Abstract
Using analytical chemistry techniques such as nuclear magnetic resonance (NMR) spectroscopy and liquid or gas chromatography–mass spectrometry (LC/GC-MS), metabolomics allows detection of most endogenous and exogenous metabolites in a biological sample. Metabolomics has a wide range of applications, and has been employed in nutrition science, toxicology, environmental studies, and systems biology. Metabolomics is particularly useful in biomedical science, and has been used for diagnostic laboratory testing, identifying targets for drug development, and monitoring drug metabolism, mode of action, and toxicity. Despite its immense potential, metabolomics remains underutilized in the study of spontaneous animal diseases. Our aim was to comprehensively review the existing literature on the use of metabolomics in spontaneous veterinary diseases. Three databases were used to find journal articles that applied metabolomics in veterinary medicine. A screening process was then conducted to eliminate references that did not meet the eligibility criteria; only primary research studies investigating spontaneous animal disease were included; 38 studies met the inclusion criteria. The main techniques used were NMR and MS. All studies detected metabolite alterations in diseased animals compared with non-diseased animals. Metabolomics was mainly used to study diseases of the digestive, reproductive, and musculoskeletal systems. Inflammatory conditions made up the largest proportion of studies when articles were categorized by disease process. Following a comprehensive analysis of the literature on metabolomics in spontaneous veterinary diseases, we concluded that metabolomics, although in its early stages in veterinary research, is a promising tool regarding diagnosis, biomarker discovery, and in uncovering new insights into disease pathophysiology.
Introduction
Metabolomics definition and broad applications
Metabolomics is an emerging “-omics” field aimed toward the comprehensive detection and quantification of metabolites and small molecules in a biological specimen.18,70 Combining advanced analytical techniques and chemometrics, 49 metabolomics enables researchers to identify a large proportion of metabolites (the metabolome) present in a sample, including amino acids, sugars, ketones, nucleotides, fatty acids, organic acids, microbial metabolites, and exogenous small molecules (including drugs, food additives, and pesticides).12,31 By analyzing these products of cellular metabolism, metabolomics reveals valuable information about an organism’s metabolic or physiologic state at the time of sampling.11,36
Metabolomics complements other omics technologies including genomics, transcriptomics, and proteomics, and there are increasing efforts to integrate these different data sets.77,86 With a tremendously wide range of applications, 42 metabolomics has been previously utilized in environmental analysis, 40 toxicology, 59 nutrition science,3,75 and systems biology. 74 In food science, metabolomics has been used in conjunction with traditional nutrition assessment methods to identify biomarkers that represent diet-related disease risks. 5 Agricultural and plant science industries have used metabolomics technologies to improve commercially significant traits and increase yield.26,38
In the biomedical sphere, metabolomics is being used to identify new disease biomarkers as well as provide novel insights into disease pathogenesis. The identification of endogenous and exogenous metabolites facilitates a better understanding of the complex changes that occur in metabolic and biochemical pathways. 47 Detecting complex changes in metabolite levels can not only aid disease diagnosis, 21 but can also monitor cellular responses to nutrition, 87 drugs, 32 toxins, 35 and environmental factors. 37 Given that altered metabolism is a key feature of cancer, metabolomics is playing an increasingly important role in cancer biology research, with uses ranging from detecting key regulatory molecules involved in carcinogenesis to identifying specific biomarkers for diagnosis.29,72 In the pharmaceutical industry, metabolomics can identify targets for drug development, 13 assist in mode-of-action studies, and monitor drug metabolism and toxicity. 66
Analytical techniques
Metabolomic analyses commonly use one or more analytical techniques to facilitate identification and quantification of as many metabolites as possible in a biological sample (Fig. 1).2,61,69,82 Nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS) are, to date, the most common techniques employed in metabolomic studies.47,58 One-dimensional NMR spectroscopy is commonly used to detect 10s–100s of metabolites in biological extracts. It is non-destructive (allowing reuse of the sample for other analyses) and can be used to quantify metabolites with high accuracy and reproducibility. Two-dimensional NMR techniques can be used to confirm or elucidate the structures of previously unidentified metabolites, as well as measure the incorporation of stable isotopes in labeling experiments. 43 MS techniques involve the ionization of derivatized or underivatized samples, and detection of corresponding charged ions (as mass-to-charge ratios). 6 MS techniques are generally more sensitive than NMR techniques, and allow greater coverage of metabolites (100s to ~1,000s) in biological extracts.
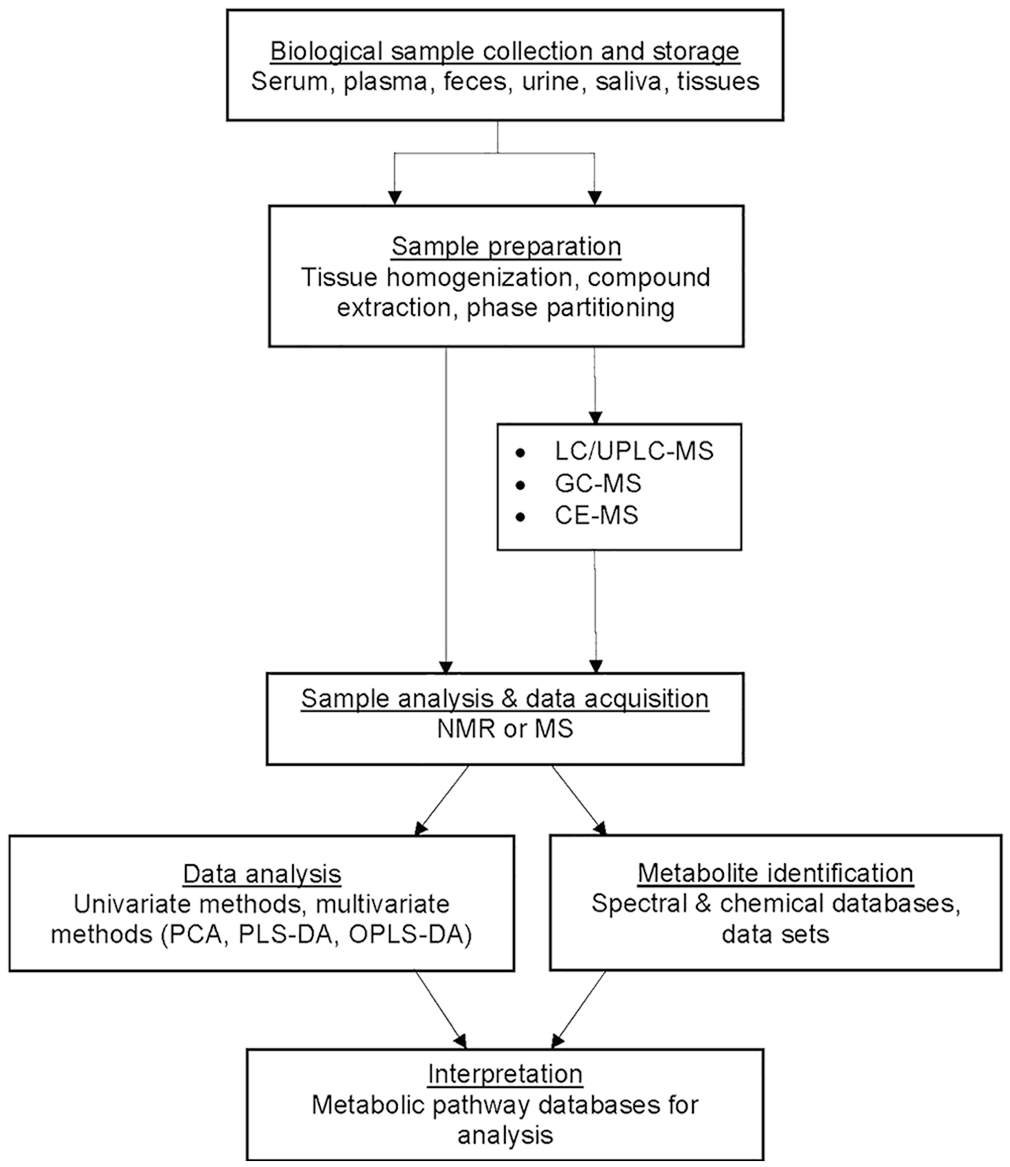
MS is often coupled with chromatographic separation techniques such as gas or liquid chromatography (GC, LC) or capillary electrophoresis (CE). Chromatographic separation of samples increases the sensitivity of MS detection (by minimizing ion suppression effects associated with complex mixtures and allowing greater sample loading) and provides orthogonal information (retention time prediction) that allows metabolite identification.2,31,47,62,82 The capabilities of GC-MS and LC-MS techniques have advanced enormously in recent years, with the use of high-resolution, accurate-mass MS instruments, such as the orbitrap MS and Fourier-transform ion cyclotron resonance MS (FT-ICR-MS). New-generation MS instruments that allow post-chromatographic separation of analytes by ion mobility instruments have the capacity to increase metabolite coverage, by allowing separation of metabolite isomers with the same mass and providing information on the shape (collision cross-section) of molecules that can be used for metabolite identification.
Given that no single platform offers complete coverage of either polar or apolar metabolites, the use of multiple complementary analytical platforms is important in initial exploratory studies on new biological systems. On the other hand, targeted metabolomic analyses, which focus on accurate quantification of a smaller number of metabolites, might be appropriate in biomarker studies. Both untargeted and targeted metabolomic approaches require the use of univariate or multivariate pattern-recognition techniques to identify differences between samples and generate new testable hypotheses.31,78,85 Other exciting opportunities in metabolomics include the use of imaging modalities (such as matrix-assisted laser desorption/ionization (MALDI) and desorption electrospray ionization (DESI) imaging–MS to detect spatial changes in metabolite levels within different tissue types.
Despite its enormous potential, metabolomics has been relatively underutilized in veterinary medicine. Our aim is to comprehensively review, tabulate, and analyze the existing peer-reviewed literature on the use of metabolomics in spontaneous veterinary diseases.
Methods
Search strategy
We searched the public databases PubMed, Web of Science, and CAB using the following search terms:
Metabolomic* or metabonomic* or metabolome or metabolic profil* or metabolomic fingerprint* or metabolite*
Veterinary* or veterinary medicine/science or livestock or small/large animal
Dog* or bitch* or canine* or canid* or canis
Cat* or feline* or felid*
Cow* or cattle or bovine or bovid*
Sheep* or ovine or small ruminant*
Horse* or equine or equid* or racehorse*
Pig* or piglet* or porcine or swine
and the combinations (1 and 2) or (1 and 3) or (1 and 4) or (1 and 5) or (1 and 6) or (1 and 7) or (1 and 8)
The use of the asterisk wildcard character allows searching for all possible suffixes. “Or” is used inclusively to search alternative terms (search results contain one or multiple phrases). The literature search (title and abstract) was conducted February 5, 2019 (Fig. 2). The titles, abstracts, or full texts were assessed for eligibility; selected articles were screened, and those that did not meet the inclusion criteria were eliminated. References of pertinent studies were also searched to identify articles for review.
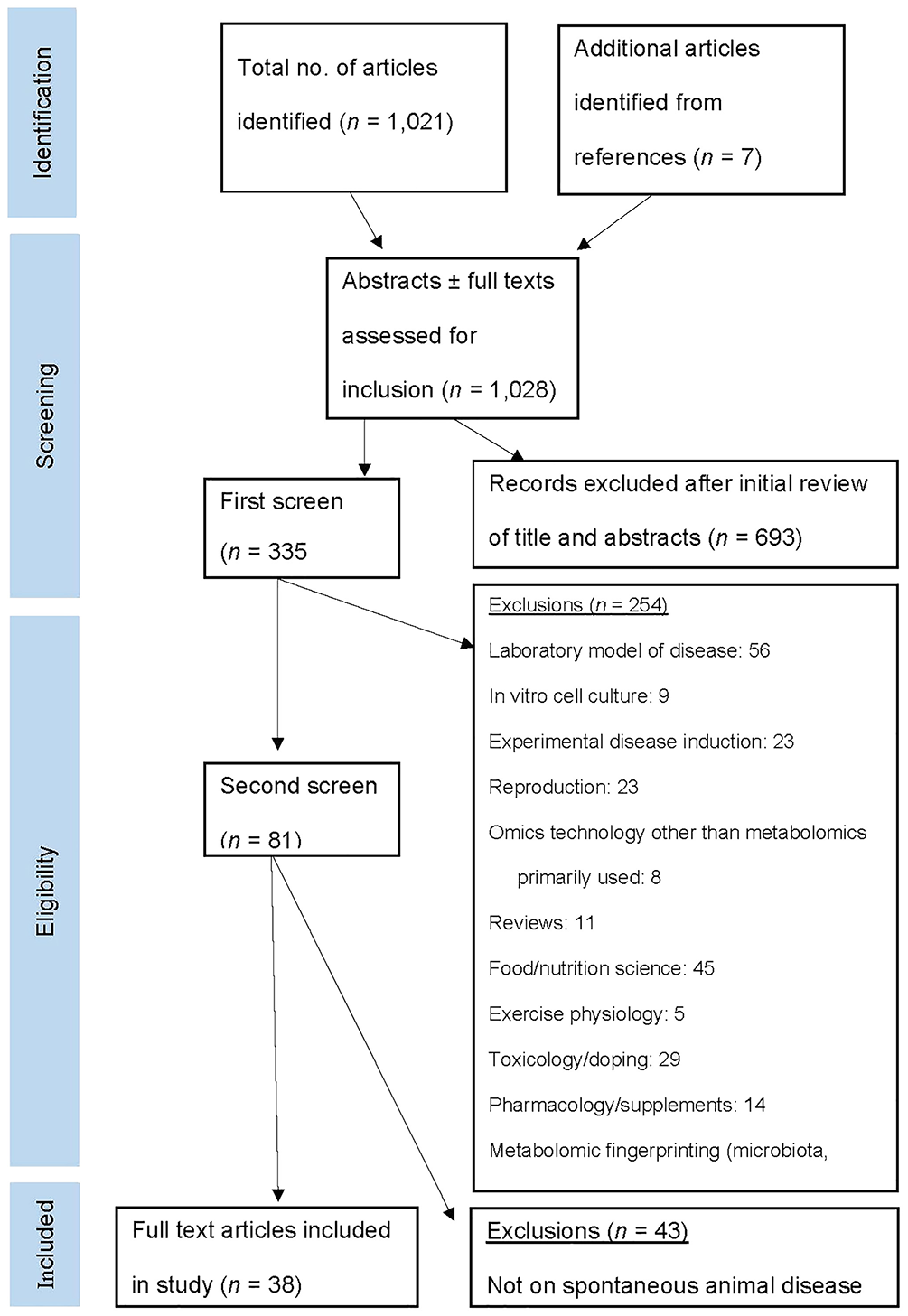
Search and selection strategy. See Table 1 footnotes for definitions of abbreviations.
Screening process and data extraction
Studies were eligible for inclusion if they were original, peer-reviewed research articles published in the English language, and if the primary aim was to apply metabolomics to investigate spontaneous animal disease. Exclusion criteria included experimentally induced disease, in vitro cell culture studies, laboratory models of disease, and studies that focused predominantly on an omics field other than metabolomics. Given practical constraints and brevity, we did not include studies on thermal stress, toxicology, or food or nutrition science in our review. Full texts of the relevant studies were retrieved, and data on species, diseases, metabolomic techniques, and results were extracted.
Results and discussion
Thirty-eight studies on metabolomics in veterinary medicine met the inclusion criteria (Table 1). The selected studies were published in 2005–2018. Eighty-eight studies were excluded because the researchers induced disease experimentally or used laboratory animals or cell culture lines to model disease. Forty-five were eliminated because they primarily focused on food or nutrition science; 29 were removed because they focused on toxicology or doping.
Thirty-eight metabolomic studies on spontaneous veterinary diseases included in our review.
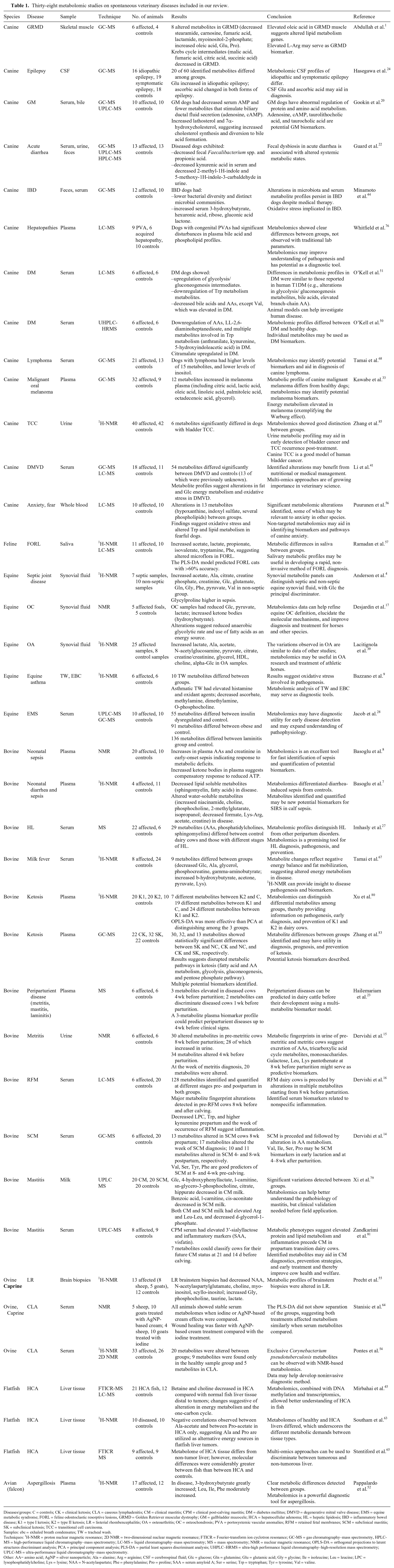
Diseases/groups: C = controls; CK = clinical ketosis; CLA = caseous lymphadenitis; CM = clinical mastitis; CPM = clinical post-calving mastitis; DM = diabetes mellitus; DMVD = degenerative mitral valve disease; EMS = equine metabolic syndrome; FORL = feline odontoclastic resorptive lesions, GRMD = Golden Retriever muscular dystrophy; GM = gallbladder mucocele; HCA = hepatocellular adenoma; HL = hepatic lipidosis; IBD = inflammatory bowel disease; K1 = type I ketosis; K2 = type II ketosis; LR = listerial rhombencephalitis; OA = osteoarthritis; OC = osteochondrosis; PVA = portosystemic vascular anomalies; RFM = retained fetal membranes; SCM = subclinical mastitis; SK = subclinical ketosis; TCC = transitional cell carcinoma.
Samples: ebc = exhaled breath condensates; TW = tracheal wash.
Techniques: 1H-NMR = proton nuclear magnetic resonance; 2D NMR = two-dimensional nuclear magnetic resonance; FTICR = Fourier-transform ion cyclotron resonance; GC-MS = gas chromatography–mass spectrometry, HPLC-MS = high-performance liquid chromatography–mass spectrometry; LC-MS = liquid chromatography–mass spectrometry; MS = mass spectrometry; NMR = nuclear magnetic resonance; OPLS-DA = orthogonal projections to latent structures discriminant analysis; PCA = principal component analysis; PLS-DA = partial least squares discriminant analysis; UHPLC–HRMS = ultra-high performance liquid chromatography–high-resolution mass spectrometry; UPLC-MS = ultra-performance liquid chromatography–mass spectrometry.
Other: AA= amino acid; AgNP = silver nanoparticle; Ala = alanine; Arg = arginine; CSF = cerebrospinal fluid; Glc = glucose; Gln = glutamine; Glu = glutamic acid; Gly = glycine; Ile = isoleucine; Leu = leucine; LPC = lysophosphatidylcholine; Lys = lysine; NAA = N-acetylaspartate; Phe = phenylalanine; Pro = proline; SAA = serum amyloid A; Ser = serine; Trp = tryptophan; Tyr = tyrosine; Val = valine.
Study characteristics
Regarding the technique used, over half of the metabolomics studies (22 of 38) used MS as the sole analytical platform. Two additional studies used FTICR-MS; only one used both NMR and MS platforms.
Regarding the species studied, 13 of the 38 studies were on dogs, 5 on horses, 12 on cows, 3 on small ruminants, 3 on fish, and 1 on birds. Interestingly, only one study focused on feline disease, suggesting that cats are grossly underrepresented in metabolomics studies despite representing a substantial proportion of veterinary patients. No swine studies met our inclusion criteria (Suppl. Table 1).
Regarding the main system investigated, diseases of the digestive system (11 of 38) made up the largest proportion of researched conditions. The second most commonly studied system was the genitourinary or reproductive system, with 7 studies. The remainder of the studies were on systemic disease (n = 3), diseases of neuromuscular or central nervous system (n = 3), the musculoskeletal system (n = 3), the respiratory system (n = 2), the lymphatic system (n = 2), and the cardiovascular system (n = 1). The remaining 6 studies were classified as “other,” and included conditions such as canine anxiety, canine diabetes mellitus, bovine milk fever, and bovine ketosis.
Inflammation (including infectious and non-infectious subcategories) represented the largest proportion of disease processes, with 15 of 38 studies; 7 studies concerned neoplasia, 2 vascular diseases, and 3 degenerative conditions. The remainder of the diseases were classified as “other,” and were subdivided into metabolic conditions (n = 8) and idiopathic (n = 3).
All bovine studies were related to reproductive diseases of economic importance, including metritis, retained fetal membranes, mastitis, and milk fever. This may reflect the growing interest in the use of multi-omics techniques for production improvement, a trend that has been demonstrated previously in agricultural sciences.
During our search, we found numerous studies that did not meet the inclusion criteria but are nonetheless relevant to veterinary science. These studies were outside the scope of our review. Evaluation of the quality of published papers was also beyond the scope of our study.
Principal findings
All studies in our review detected statistically significant differences in the metabolome of diseased and non-diseased states, suggesting that nearly all spontaneous diseases of veterinary interest are characterized by altered host cellular and/or microbial metabolism. In fact, most studies identified 10 or more metabolite differences in the diseased subjects compared with controls. Seventeen studies identified specific metabolites that could serve as biomarkers of the disease of interest in a clinical setting. A biomarker is a measurable biochemical indicator of a biological state, including normal or pathologic processes;46,71 in addition to their diagnostic utility, biomarkers can also help monitor a patient’s response to treatment. As an example, a 2018 study used a targeted metabolomics approach to evaluate the pathogenesis of retained fetal membranes in dairy cows, and to identify potential biomarkers that may serve as early predictors of disease. 16 Multiple metabolite alterations were identified as early as 8 weeks prepartum; however, many of the metabolites reflected the presence of inflammation and may not be specific to retained fetal membranes. This highlights the importance of establishing whether changes in the metabolome are specific to disease when evaluating the utility of metabolomics in biomarker discovery. 34 Other challenges in developing biomarkers in veterinary medicine include validation and qualification of biomarkers, 46 which no study in our review achieved.
Our study also reveals that metabolomics in veterinary medical research can complement our understanding of human disease. As an example, a 2012 study on canine transitional cell carcinoma found that affected dogs had increased levels of citrate and beta-hydroxybutyrate in their urine. 85 These aforementioned metabolites are similarly elevated in the serum of humans with esophageal adenocarcinomas, suggesting changes in Krebs cycle activity in epithelial malignancies,84,85 regardless of the host species. Similarly, a 2017 study identified similar serum metabolites in canine diabetes mellitus and human type I diabetes, including changes in glycolytic intermediates and elevated levels of branch-chain amino acids. 51
Reflecting the increasingly recognized importance of the microbiome, multiple studies explored the relationship between host health and the gastrointestinal microbiota.22,44 As an example, dogs with acute diarrhea were found to have decreased fecal concentrations of Faecalibacterium spp. and propionic acid, a short-chain fatty acid (SCFA). 22 Although host–microbiome interactions are complex and dynamic, this finding suggests that dysbiosis in acute diarrhea may have a direct impact on SCFA concentrations. 22 Additionally, as circulating metabolites may be derived from microbes rather than host cells,25,48 observed metabolite alterations may reflect microbiome changes rather than host cellular changes in disease states. Further research on the gut microbial–host co-metabolism is needed to improve our understanding of disease pathogenesis and potential treatments.
Many included studies did not acknowledge or control for concurrent processes that may arise in disease, such as inappetence or dehydration. Therefore, it is challenging to determine whether the observed metabolite alterations are a result of the disease or the result of a concurrent process. Given that diseased animals often exhibit reduced feed intake and lethargy, researchers should consider potential confounding effects when evaluating the strength of association between metabolite changes and disease.
Limitations of metabolomics
Although metabolomic techniques have evolved since its inception, there are a number of limitations that hinder its widespread use. At present, most metabolomic studies only identify a minority of metabolites in biological samples, reflecting the complexity of sample analysis, the presence of multiple adducts and isotopes for each species, and the difficulty of validating 100s of metabolites with suitable standards. In addition, some classes of metabolites are either difficult to detect using current instrumentation or are present below the level of detection. Incomplete coverage of key metabolic pathways can complicate the interpretation of data. Although the identification of unknown or poorly defined metabolites remains one of the biggest challenges for metabolomics, the detection of new or unanticipated metabolites also presents new opportunities for understanding disease processes and detecting new disease biomarkers.60,73
Another challenge in metabolomics is determining the significance or role of identified metabolites. Non-targeted metabolomic analyses generate enormous data sets that may contain vast amounts of clinically irrelevant information. The complexity of the metabolome and limitations in computational software technologies and algorithms make it challenging to extract relevant data. 19 Transforming these data into valid interpretations and conclusions requires an in-depth understanding of metabolic pathways and the interconnectivity of metabolites and biological systems. 30 In many cases, it is difficult to conclude how a change in metabolite steady-state levels translates to changes in metabolic fluxes through one or more associated pathways, although this is increasingly being addressed by coupling metabolomic approaches with stable isotope labeling. Another point to consider is that some metabolites are only biologically significant in the presence of other metabolite(s), which makes pattern-recognition analyses a particularly important component of metabolomics.
Additional challenges that need to be addressed are the development and optimization of sample collection and storage protocols for veterinary studies. Finally, metabolomic data alone is often insufficient in gaining a global understanding of physiologic processes; therefore, integrating multiple omics technologies (such as genomics, proteomics, or transcriptomics) may provide a more holistic perspective than any single omics field alone.10,53
Conclusion
Our literature search revealed that metabolomics has been applied widely in various animal science disciplines, but relatively few studies focused on spontaneous animal disease. Employing techniques such as NMR spectrometry and MS, metabolomics enables characterization and analysis of numerous metabolites in a biological sample. Metabolomics has immense potential in the study of spontaneous veterinary disease and may facilitate biomarker discovery and improve our knowledge of disease pathogenesis. Other opportunities include tracking response to treatment, pharmaceutical development, and toxicologic studies. Although there are relatively few metabolomics studies to date, we anticipate that many more will be performed in the future.
Supplemental Material
Supplemental_material – Supplemental material for Metabolomics in the study of spontaneous animal diseases
Supplemental material, Supplemental_material for Metabolomics in the study of spontaneous animal diseases by Helena Tran, Malcolm McConville and Panayiotis Loukopoulos in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors declared that they received no financial support for their research and/or authorship of this article. MJM is a NHMRC Principal Research Fellow.
Supplementary material
Supplementary material is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
