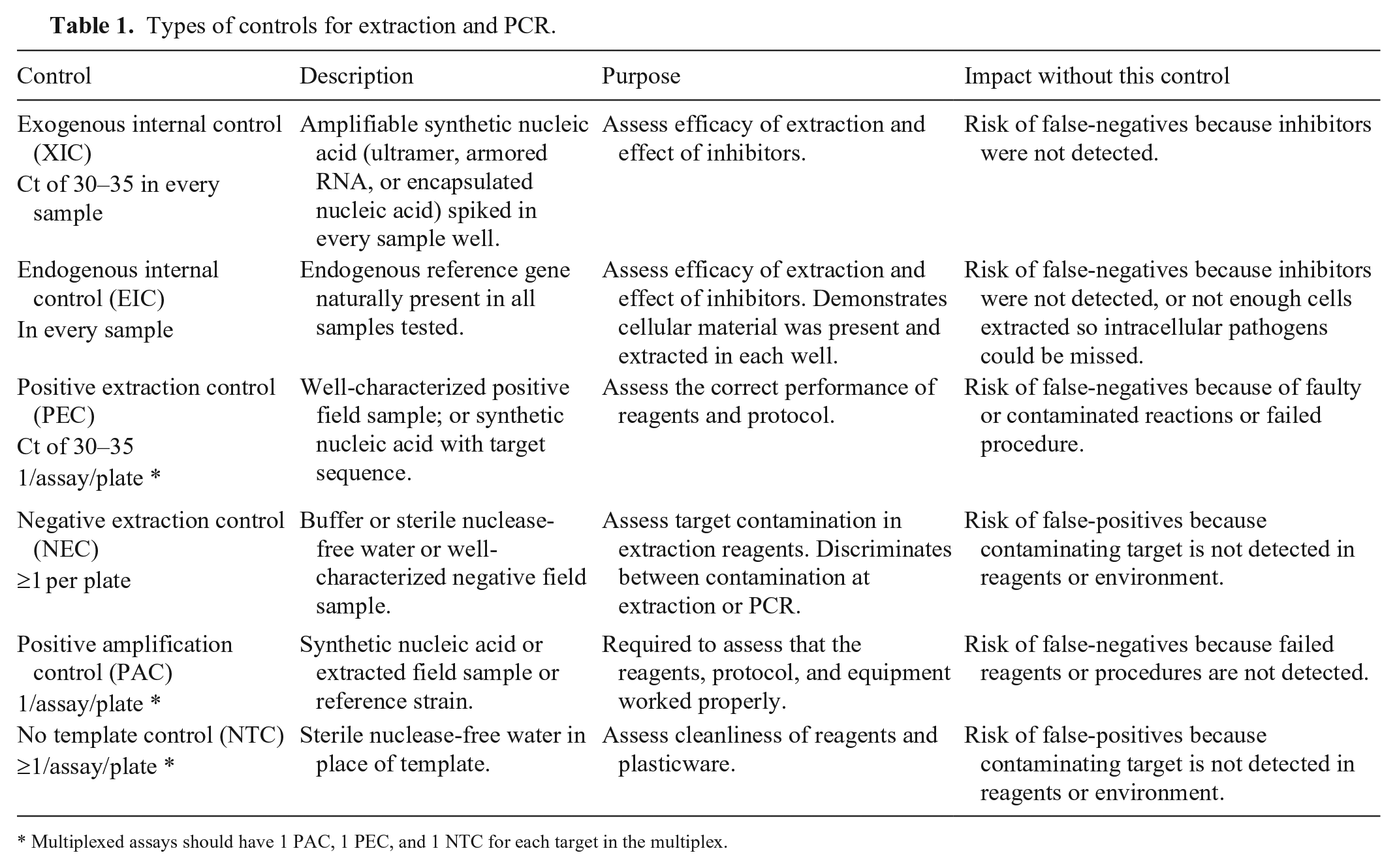
Multiplexed assays should have 1 PAC, 1 PEC, and 1 NTC for each target in the multiplex.
PCR performance
Monitoring a validated assay using a battery of controls is an essential activity in order to be congruent with OIE recommendations, the ISO:IEC 17025:2017 standard, and with AAVLD accreditation guidelines.1,85–87 At a minimum, it is recommended that a positive control and a negative control be included in each run for each assay. Selection of suitable controls is essential for ensuring full confidence in assay results. Reference strains suitable for positive controls are recommended by the OIE 88 and will assist with harmonization with the AAVLD accreditation standard. 1 The American Type Culture Collection (ATCC) site (https://www.atcc.org) also provides links to commercial firms and other national and international sources of standards. Instructions for determining the appropriate concentration for routine use of positive amplification controls (PACs) are described elsewhere.6,88 Briefly, a provisional control concentration is determined based on a recommended 15 independent determinations. After ≥1 mo of performing the assay, an established value with acceptable standard deviation is determined. An example of the proper determination of an exogenous internal control (XIC) is illustrated in an accompanying article in this focus issue. 71 VDLs often run multiplex assays in which more than 1 target can be detected. In these cases, there should be a PAC and a no template control (NTC) for each target (Table 1). Inclusion of a copy number control (transcribed RNA or synthetic DNA) is not required unless true quantification with a standard curve is being performed.
Standard operating procedures (SOPs) for the performance of rtPCR should include appropriate detail, not only with regard to correct handling of reagents (e.g., appropriate dilutions from stock solutions to working solutions) but also with regard to thorough decontamination of biosafety cabinets, dead air boxes, other workbenches, and equipment between setups and at the end of the day. A master mix calculator (e.g., Excel sheet with appropriate functionality and protection of cells containing the formulas) helps avoid calculation errors when determining volumes of the different reagents needed for master mixes. Several companies provide generic calculators that can be assessed by using the online search term “PCR master mix calculator”, whereas other companies provide calculators specific to their products.3,72 Of note, the order of adding reagents is of such importance to the enzymatic reaction that the master mix order must be followed. The enzyme is added last, unless already in a commercial master mix, to ensure the proper buffer concentration is present for the enzyme to retain activity.
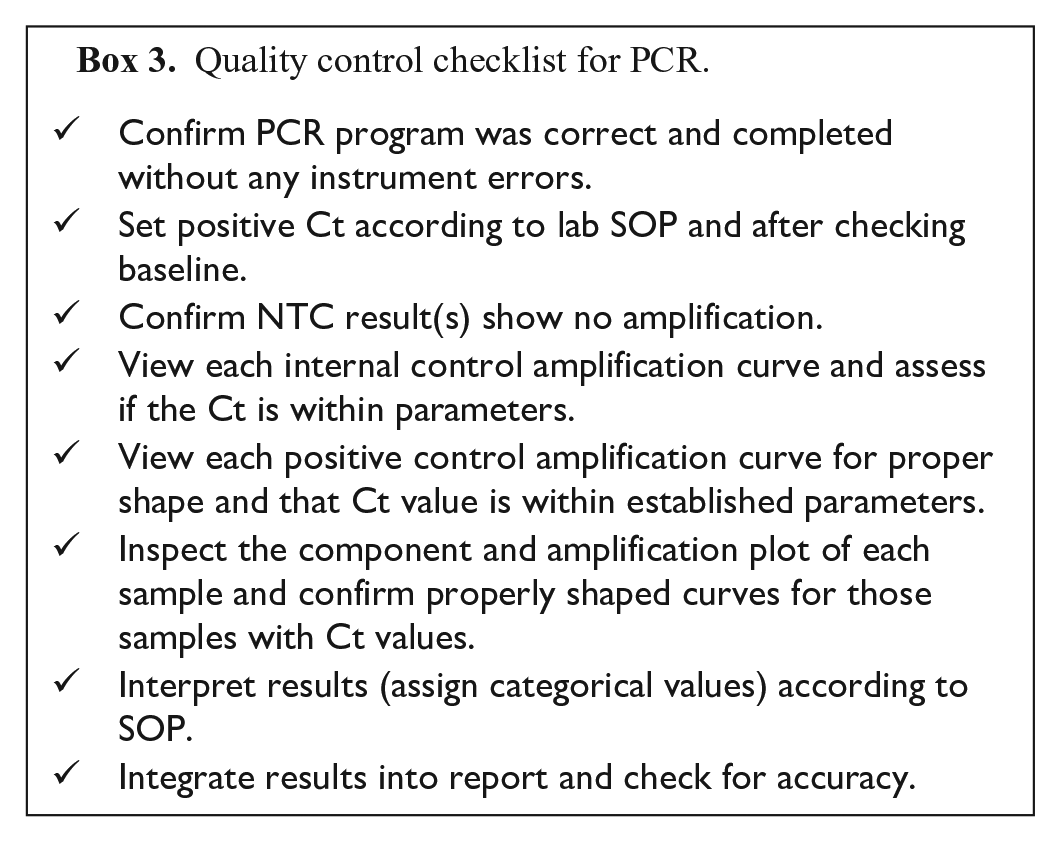
Interpretation of the PCR reaction must be described in the SOP. Threshold settings may be automatic or user-defined, dependent on the instrument brand. If user-defined, a set threshold may be dictated, or a value within 5 or 10% of the positive control can be used. 72 A list of quality control items to review for each run is included (Box 3). A critical component of quality control for each run is the inspection of the amplification plot and component plot of each sample to ensure that the software is calling the result correctly. When extremely strong signals are obtained, the automatic baseline setting provided by rtPCR software may be incorrect. For the thermocyclers used commonly in VDLs, the baseline is set from 3 to 15 cycles, and fluorescent signals obtained sooner than cycle 15 cause the software to generate nonsense amplification plots. Double-checking the baseline setting is advisable to ensure appropriateness for individual assays and signal strength of each sample.
Quality control checklist for PCR.
Postanalytical
Long-term monitoring is the most critical step in any validation process.6,85 Relevant reagent and kit lot numbers should be charted. All amplification control Ct values and inhibition issues should also be charted (Box 4). Trend charts should be reviewed daily to detect any emerging issues (e.g., assay failures, presence of contamination). Assessment of sequence suitability for each target should be performed routinely, with the LTC recommending at least annually to ensure that the assay primers and probe retain their sensitivity and specificity.7,15,28,38,86
Items for trend analysis.

Reporting of rtPCR results has been controversial, and there is no consensus on whether to report the Ct level as well as the interpretation (i.e., positive, inconclusive, not detected, etc.). Reporting of Ct levels requires substantial initial client education, but once this hurdle is cleared, clients may find that the Ct values help to interpret the significance of their results. However, client education must also emphasize that Ct interpretation can only be used for the specified assay performed in a particular laboratory unless the assay has been shown to be highly reproducible across laboratories. 88
Future directions
The LTC has provided here a set of best practices for performing rtPCR. The LTC recommendations outlined in this document were based on OIE chapters, publications, and collective experience of the LTC membership. Specific recommendations such as the use of amplification controls, extraction controls, and NTCs are universally found in guidance documents. Other practices, such as the use of internal controls, are strongly recommended but given the cost of the required method comparability studies, these may not be included as requirements in the relevant guidance documents. There were several areas identified while developing the LTC guidance documents presented in this special issue that will require future LTC activity, including: assessment of universal transport systems; assessment of decontamination systems; evaluation of newer amplification systems such as isothermal and digital PCR; all with the goal of developing additional best practices.
Footnotes
Acknowledgements
We are grateful for discussions and troubleshooting sessions with colleagues in our laboratories as well as discussions with LTC members. We thank Amy Glaser (retired from Population Medicine & Diagnostic Sciences, Cornell University) for thoughtful contributions over many years of developing these best practices.
Declaration of conflicting interests
The authors declared no potential conflict of interest with respect to the research, authorship, and/or publication of this manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
