Abstract
Male-to-female sex reversal in horses is a developmental disorder in which phenotypic females have a male genetic constitution. Male-to-female sex reversal is the second most common genetic sex abnormality, after X chromosome monosomy. All male-to-female sex reversal cases studied to date have been found to be infertile. Therefore, a screening test is particularly useful in laboratories doing DNA genotyping in horses. Our laboratory has tested > 209,000 horses for parentage using a panel of microsatellite markers and the sex marker gene amelogenin (AMEL). Suspect XY sex reversal cases are reported females with a male profile by AMEL testing. After routine genotyping, 49 cases were detected and further tested using the sex-determining region Y (SRY) gene, confirming the XY SRY-negative genotype of suspect sex reversal cases. When some inconsistencies arose in the initial result, a molecular panel of X- and Y-linked markers was analyzed for these samples. Of the 49 cases, 33 were confirmed as XY SRY-negative. The remaining 16 cases were identified as false-positives as a result of anomalies of AMEL testing in horses.
Keywords
Parentage verification is a common part of pedigree control programs. Our laboratory in Argentina tests annually ~ 15,000 horses of different breeds for identification and parentage purposes using a panel of polymorphic microsatellites and the sexing marker AMEL, a gene with distinct X and Y alleles (AMELX and AMELY).5,6 Since 2007, we have genotyped > 209,000 horses using AMEL, and have identified potential sex reversal cases.
Sex reversal disorders in mammals are described as a disagreement between sex chromosome constitution and gonadal and phenotypic/behavioral sex. Male-to-female, XY SRY-negative is the most frequent finding among sex reversal cases in horses. 10 Given that such horses are infertile, a molecular screening protocol based on AMEL is useful to detect inconsistencies between the reported and tested sex. When a declared female has a male profile by AMEL testing, the male virilization gene, SRY (sex-determining region Y gene), is tested. We found 49 horses suspected to have XY SRY-negative sex reversal phenotypes. Interestingly, some of these horses registered offspring and, in some cases, both the dam and the female foal shared the same anomalous AMELY genotype (Table 1). Therefore, we decided to investigate all 49 cases with an additional panel of X- and Y-linked markers used as the sex confirmatory panel.1,8
XY sex reversal suspect horses from different breeds detected by amelogenin (AMEL) testing. The sex panel of X- and Y-linked markers was used to confirm the sex reversal cases. Confirmed cases were identified as male, SRY (−); false-positive cases were named as female, AMELY (+).
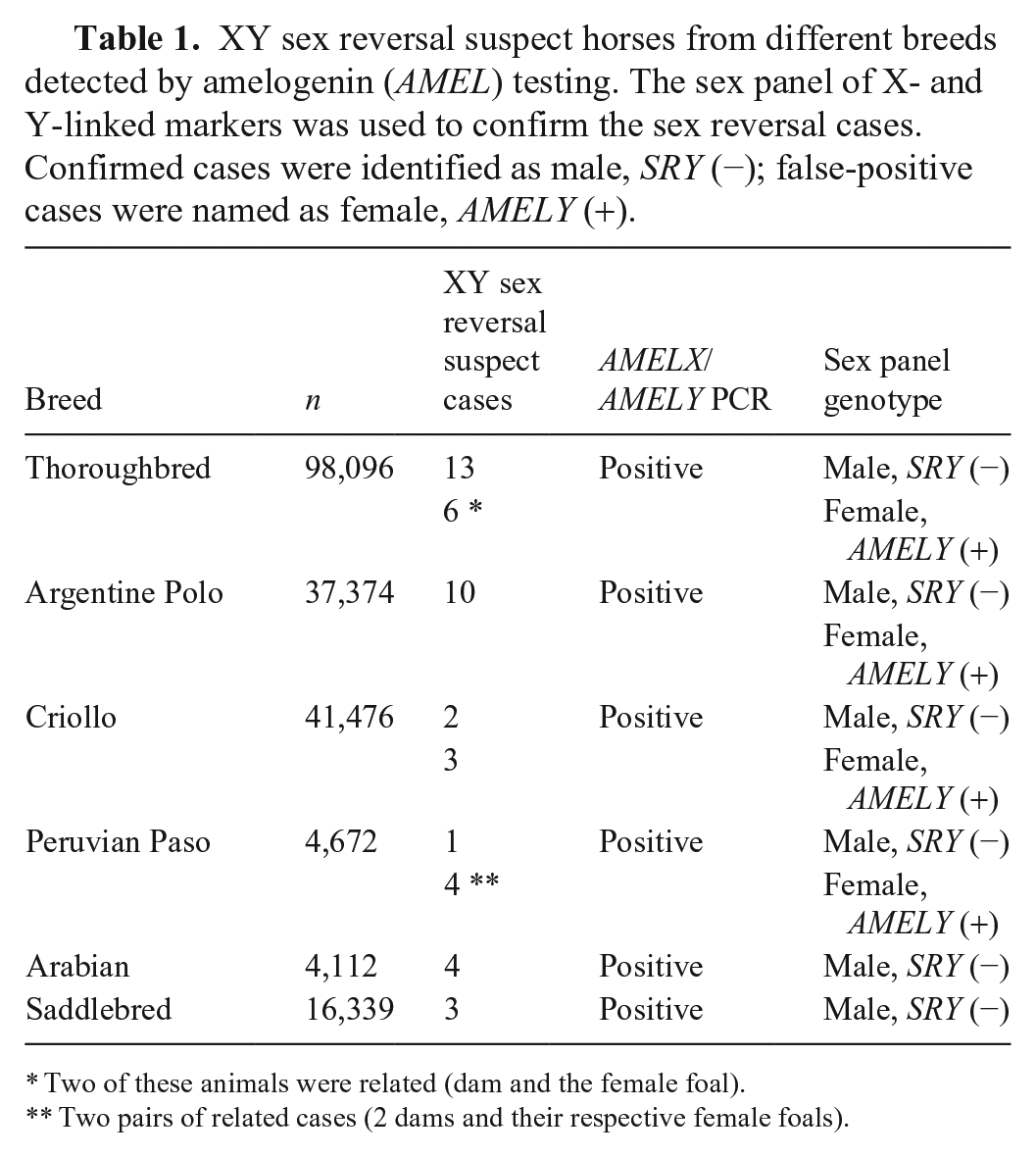
Two of these animals were related (dam and the female foal).
Two pairs of related cases (2 dams and their respective female foals).
DNA was extracted from 5 hair roots using 100 µL of lysis buffer (10 mM Tris HCl, pH 8.0, 50 mM KCl, 2.5 mM MgCl2, 0.5% Tween 20, 10 µg proteinase K; Invitrogen) and incubated at 60°C for 45 min followed by heating at 95°C for 45 min. The solution was tested directly for parentage with 15 horse microsatellite markers standardized according to the International Society of Animal Genetics (ISAG). 2 Microsatellites and AMEL were multiplexed in 2 mixes, as described previously (mix 1: VHL20, HTG4, AHT4, HMS7, ASB17, AHT5, HMS6, ASB23, TKY28, AMEL; mix 2: ASB2, HMS2, HTG10, HMS3, UCDEQ425, TKY325), grouped by their annealing temperatures (60°C and 56°C, respectively). 8 XY sex reversal suspect cases were initially analyzed with a mix of AMEL and Y-linked SRY genes (annealing temperature 58°C) and further tested with an extended sex panel of X- and Y-linked markers that were multiplexed into: mix 1 (X-linked microsatellites: TKY38, TKY270, Lex003, and Lex026; Y-linked microsatellite: ECAYA16; annealing temperature 50°C) and mix 2 (AMEL gene, Y-linked microsatellites: ECAYH12, ECAYM2, and SRY gene; X-linked microsatellites: Lex003 and UCDEQ502; annealing temperature 58°C). 8 Lex003 was duplicated to verify the sample identity, and marker genes SRY and AMEL were included in mix 2 for continuity with the first panel.
Each 25-µL reaction contained 20 ng of genomic DNA, 1× PCR buffer, 2.2 mM MgCl2, 0.3 mM of dNTPs (PCR buffer, MgCl2, and dNTPs; Invitrogen), 2% DMSO (dimethylsulfoxide; MilliporeSigma), 0.15–0.5 µM of each primer (Applied Biosystems), and 1 U Platinum Taq DNA polymerase (Invitrogen). The PCR protocol consists of 10 min at 95°C, followed by 31 cycles of 30 s at 95°C, 30 s at appropriate annealing temperature, 30 s at 72°C, and a final extension step of 72°C for 60 min. PCR products were separated by electrophoresis (3130xl; Applied Biosystems) and analyzed using GeneMapper v.4.0 (Applied Biosystems).
We analyzed the 49 suspect reversal cases using the sex panel; 33 samples had the profile expected for XY SRY-negative sex reversal cases (Fig. 1A) with products for all Y-linked markers and homozygous genotypes for X-linked markers. These profiles indicated a male XY chromosomal constitution with SRY mutation, yielding a non-virilized male. The additional 16 samples had a profile compatible with a female genotype (Fig. 1B), with no amplification of Y-linked markers (including SRY) and heterozygous genotypes for several X-linked microsatellites. AMEL was the only anomaly, showing both AMELX and AMELY alleles. Based on those results, these 16 samples were assessed as false-positive XY sex reversal cases, explaining the finding of offspring among cases tested (Table 1). The higher rate of these cases in some breeds may be explained by the larger number of horses genotyped, except for the Peruvian Paso with 4 cases, although they were related with 2 mares and their offspring. The false-positive rate was high (32.6%) for suspect XY SRY-negative cases, although extremely rare among all horses tested (0.0077%). This low rate may explain the absence of previous reports describing this anomaly in AMEL testing.
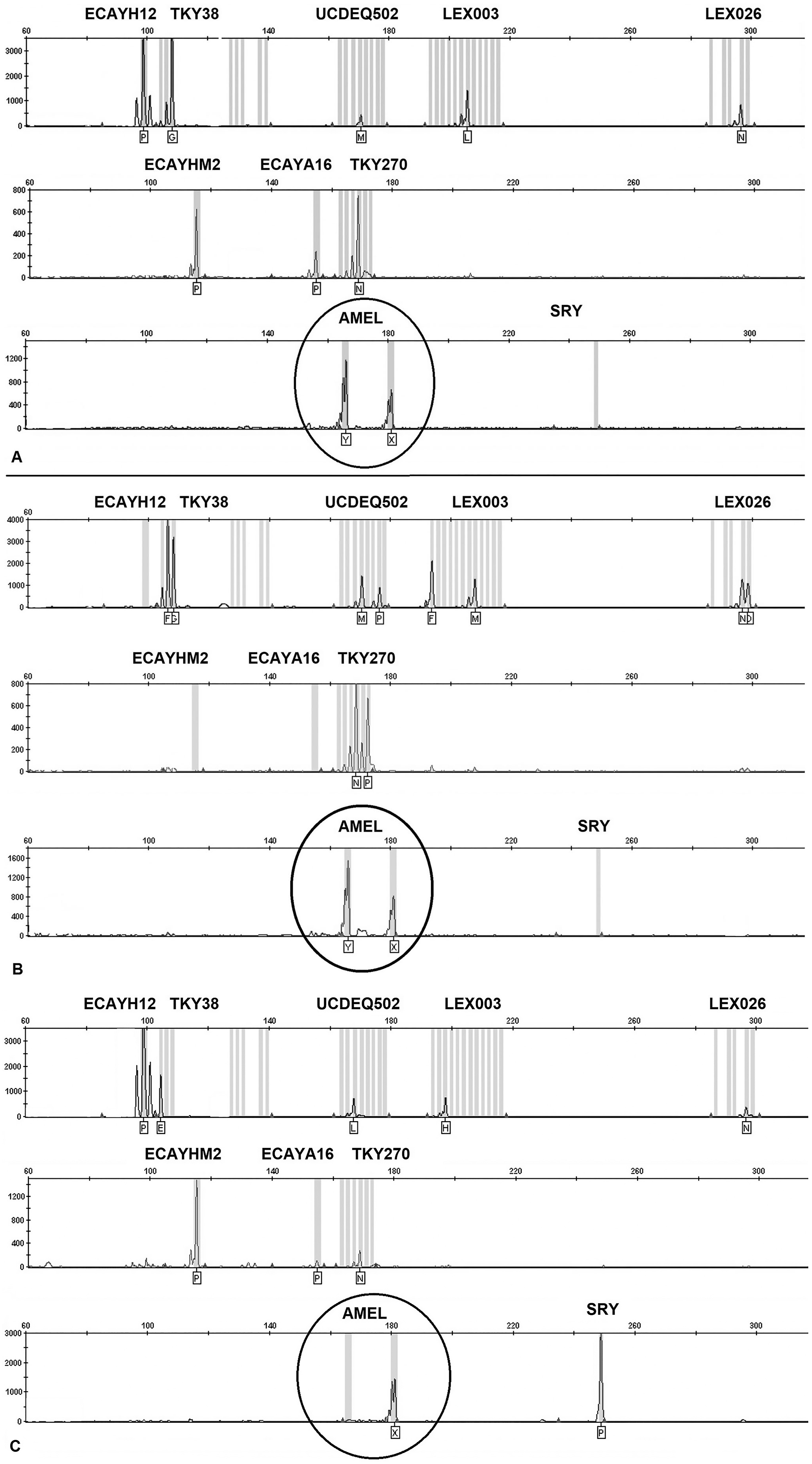
Electropherograms of X-linked markers (TKY38, UCDEQ502, Lex003, Lex026, TKY270, and AMELX) and Y-linked markers (ECAYH12, ECAYM2, ECAYA16, SRY, and AMELY):
We suspect that the presence of AMELY in these females might be the result of its translocation (in the paternal line) from the Y to another chromosome, given that AMELY is located in the distal extreme of the male chromosome in horses. 9 There was no similar proportion of SRY-positive XX males. Only one Thoroughbred horse was detected with a male profile by the sex panel, and it lacked the AMELY allele (Fig. 1C). The absence of AMELY in this case might be the result of amplification failure.3,4
AMEL genotyping anomalies have been reported in some human populations, although associated with AMELY or AMELX dropout.3,4 To our knowledge, AMELY-positive amplification in female horses yielding erroneous sex identification has not been reported previously in horses. Given that AMEL is included in many commercial parentage kits, and is used in forensic cases and for preimplantation sex diagnosis,5,7,8 awareness of this possibility would be helpful in the interpretation of anomalous results.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
