Abstract
Porcine epidemic diarrhea, a disease caused by porcine epidemic diarrhea virus (PEDV), results in large economic losses to the global swine industry. To manage this disease effectively, it is essential to detect PEDV early and accurately. We developed a sensitive and accurate droplet digital PCR (ddPCR) assay to detect PEDV. The optimal primer-to-probe concentration and melting temperature were identified as 300:200 nM and 59.2°C, respectively. The specificity of the ddPCR assay was confirmed by negative test results for common swine pathogens. The detection limit for the ddPCR was 0.26 copies/μL, which is a 5.7-fold increase in sensitivity compared to that of real-time PCR (rtPCR). Both ddPCR and rtPCR assays exhibited good linearity, although ddPCR provided higher sensitivity for clinical detection compared to that of rtPCR. Our ddPCR methodology provides a promising tool for evaluating the PEDV viral load when used for clinical testing, particularly for detecting samples with low-copy viral loads.
Porcine epidemic diarrhea virus (PEDV; single-stranded RNA;
Many PEDV detection methods exist and include clinical observation, histologic observation, neutralization tests, immunofluorescence, and immunohistochemistry. These methods are time-consuming and unsuitable for large-scale clinical testing. 14 Molecular methods have become more commonly used for detecting PEDV. A reverse-transcription PCR (RT-PCR) assay has been developed to allow for detection of PEDV, but this method requires in-gel analysis of the PCR products. 12 Loop-mediated isothermal amplification is a novel DNA amplification method that has been developed11,14; however, this method produces false-positive results because of self-primer interactions. Real-time PCR (rtPCR) is used widely in clinical testing, and quantification using this method relies on a standard curve, quantification cycles (Cq), and the need to establish a Cq threshold line that can result in data bias. Therefore, a more accurate method needs to be established to detect PEDV infection.
Droplet PCR (dPCR) has come into use for precise quantification, given that this method can provide an absolute measurement of nucleic acid concentration without the use of standard curves. 1 Additionally, dPCR is more tolerant to inhibitors, and this allows for improved accuracy and precision of quantification of the target. 4 dPCR has been widely used in many areas such as food authentication, identification of genetically modified organisms, and clinical testing.4,6,17 We established a method for detection and quantification of PEDV using droplet digital PCR (ddPCR).
The primers and probes were designed for rtPCR and ddPCR assays (Oligo Primer Analysis software; Molecular Biology Insights), based on the nucleotide sequences of the
For the rtPCR system, the same primers and probe used for ddPCR were used (ABI QuantStudio 6 Flex real-time PCR system; Thermo Fisher Scientific). Real-time PCR reactions contained 2× AceQ qPCR probe master mix (Vazyme Biotech), 300 nmol/L of PEDV primers, 200 nmol/L of hydrolysis probes, 1 μL of template DNA, and sterile distilled water to provide a final volume of 20 μL. The reaction conditions for rtPCR included 95°C for 30 s, 35 cycles of 95°C for 5 s, and 60°C for 35 s.
For comparison, copy numbers were calculated for rtPCR based on the concentration calculated by standard curve using a calculator (http://scienceprimer.com/copy-number-calculator-for-realtime-pcr). Each sample was tested in triplicate to evaluate intra- and inter-assay repeatability. All statistical analyses were performed using the SPSS program (v.13.0; IBM).
To test if the ddPCR reaction system for PEDV could be improved, the primer-to-probe concentration and annealing temperature were optimized. 8 To select an optimal annealing temperature, PEDV complementary DNA (cDNA; 3.9 × 107 copies/μL) was annealed at temperatures of 55, 56, 57, 58.2, 59.2, and 60°C. Our results indicated that 59.2°C provided the optimal annealing temperature (Fig. 1). The fluorescence amplitude difference between the positive and the negative droplet clusters was the highest at this temperature. The primer concentration was optimized using PEDV cDNA (3.9 × 106 copies/μL for each reaction mixture). The optimal concentration ratio was 300:200 nM because this ratio of reagents resulted in optimal separation between positive and negative droplets (Fig. 2). When the concentration ratio was 400:400 nM or 900:250 nM, the rain effect (intermediate fluorescence of some droplets) was more pronounced and could affect the accuracy of the results. All results were analyzed using QuantaSoft software.
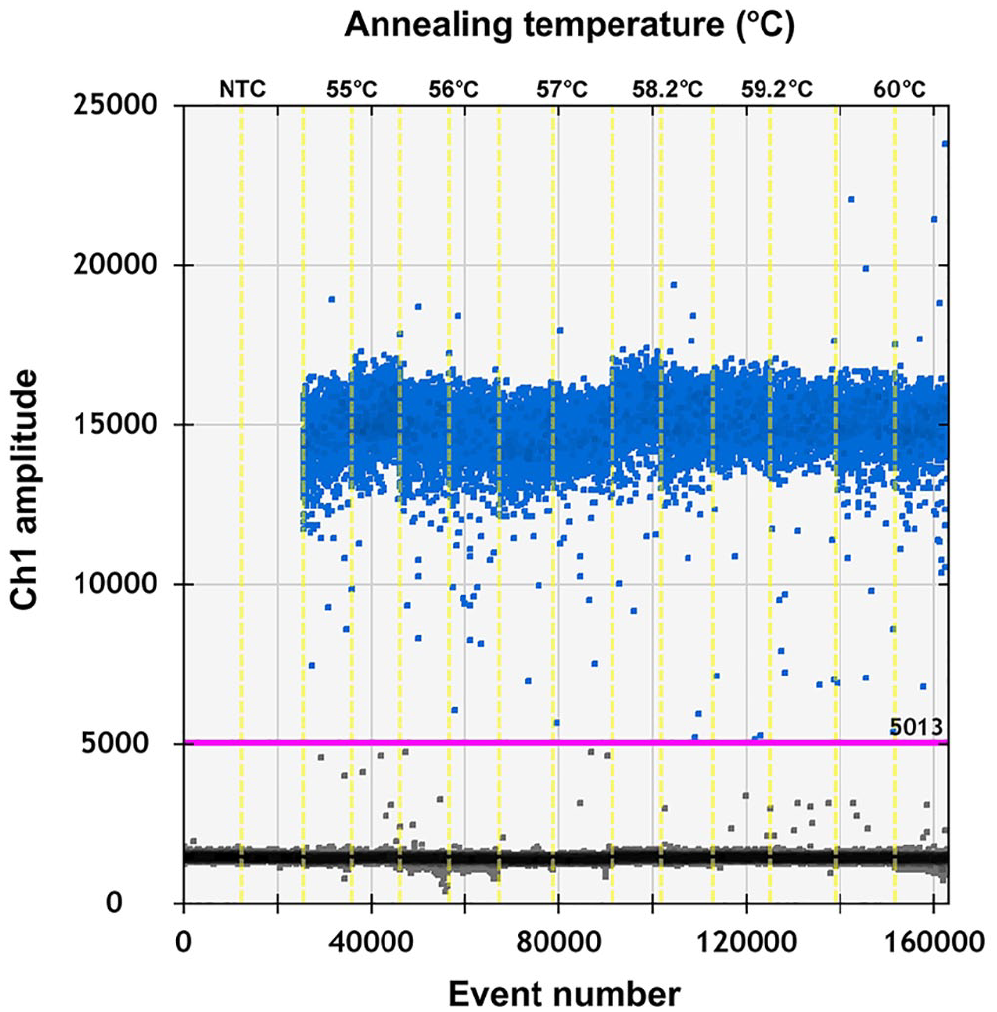
Influence of annealing temperature (55, 56, 57, 58.2, 59.2, and 60°C) on the porcine epidemic diarrhea virus droplet digital PCR system.
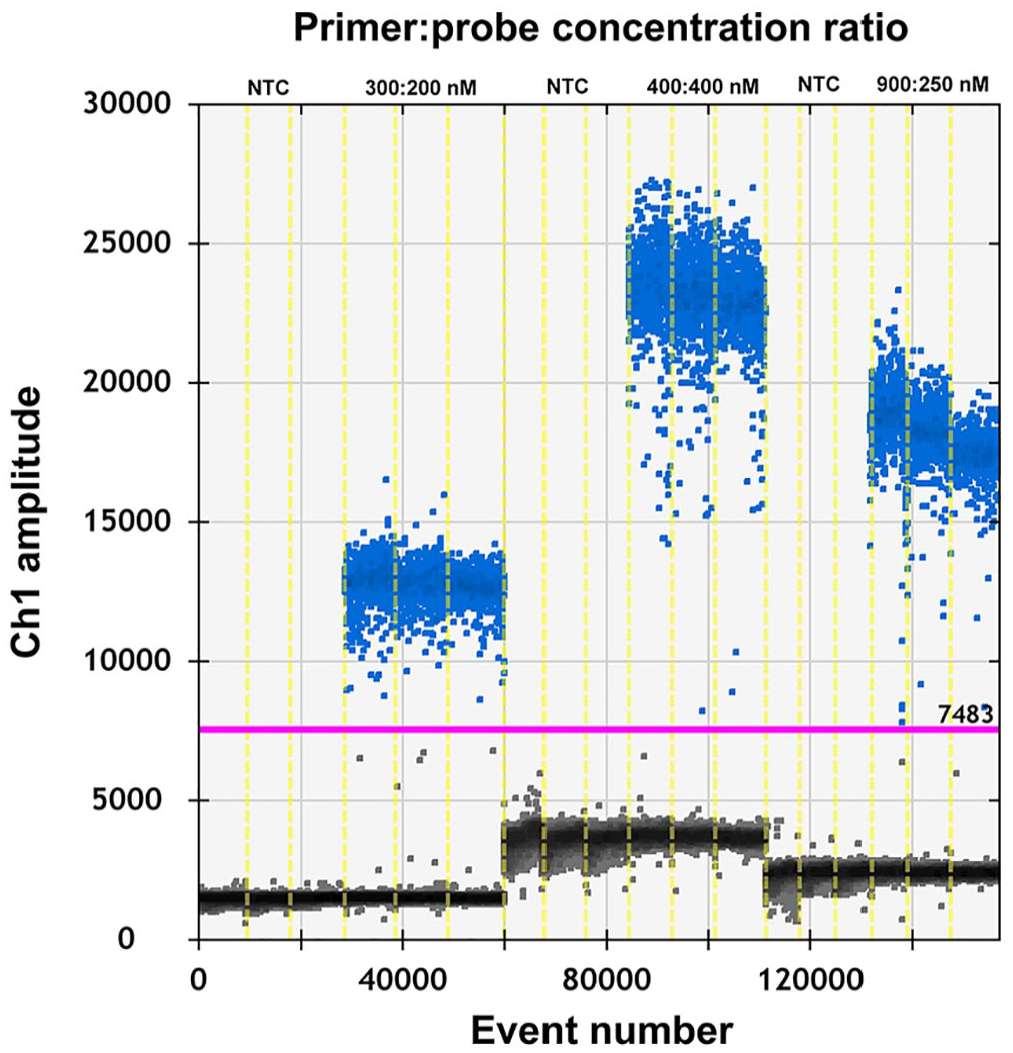
Influence of primer-to-probe concentration ratio (300:200 nM, 400:400 nM, and 900:250 nM) used in the porcine epidemic diarrhea virus droplet digital PCR system.
In specificity tests of the rtPCR and ddPCR assays, the nucleic acid extracts used as reaction templates included: porcine pseudorabies virus (
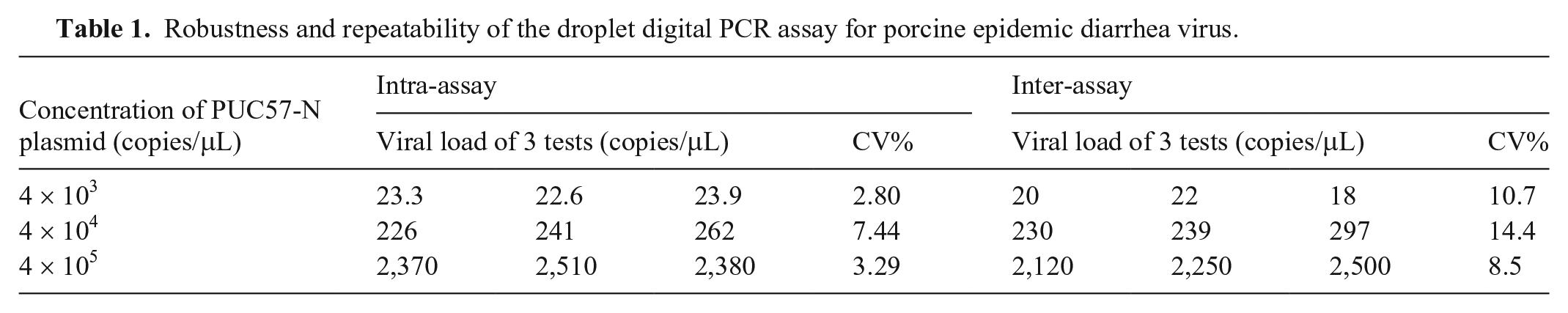
A total of 147 clinical specimens (porcine small intestine, feces, and serum) were obtained from 10 pig farms in China. Field strains and clinical samples were obtained from a commercial company (Yongshun Biological Pharmaceutical). Total RNA/DNA was extracted (RaPure viral RNA/DNA kit; Magen Technologies), and the RNA derived from PRRSV, CSFV, PEDV, PDCoV, TGEV, PRCV, SADS-CoV, and clinical specimens was reverse transcribed (Primer Script RT reagent kit; TaKaRa) according to the kit manufacturer’s instructions. In the repeatability tests of the ddPCR reaction system, 10-fold diluted PEDV plasmids (PUC57-N) were simultaneously tested in triplicate. The coefficients of variation (CV) and the concentrations (copies/μL) were calculated. These experiments revealed that the intra-assay CV for concentration was 2.8–7.44% and that the CV of the inter-assay was 8.5–14.4%, which indicated that the repeatability of the ddPCR reaction system was high (Table 1).
Robustness and repeatability of the droplet digital PCR assay for porcine epidemic diarrhea virus.
To compare the sensitivity, linearity, and quantification agreement between ddPCR and rtPCR, 10-fold diluted templates of plasmid PUC57-N (4 × 108–4 × 101 copies/μL) were used to conduct parallel tests (Table 2). The detection limit of ddPCR was 0.26 copies/μL; the detection limit of rtPCR was 1.47 copies/μL. Thus, the sensitivity of the ddPCR assay was 5.7 times higher than that of rtPCR. In this experiment, both rtPCR (
Comparison of real-time PCR (rtPCR) and droplet digital PCR (ddPCR) assays using serially diluted porcine epidemic diarrhea virus plasmids.
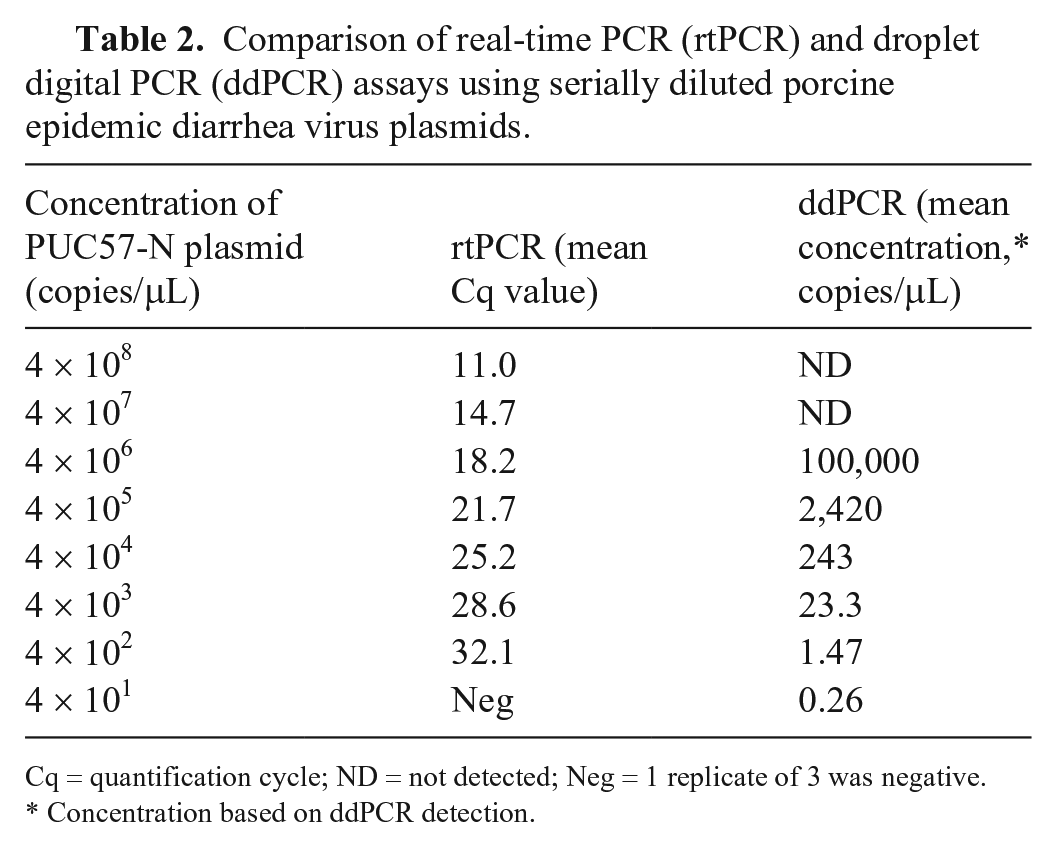
Cq = quantification cycle; ND = not detected; Neg = 1 replicate of 3 was negative.
Concentration based on ddPCR detection.
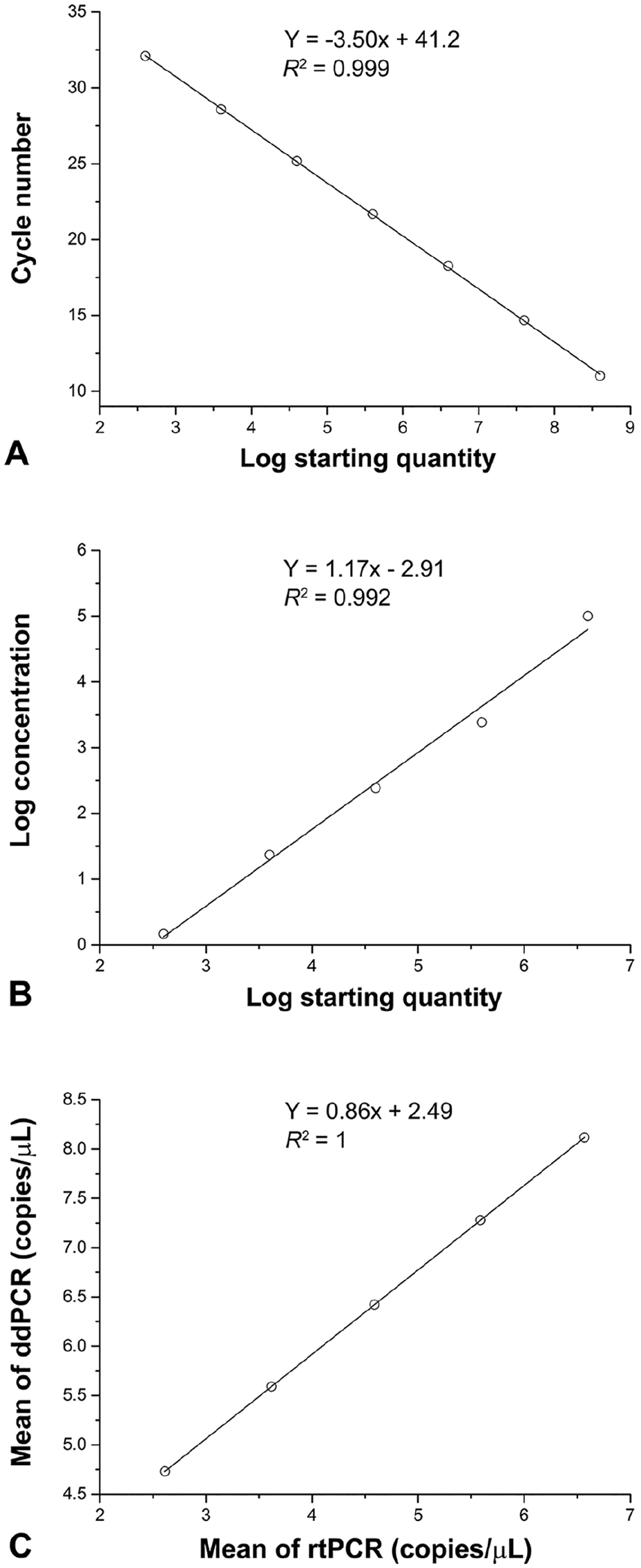
Sensitivity comparison between droplet digital PCR (ddPCR) and real-time PCR (rtPCR) using serially diluted porcine epidemic diarrhea virus (PEDV) plasmids.
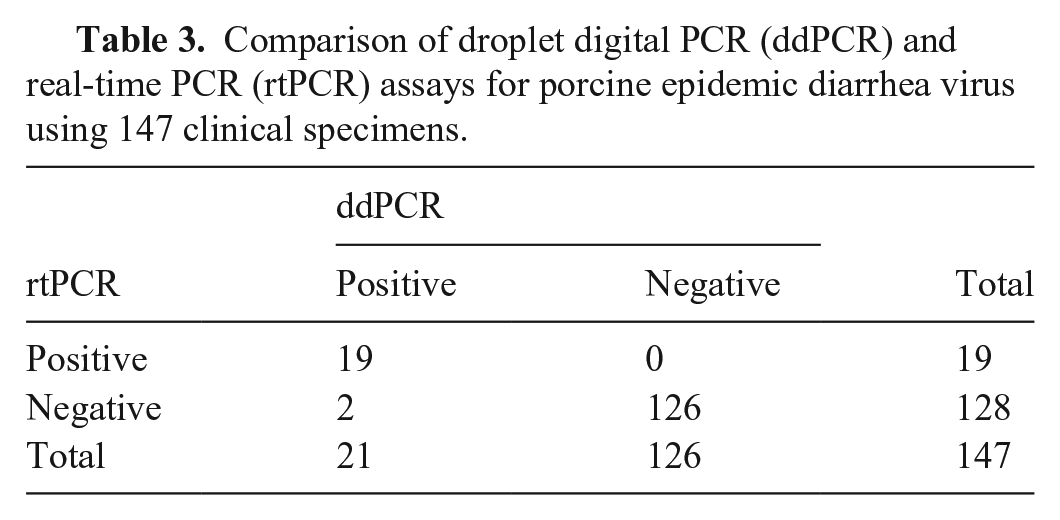
The 147 clinical specimens were tested simultaneously using ddPCR and rtPCR to compare the sensitivities of the assays. PEDV was detected with a positive rate of 14.3% (21 of 147) by ddPCR and 12.9% (19 of 147) by rtPCR. The overall coincidence rate was 98.6%, the kappa value was 0.942, and 95% CI was 0.863~1.022. Two of the samples identified as negative by rtPCR were positive by ddPCR (Table 3). The conflicting results detected by ddPCR and rtPCR were subsequently sequenced, and the sequencing results verified that these samples were positive for PEDV. The conflicting results from these samples could be a result of the higher sensitivity provided by the ddPCR assay compared to that of the rtPCR method.
Comparison of droplet digital PCR (ddPCR) and real-time PCR (rtPCR) assays for porcine epidemic diarrhea virus using 147 clinical specimens.
Our results indicated that the 2 methods had a high coincidence and that the ddPCR assay provided a more sensitive method for the precise quantification of PEDV compared to that of the rtPCR system, particularly for detecting lower concentrations of PEDV. 9 Additionally, the ddPCR system quantified the DNA in a highly reproducible manner without relying on a standard curve.2,8 However, based on analyzing 96 samples, the ddPCR took 3.3 times longer to complete than the rtPCR, the operation of ddPCR was more complicated than the rtPCR, and the overall cost (consumables and labor) of ddPCR was 2 times higher than rtPCR.2,18 In terms of high-throughput detection, the ddPCR instrument has only 2 fluorescence channels, testing a maximum of 4 target genes; the rtPCR instrument has 5 fluorescence channels, testing a maximum of 5 target genes.3,10 In general, the ddPCR system was verified as a sensitive and accurate method for detecting PEDV in clinical molecular virology.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest in regard to the research, authorship, and/or publication of this article.
Funding
This work was supported by the National Key Research and Development Program of China (grant 2016YFD0500600) and the Natural Science Foundation of Guangdong Province of China (grant 2017A030310175).
