Abstract
With the cost of next-generation sequencing (NGS) decreasing, this technology is rapidly being integrated into the workflows of veterinary clinical and diagnostic laboratories nationwide. The mission of the U.S. Department of Agriculture–National Animal Health Laboratory Network (NAHLN) is in part to evaluate new technologies and develop standardized processes for deploying these technologies to network laboratories for improving detection and response to emerging and foreign animal diseases. Thus, in 2018, the NAHLN identified the integration of NGS into the network as a top priority. In order to assess the current state of preparedness across NAHLN laboratories and to identify which have the capability for performing NGS, a questionnaire was developed by the NAHLN Methods Technical Working Group and submitted to all NAHLN laboratories in December 2018. Thirty of 59 laboratories completed the questionnaire, of which 18 (60%) reported having some sequencing capability. Multiple sequencing platforms and reagents were identified, and limited standardized quality control parameters were reported. Our results confirm that NGS capacity is available within the NAHLN, but several gaps remain. Gaps include not having sufficient personnel trained in bioinformatics and data interpretation, lack of standardized methods and equipment, and maintenance of sufficient computing capacity to meet the growing demand for this technology.
The term
The U.S. Department of Agriculture’s National Animal Health Laboratory Network (NAHLN) was established in 2002, and is a nationally coordinated network and partnership of federal, state, and university-associated animal health laboratories. NAHLN’s mission is to provide animal health laboratory testing to detect emerging and foreign animal diseases (FADs) such as foot-and-mouth disease, African swine fever, and influenza in avian and swine species, among other NAHLN scope diseases. Importantly, the NAHLN also evaluates new technologies and develops standardized processes for deploying these technologies to network laboratories to improve FAD and emerging disease detection and response. The most recent NAHLN Strategic Plan identified the need to incorporate NGS into the NAHLN to support disease identification and surveillance activities.
Laboratories within the NAHLN are classified as levels 1–3 (L1–3), affiliate, or specialty, based on criteria established in the 2012 NAHLN Concept Paper. 12 Level 1–3 laboratories are publicly funded, state and university-associated animal health diagnostic laboratories, whereas affiliate laboratories are either federal or publicly funded laboratories whose mission falls outside the scope of animal agriculture. Specialty laboratories are private laboratories that have a specific testing capability otherwise unavailable within the NAHLN, needed to support APHIS surveillance and disease eradication efforts. Branch laboratories are defined in the NAHLN as multiple laboratories within a state or territory that share one or more of the following: administrative oversight, a quality system, or NAHLN funding mechanism. In December 2018, 59 laboratories and 2 National Veterinary Services Laboratories (NVSL; reference laboratories) constituted the NAHLN: 15 L1 laboratories; 6 L1 branch laboratories; 28 L2 laboratories; 2 L2 branch laboratories; 7 L3 laboratories; and 1 affiliate laboratory.
In 2018, a questionnaire was developed by a 7-member subcommittee of the NAHLN Methods Technical Working Group, with the goals of 1) determining the current state of NGS in the NAHLN laboratory community, 2) identifying the current use and capabilities of NAHLN laboratories performing NGS, and 3) identifying potential equipment platforms, bioinformatics tools, and pipelines that would allow standardization of NGS as a detection or disease response tool in the NAHLN. Thirty-two questions were formulated and divided into 3 general topic areas: reagents and equipment, analysis tools, and cost. Within the reagents and equipment topic, 14 questions targeted NGS technical procedures and addressed equipment, reagents, personnel, sample type(s), and current applications. The analysis tools section included 16 questions on quality control practices, software, data processing, analysis, storage, data sharing, and practices. The final 2 questions addressed fees and specific drivers behind NGS use in the laboratory. The estimated time for completion of the questionnaire was 15 min. Laboratories were given 2 wk from time of receipt to complete and return the questionnaire.
Responses were received from 30 NAHLN laboratories: 13 L1 laboratories, 15 L2 laboratories, and 2 L3 laboratories. No responses were received from any NAHLN branch or affiliate laboratories.
Overall, 18 responding NAHLN laboratories (60%) reported having sequencing capabilities, defined as either in-house or via a core facility (Table 1). This is similar to the 2016 survey of U.S. veterinary diagnostic laboratories, 2 in which 39% of surveyed laboratories reported the ability to conduct sequencing. Similar to the 2016 study, approximately one-third of the laboratories in the NAHLN still employ Sanger sequencing at some level.
Next-generation sequencing (NGS) capacity within the National Animal Health Laboratory Network as of December 2018.

This question did not distinguish between in-house or core facility sequencing capabilities.
Laboratories could select as many answers as applied.
Laboratories were asked for information about which NGS sequencing platforms they used. Thirteen laboratories reported access to and use of multiple sequencing platforms. Overall, Illumina and Oxford Nanopore Technologies were the 2 NGS manufacturers cited most frequently by all facilities (Table 2), followed by Thermo Fisher. Across the Illumina sequencing platforms, 2 laboratories reported using the HiSeq, 15 had a MiSeq, 6 used an iSeq, and 1 used a NextSeq. For the Oxford Nanopore sequencers, 2 laboratories reported using a MinION, and 2 had access to a PromethION. Two laboratories reported using the Thermo Fisher Ion Torrent sequencing platform, and 1 laboratory had access to a PacBio sequencing platform. These results align with recent surveys of both human public health laboratories 10 and veterinary diagnostic laboratories, 2 in which Illumina platforms were the most frequently employed.
Types of sequencing platforms used by NAHLN laboratories.
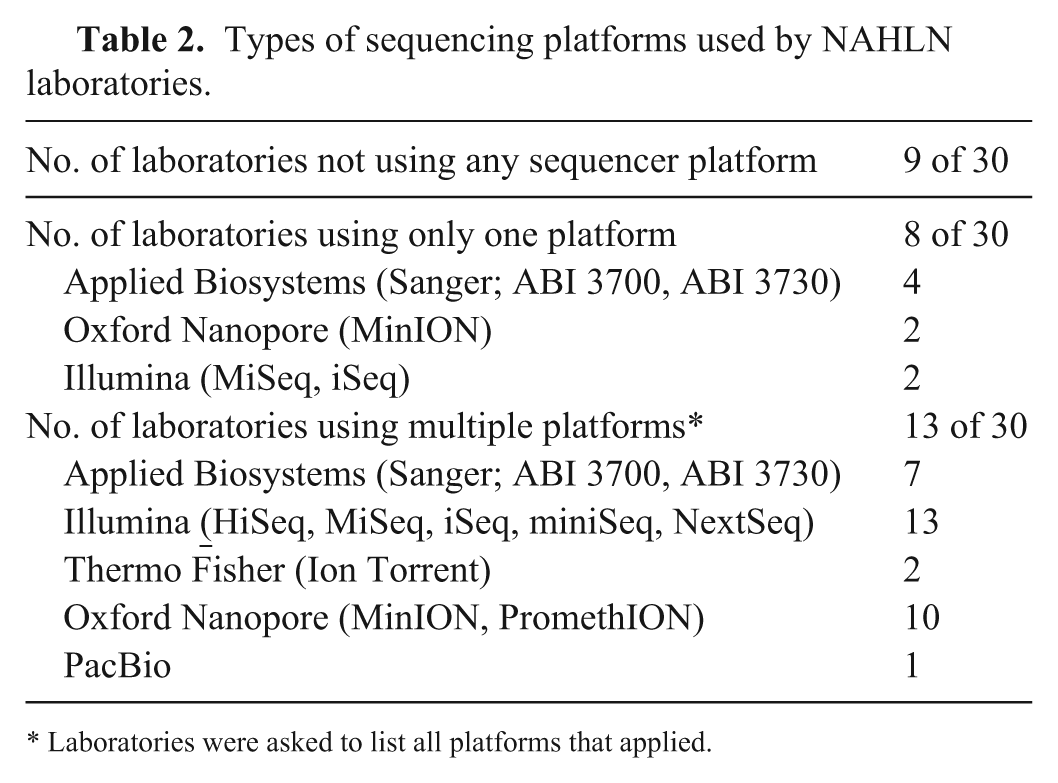
Laboratories were asked to list all platforms that applied.
This questionnaire also asked which commercial reagents were being used by laboratories given that standardization of protocols, including the use of specific reagents or kits, is a cornerstone of the NAHLN. The most common nucleic acid preparations employed were silica membrane spin-column technologies (10 of 15 laboratories responding), with significantly fewer laboratories using magnetic bead-particle technology (5 of 15 laboratories). Library preparation was most often performed using transposon-based fragmentation (14 of 22 responding laboratories), followed by enzymatic shearing (5 of 22 laboratories) and ligation-based fragmentation (3 of 22 laboratories). NAHLN laboratories were also queried about the manufacturers of sequencing kits that they used. Illumina kits were used most commonly (13 of 16 responding laboratories), followed by Oxford Nanopore (3 of 16 laboratories). This is unsurprising, given that the use of reagents from the same manufacturer as the sequencing platform ensures that workflows are optimized.
Information was requested on the volume of NGS testing performed by laboratories, and how NGS was being applied within each institution. On average, laboratories tested between 6 (minimum) and 55 (maximum) samples per run, with a range of 1–192. The number of laboratories performing metagenomics or WGS of individual bacterial or viral isolates was also requested. Eleven of 30 laboratories reported doing either targeted or shotgun metagenomics, and 12 indicated they were using NGS for WGS of bacterial and/or viral isolates.
Additional questions were asked to gain further understanding of applications. Overall, 16 laboratories use NGS technology for pathogen characterization, 11 for pathogen detection, 9 for virus discovery, 7 for outbreak situations or molecular epidemiology, and 4 for other purposes. Comparatively, bacterial identification, antimicrobial resistance profiling, and molecular epidemiology are frequent NGS applications for both human and veterinary clinical laboratories.1,2,4,9
Bioinformatics expertise can be a bottleneck for incorporating NGS into the clinical laboratory setting.1,10 Thus, our questionnaire queried personnel resources available to laboratories for conducting NGS. Most laboratories reported having at least some personnel trained in NGS techniques (Table 1). Bioinformatics expertise across all NAHLN laboratories was self-assessed at the beginner or intermediate level for 17 of the 22 laboratories responding to this question, suggesting that additional bioinformatics training may be needed to ensure standardized deployment of sequencing within the NAHLN. Although fewer laboratories reported having personnel dedicated for NGS bench work than bioinformatics or data analysis, our results may not reflect laboratory staff who are trained for routine molecular testing that also perform NGS bench work.
A founding principle of the NAHLN is for laboratories to operate within a quality management system, to provide confidence in test performance. Moreover, laboratories working under a formal quality management system such as ISO 17025 require extensive test validation to demonstrate that employed methods provide high-quality results. 11 Thus, our questionnaire also queried quality control parameters employed by laboratories performing NGS work. Although no clear consensus was identified regarding the use of standardized parameters to confirm NGS results, some themes emerged that may be explored for use within the NAHLN. These include evaluating raw data FastQ scores, assessing internal controls, and using third-party bioinformatics tools such as FastQC 15 or QUAST. 7 Additionally, the use of Clinical Laboratory Improvements Act guidelines may be considered given that these practices are being utilized by some public health laboratories. 8 The use of proficiency testing to help standardize NGS results was also cited by several reports as a critical aspect of overall quality management of NGS data.4,11,16
Standardized bioinformatics data analysis tools, or “pipelines,” are needed for normalizing NGS methods across NAHLN laboratories. Implementation of such pipelines is often impeded by the level of experience of personnel and types of tools available to them.5,10 The questionnaire responses showed differences in skill levels and tools used in NAHLN laboratories, with most using at least one bioinformatics analysis tool. Most laboratories reported having either beginner or intermediate level expertise; advanced expertise was reported by only 5 laboratories. Eight laboratories did not respond to this question. The tools used varied by reported skill level (Fig. 1). Laboratories at a beginner level were more likely to use commercial software tools; 7 laboratories using open-source software were at an intermediate level of expertise (Fig. 1). Comparatively, only 9 laboratories reported knowledge of scripting languages necessary for developing pipelines. Of these, 4 of 9 had intermediate, and another 4 of 9 had advanced, experience. Knowledge of additional skills such as Structured Query Language (SQL) were reported by 7 laboratories. Although these results are not unexpected, they present challenges for developing and integrating standardized bioinformatics analysis tools within the NAHLN.
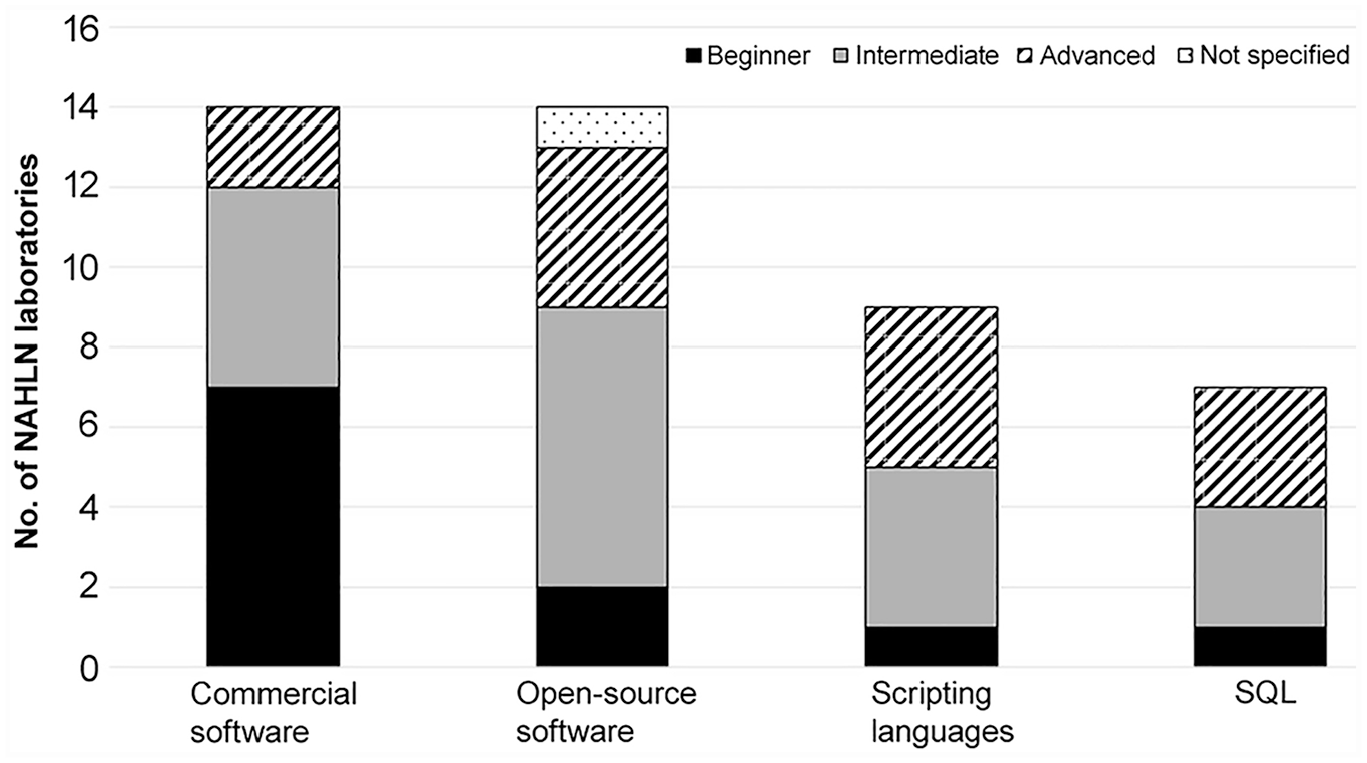
Next-generation sequencing bioinformatics data analysis tools used by NAHLN laboratories. The number of laboratories using each type of bioinformatics tool (commercial software, open-source software, scripting languages, or SQL) are further divided into bioinformatics skill levels, as reported by the laboratories.
Computational resources can also be a limiting factor for incorporating NGS into clinical laboratories. 10 Overall, 19 of 30 responding NAHLN laboratories indicated that they were satisfied with both their computer processing capacity and data analysis capabilities (Table 1). However, most of these laboratories also reported doing low levels of NGS sequencing, suggesting that, as laboratories find more applications for NGS, current computational resources may become limited.
The information gathered from our questionnaire indicates that many NAHLN laboratories have implemented NGS at some level; however, in order to fulfill the NAHLN’s mission of detecting foreign and emerging animal diseases, identifying additional resources for implementing this technology across all laboratories in the network will be needed. The gaps identified through our questionnaire include having sufficient personnel trained in bioinformatics and data interpretation, lack of standardized quality control methods and bioinformatics pipelines, and maintaining sufficient computing capacity to meet the growing demand for this technology. Nonetheless, the opportunities for improved testing that NGS affords to the veterinary community will help drive this forward and continue to make this a priority for implementation.
Footnotes
Acknowledgements
We thank all members of the NAHLN Methods Technical Working Group subcommittee on NGS for their input and critical review of the questionnaire. The findings and conclusions in this publication have not been formally disseminated by the USDA and should not be construed to represent any agency determination or policy.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. Mention of tradenames or commercial products is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
