Abstract
Kobuvirus infections are common among humans, rodents, carnivores, pigs, and ruminants. We report herein the complete genome sequence of a novel caprine kobuvirus (MN604700) from diarrheic kids in Minnesota. Whole-genome sequencing revealed a kobuvirus genome of 8,139 nt with a single ORF region encoding a polyprotein of 2,480 amino acids. Further analysis revealed nt substitutions along the genome compared with that of the caprine kobuvirus reference strain, with 93% identity. Phylogenetic analysis indicated that the clade of the caprine kobuvirus was most closely related to porcine kobuviruses rather than bovine or ovine kobuviruses. Using primers designed from this genome, caprine kobuvirus was identified in the stools of other goats. Sanger sequencing of PCR products indicated
Introduction
Genus
Kobuvirus was first isolated in Japan in 1989 from humans with gastroenteritis following the consumption of raw oysters. 27 Subsequently, the virus was isolated from a wide range of hosts including human, ferrets, mice, foxes, bats, dogs, cats, cattle, sheep, pigs, and goats with and without clinical signs from many countries of the world.3,11,14,16,18,20,29,30 Caprine kobuvirus was detected in a black goat with diarrhea in the Republic of South Korea.13,21 In Italy, kobuvirus was detected in both healthy and diarrheic goats, with a higher percentage in diarrheic goats. 17
Most of the kobuviruses cause subclinical infections in their hosts but, in some cases depending on the immunocompetence of the animals, diarrhea and fever caused by viremia are observed.26,27 Next-generation sequencing (NGS) has played a great role in identification of
Materials and methods
Samples
In March 2017 and October 2019, the Minnesota Veterinary Diagnostic Laboratory (St. Paul, MN) received small and large intestinal samples from goat kids (10–14 d old) with a history of diarrhea followed by acute death. The submitting veterinarian reported that multiple kids were dying after developing diarrhea. Samples were received as segments of unfixed intestine and contents in Ziplock bags plus segments of formalin-fixed intestine. On gross examination, the large and small intestines were dilated and filled with gas and yellow, watery contents. No blood or mucus were observed in the submitted samples. Curdled milk was present in the abomasum; all other tissues appeared normal.
Bacteriologic and parasitologic examination
Swabs of intestinal contents were inoculated in cooked meat enrichment medium to detect anaerobic microorganisms (mainly
Histopathology
Pieces of formalin-fixed intestine were processed routinely, stained with hematoxylin and eosin, and examined under a light microscope. 28
Electron microscopy
For negative-contrast transmission electron microscopy (TEM), fecal samples from 4 goats were placed in a 20-mL tube, and double-distilled water was added to a final volume of 15 mL. After 10 min at room temperature, 1 mL of the fecal suspension was filtered through 0.45-µm syringe filters (Thermo Fisher). Fifty µL of filtered suspension was transferred to airfuge tubes (Beckman Coulter) with formvar-coated, 200-mesh copper grids (Electron Microscopy Sciences) and centrifuged at 148,000×
Next-generation sequencing
Total nucleic acid was extracted from intestinal contents (QIAamp MinElute virus spin kit; Qiagen) followed by submission to the University of Minnesota Genomic Center (UMGC) for complementary DNA synthesis, library preparation, and 250-bp, paired-end cycle sequencing (MiSeq; Illumina). CLC Genomic Workbench v.11.0.1 (Qiagen) was used to analyze the sequence reads. De novo assembled contigs were further confirmed by BLASTn and BLASTx analysis on NCBI (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). The ORF finder tool (www.ncbi.nlm.nih.gov/orffinder/) was used to find ORFs in the obtained sequences. Prediction of the presence of cleavage sites and structural and nonstructural proteins was done based on nucleotide (nt) and amino acid (aa) alignments, with reference strain KF793927.1 of caprine kobuvirus in GenBank.
Phylogenetic analysis
The complete genome sequence of caprine kobuvirus obtained in our study was aligned with published sequences of kobuviruses in GenBank using the Clustal W method in MEGA-X.
12
The maximum-likelihood method for analysis of the complete genome (GTR+G+I),
Primer design and amplicon sequencing
From the newly identified sequence obtained by NGS, we designed primers that could amplify the
Results
Bacteriologic, parasitologic, and histologic investigation
The bacteriologic examination did not result in any significant growth of anaerobic and/or aerobic bacteria. Parasitologic examination yielded no parasitic ova or oocysts. Histopathology revealed intestinal crypt hyperplasia and infiltration of lamina propria of the colon with numerous degenerate and nondegenerate neutrophils (Fig. 1). Postmortem autolysis precluded examination of the surface epithelium for erosions. Evidence of a coccidial infection was not detected in the small or large intestine.
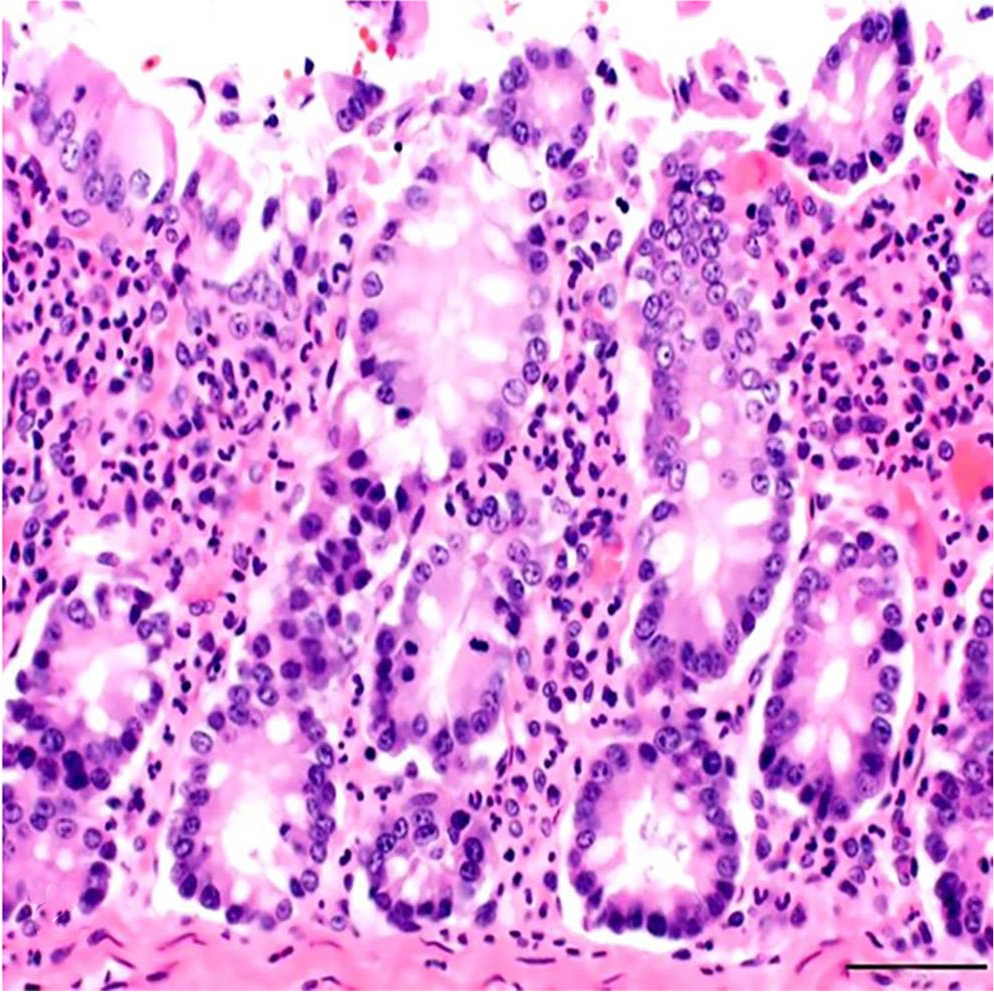
Colon of a goat with kobuviral infection. The lamina propria is markedly infiltrated with neutrophils. H&E. Bar = 50 µm.
Electron microscopy
Large numbers of nonenveloped, spherical particles (30.19–35.55 nm diameter; mean: 33.44 nm, SD: 1.43 nm) were visualized on TEM (Fig. 2).
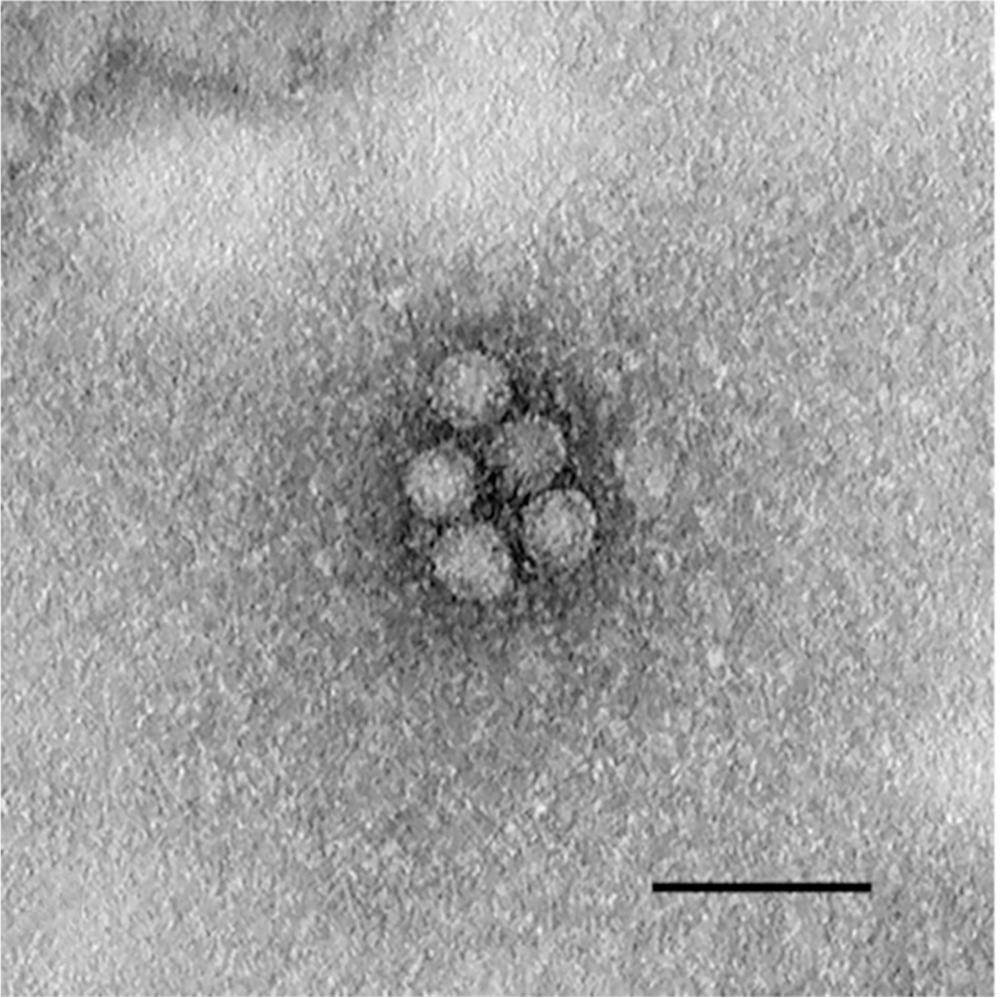
Ultra-microphotograph of negative-contrasted kobuvirus from goat 1. Nonenveloped, spherical viral particles. Bar = 100 nm.
Molecular investigation
NGS data analysis revealed that the distribution of sequences in the sample was 20% bacteria, 20% eukaryotic, and 49% unknown. However, we do not have the goat (host) genome in our reference dataset yet. Thus, a major portion of the “unknown” sequences are likely goat sequences. Only 1% of all sequences were classified as viral (which is common in metagenomic sequencing). Among the viral sequences, 34% (4,039 reads) of all viral sequences were classified as caprine kobuvirus. In addition, ~3% of viral sequences belonged to the
The almost complete kobuvirus genome sequence was assembled with 8,139 nt including 665 nt in 5′-UTR, 7,440 nt for ORF, and 34 nt in 3′-UTR. The single ORF region encodes a polyprotein of 2,480 aa beginning with a methionine codon and ending with an alanine codon. The complete genome sequence (caprine kobuvirus/MN-USA/2017) was submitted to GenBank (accession MN604700). Detection of nt variations with the closest caprine kobuvirus sequence (KF793927) revealed the insertion of 5 and 3 nt in
Substitution of nucleotides and amino acids along the caprine kobuvirus genome compared with the closest reference strain, KF793927.
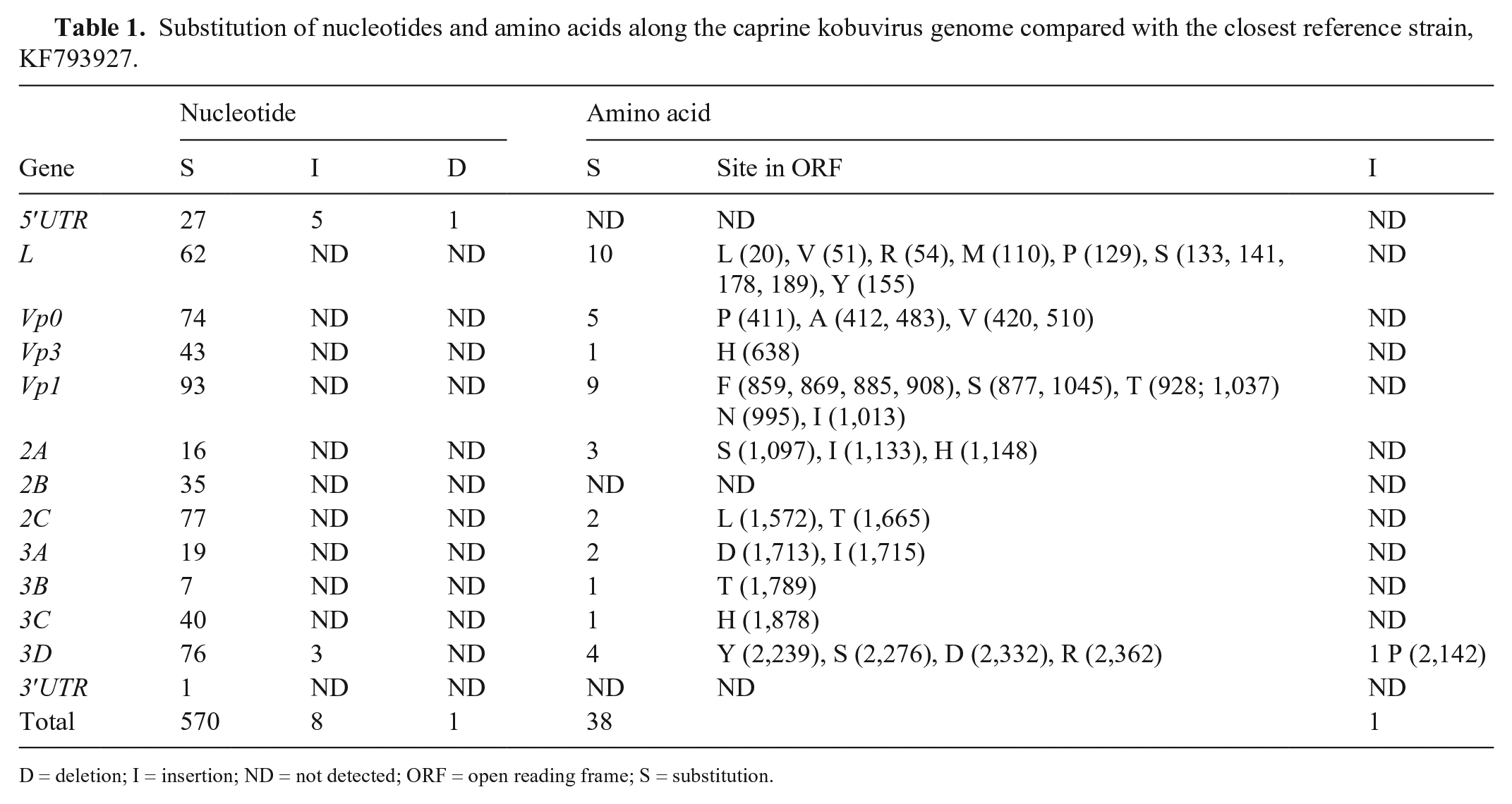
D = deletion; I = insertion; ND = not detected; ORF = open reading frame; S = substitution.
Phylogenetic analysis revealed that these caprine kobuviruses formed a new clade and were closely related to porcine kobuvirus rather than to bovine, and sheep kobuviruses, which are most related to each other (Figs. 3, 4). The
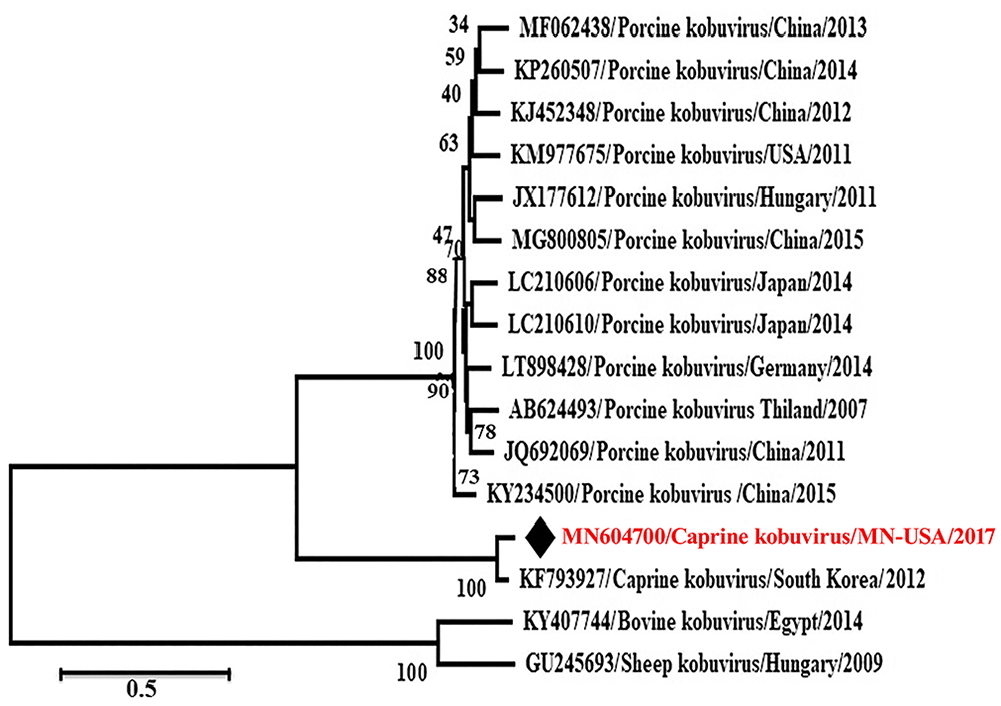

Phylogenetic analysis of MN604700 strain (caprine kobuvirus/MN/2017) complete open reading frame with other related viruses through the maximum-likelihood method by using the GTR+G+I model.
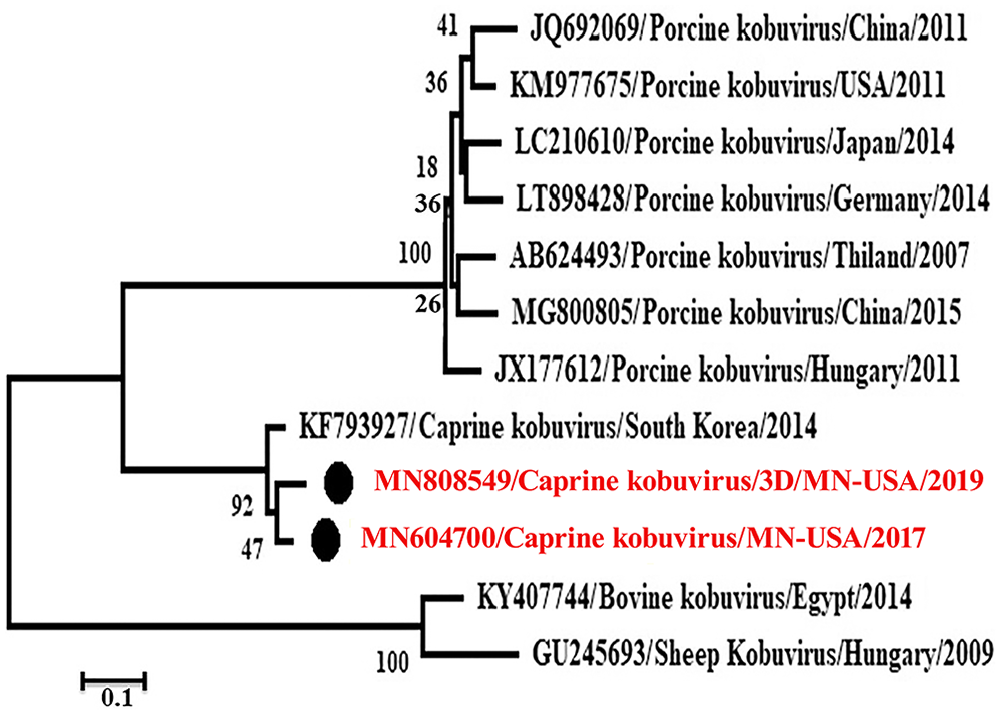
Phylogenetic analysis of the
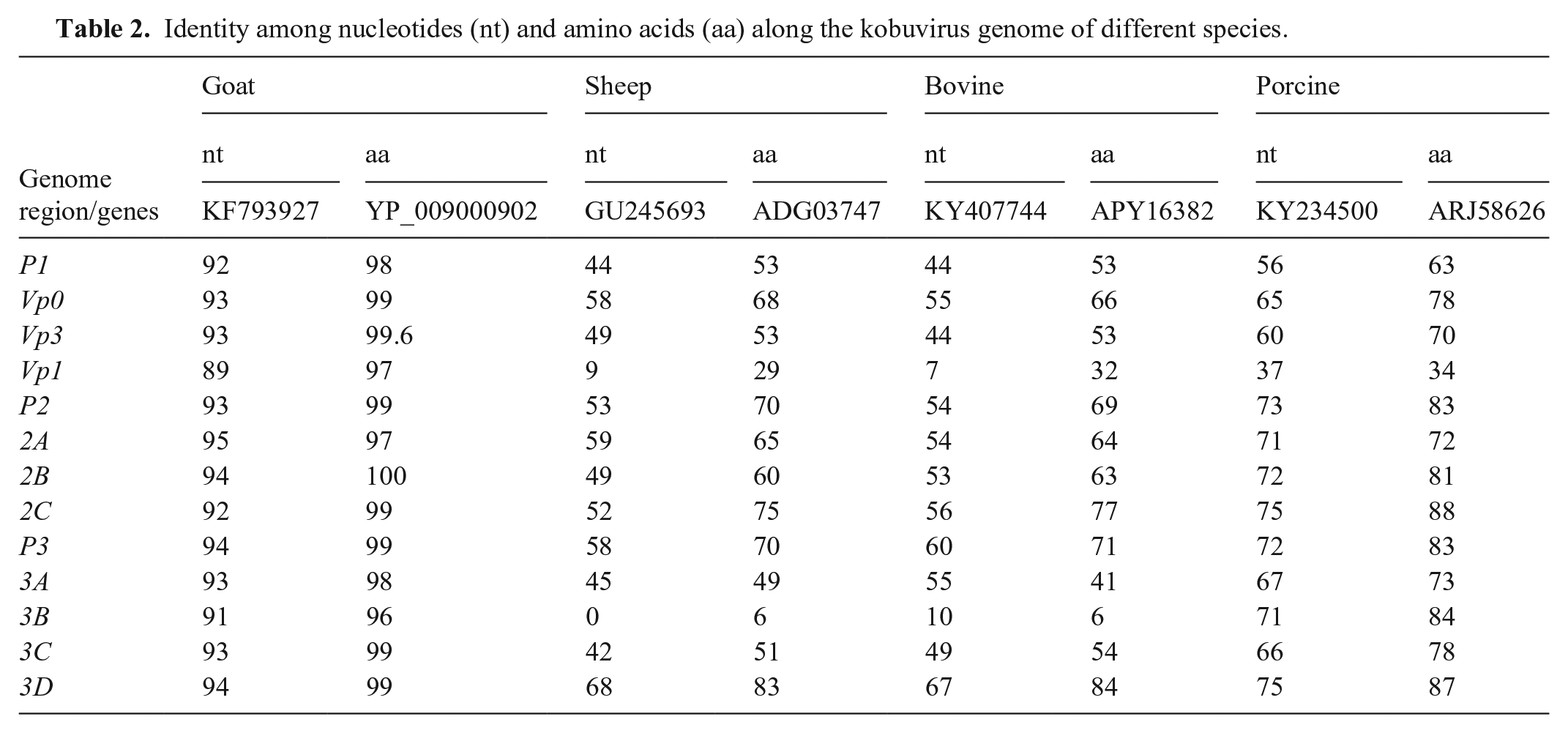
Identity among nucleotides (nt) and amino acids (aa) along the kobuvirus genome of different species.
The designed primers were used for testing samples from diarrheic goats obtained in 2019. Primers of the
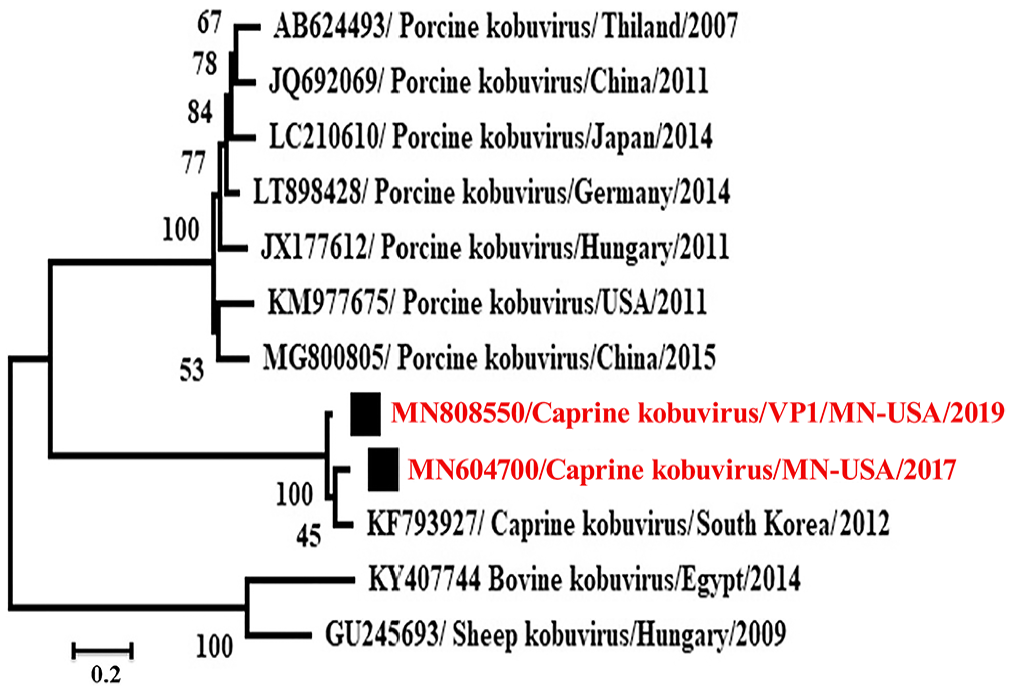
Phylogenetic analysis of the
Discussion
We detected the presence of a kobu-like virus by using electron microscopy, which is considered an optimal preliminary method for finding an unknown causative agent because it does not use reagents specific for a particular pathogen. However, it cannot be the only approach for virus detection because it requires a high concentration of virus in the sample for optimal sensitivity.4,34
Histopathology showed lamina propria infiltration with neutrophils, which would be consistent with intestinal erosion. Intestinal erosions are difficult to verify histologically unless optimally preserved intestinal samples are used. Molecular confirmation by WGS revealed the presence of kobuvirus in the above lesions. Analysis of the viral genome identified it as a caprine kobuvirus, with the same gene arrangement as the previously identified caprine kobuviruses.13,21 Incompatible with previous studies of
It is believed that each species’ kobuvirus is different from the others, and that there is a low possibility of cross-species transmission. However, historical transmission between rabbits and bats and between rodents and bats has been reported, which does raise the possibility of cross-species transmission in the future.2,15 Phylogenetic analysis shows that the clade of sheep kobuviruses is adjacent to that of bovine kobuviruses (Fig. 3), although there is no clear evidence of cross-transmission. The recombination rate in the virus is high; a previous study implied that a new porcine kobuvirus arose as a result of recombination between previously identified porcine and bovine kobuviruses. 10 Similarly, 3 strains of sheep kobuviruses in Brazil were near the bovine kobuvirus lineage on the phylogenetic tree. 5 Caprine kobuvirus was also identified in apparently healthy roe deer with 84.2–87.6% nt identity with caprine kobuvirus strain 12Q108. 7 The common finding of kobuvirus among a large class of wild and domestic animals all over the world indicates the widespread nature of these viruses and their potential to cause enteric disease.
Sequences of the
Identification of goat kobuvirus subsequently in 2019 indicates that the virus has been circulating in Minnesota for up to 2 y, sharing in the causation of goat diarrhea. The low identity of
Footnotes
Acknowledgements
We thank Wendy Wiese and Lotus Smasal of the Virology Section of the Minnesota Veterinary Diagnostic Laboratory.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
