Abstract
Evaluation of the diagnostic sensitivity (DSe) and specificity (DSp) of tests for infectious diseases in wild animals is challenging, and some of the limitations may affect compliance with the OIE-recommended test validation pathway. We conducted a methodologic review of test validation studies for OIE-listed diseases in wild mammals published between 2008 and 2017 and focused on study design, statistical analysis, and reporting of results. Most published papers addressed Mycobacterium bovis infection in one or more wildlife species. Our review revealed limitations or missing information about sampled animals, identification criteria for positive and negative samples (case definition), representativeness of source and target populations, and species in the study, as well as information identifying animals sampled for calculations of DSe and DSp as naturally infected captive, free-ranging, or experimentally challenged animals. The deficiencies may have reflected omissions in reporting rather than design flaws, although lack of random sampling might have induced bias in estimates of DSe and DSp. We used case studies of validation of tests for hemorrhagic diseases in deer and white-nose syndrome in hibernating bats to demonstrate approaches for validation when new pathogen serotypes or genotypes are detected and diagnostic algorithms are changed, and how purposes of tests evolve together with the evolution of the pathogen after identification. We describe potential benefits of experimental challenge studies for obtaining DSe and DSp estimates, methods to maintain sample integrity, and Bayesian latent class models for statistical analysis. We make recommendations for improvements in future studies of detection test accuracy in wild mammals.
Introduction
Improved understanding of the ecology of wildlife disease and the role of wildlife in the evolution and transmission of emerging infectious diseases is a core component of modern One Health initiatives. Much of this information is acquired through ecologic and epidemiologic studies in populations, and such studies require accurate and reproducible detection tests to provide information about disease or infection status of individuals and populations. Information gained in these studies is vital for guiding and monitoring health management policies including control of important animal and zoonotic diseases. The concept of a test being fit for a specified purpose (e.g., screening or confirmatory diagnosis) has been endorsed and promoted by the World Organisation for Animal Health (OIE) and others who work on test validation science.36,40
To enhance the harmonization of tests used in international trade, the OIE developed a standardized validation pathway 121 with 4 phases, representing analytical (stage 1), diagnostic (stage 2), among-laboratory reproducibility studies (stage 3), and deployment of tests to multiple laboratories (stage 4). Adherence to the OIE test validation pathway, development of reporting standards for test accuracy studies,34,35 and design standards in some animal species or taxa57,77 have improved the utility of studies by better defining source populations, use of structured experimental designs including sampling methods, and accounting for strengths and weaknesses of available tests for a specific testing purpose. The focus of these improvements, as outlined in the OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals, has been on infectious diseases of livestock and aquatic animals, but a specific chapter (2.2.7) 122 is directed at validation strategies in wild animals.
The large numbers of wildlife species, the diversity of pathogens that infect them, and the emergence of new diseases present unique challenges for test validation. Consequently, detection testing algorithms are often inadequately validated. Diseases in wildlife are frequently novel and can include multiple host species. Moreover, wildlife diseases, especially newly emerging ones, also are often studied inadequately. Compared with livestock, much less is known about the accuracy of detection tests when used in wildlife, and the question remains whether there are significant differences in test accuracy (e.g., diagnostic sensitivity [DSe] and specificity [DSp]) when used in different wild mammal species.
Often it is assumed that test protocols that have been validated for related species can be applied equally in wildlife, for example, bovine tuberculosis (Mycobacterium bovis). However, frequently there are flaws in the original test validation studies, with incomplete reporting of relevant animal- and population-level information. Access to adequate numbers and quality of specimens, information about infection status of populations, and limited knowledge of host–pathogen biology, pathogenesis, epidemiology, and population dynamics of naturally occurring infections are frequent limitations in wildlife. Also, inadequate funding may limit the scope and size of validation studies, especially when tests are used in multiple wild mammal species.
Our objectives are to: 1) review studies published between 2008 and 2017 on laboratory test validation with purposes of disease management and conservation in wild mammals, focusing on study design and statistical analysis methods; 2) document strengths, weaknesses, and design trade-offs of test accuracy studies in wild mammals including situations in which animal experiments may add value; and 3) make recommendations for improvements in design and statistical analysis, reporting of validation studies, and interpretation of test results. In addition, 2 coauthors (D.E. Stallknecht, D. Blehert) provide insights through case studies that demonstrate how new pathogens or viral serotypes or genotypes can be detected and testing algorithms changed, and how tests evolve for a new disease once the causative agent has been identified.
Methodologic review
Literature search, appraisal, and review process
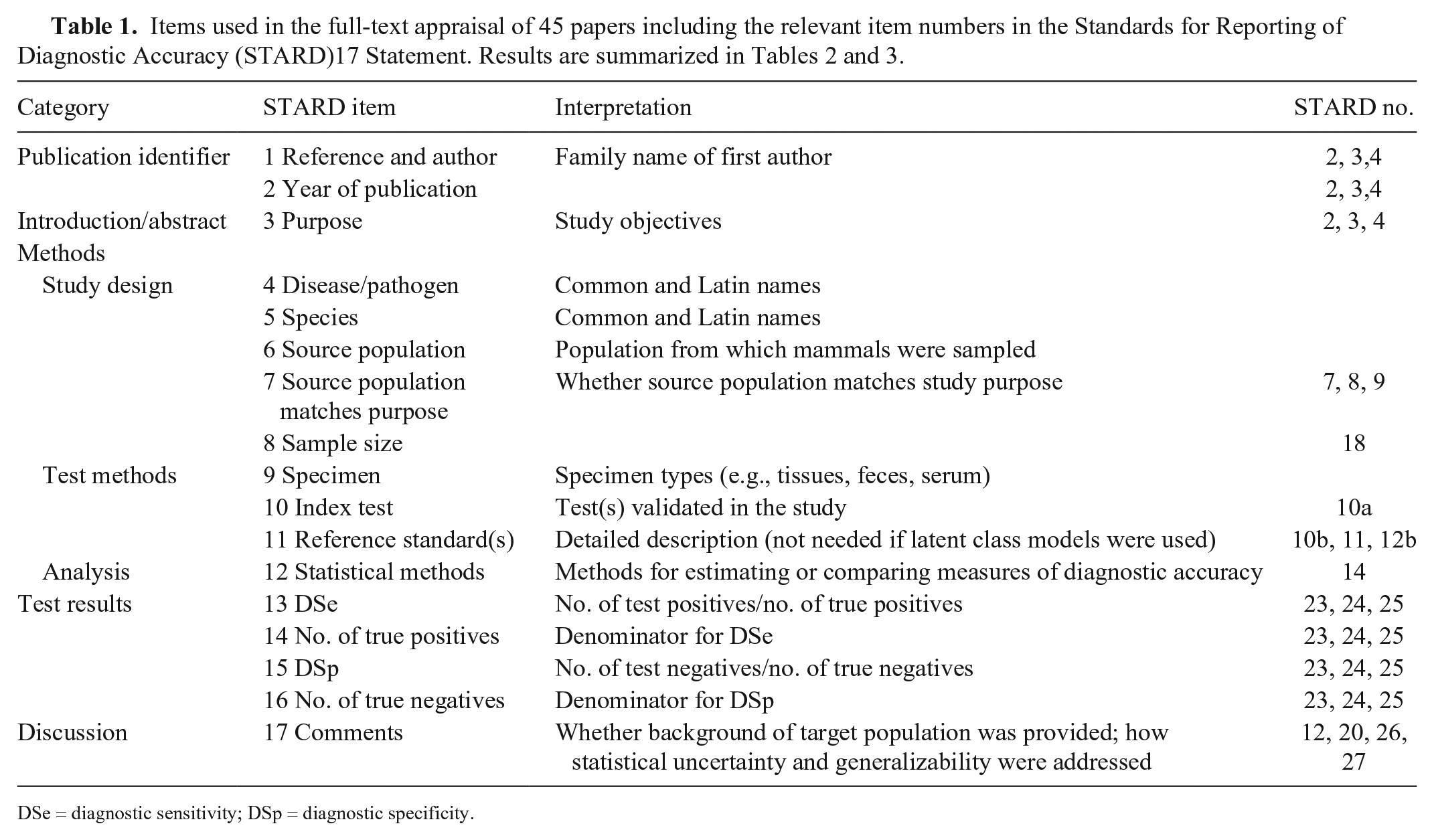
Our methodologic review was based on checklist items in the Preferred Reporting Items for a Systematic Review and Meta-analysis of Diagnostic Test Accuracy Studies (PRISMA-DTA). 70 Specifically, items in the Methods section of the list were used in the identification, screening, inclusion, and summary of relevant literature, including search, study selection, data collection, and synthesis of results. 70 We first conducted a systematic search to locate existing studies reporting test accuracy for OIE-listed diseases in wild mammals published between 2008 and 2017, using a standard set of search terms and disease names (Suppl. Fig. 1; Suppl. Data 1, 2). We characterized each study using a 17-item template (Table 1), which was based on modifications of the 27 items recommended in the Statement for Reporting Studies of Diagnostic Accuracy (STARD). 17
DSe = diagnostic sensitivity; DSp = diagnostic specificity.
We summarized results of full-text review based on whether the study was related to infection with Mycobacterium bovis (n = 15; Table 2) or pathogens other than M. bovis (n = 30; Table 3). We hypothesized that the long history of research involving development and validation of tests for M. bovis might contribute to substantial differences between these 2 groups. Historically, test accuracy studies for M. bovis focused on disease control and veterinary public health, including identification of infected cattle populations and removal of test-positive cattle. Since ~2010, there has been an increased focus on M. bovis in wild mammals because of their possible role as reservoir species as well as conservation concerns. In contrast, test validation studies for non–M. bovis diseases were mostly for outbreak investigation or cross-sectional studies involving prevalence estimation. Nineteen diseases were included in the non–M. bovis studies. Diseases with more than a single manuscript recording were African swine fever (n = 4), Sarcoptes scabiei (n = 3), Anaplasma spp. infection (n = 2), brucellosis (n = 2), chronic wasting disease (n = 2), Johne’s disease (n = 2), rabies (n = 2), and Toxoplasma gondii infection (n = 2; Table 3).
Summary of selected test validation studies (n = 15) for Mycobacterium bovis in free-range, captive, and farmed wild mammals published between 2008 and 2017.
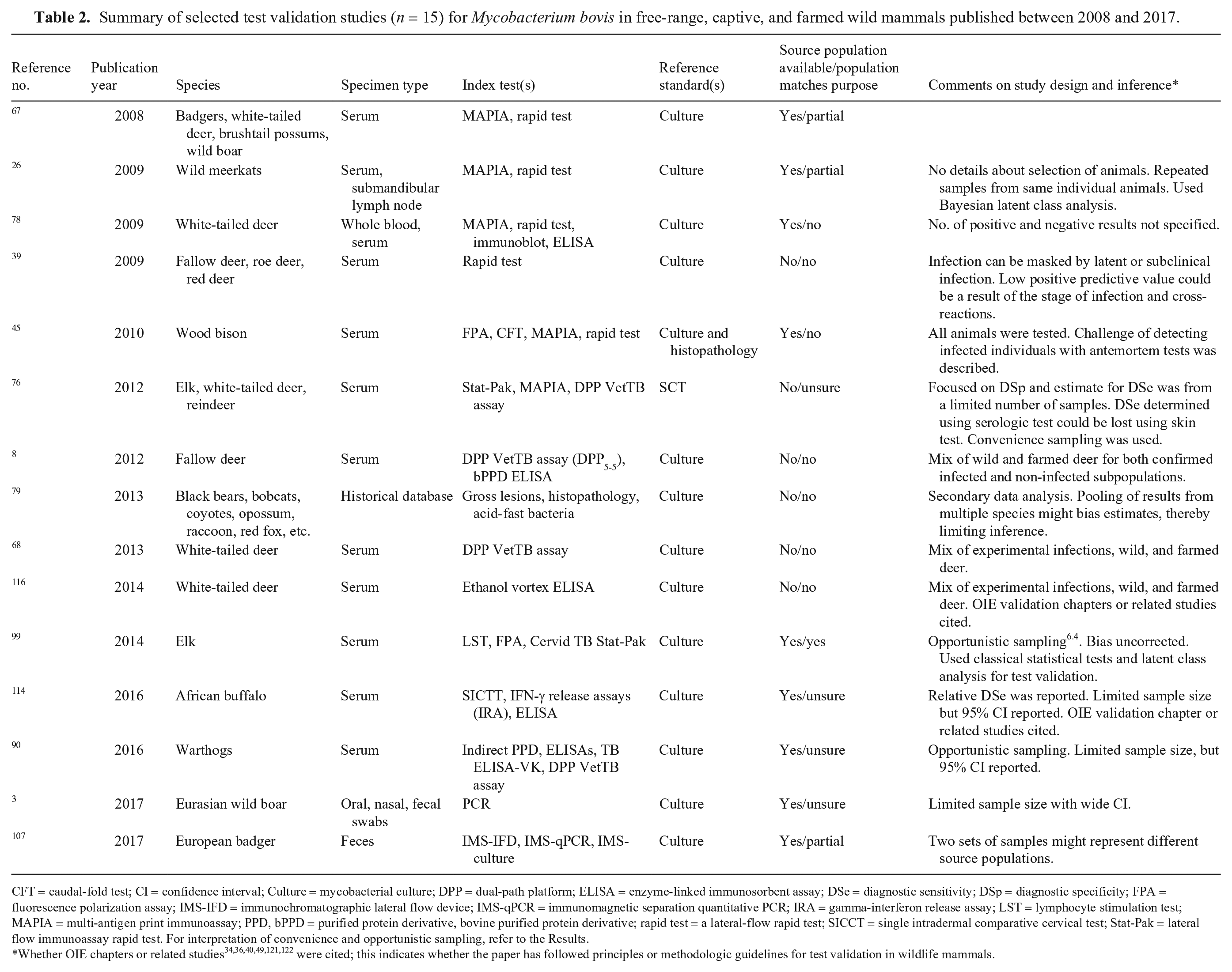
CFT = caudal-fold test; CI = confidence interval; Culture = mycobacterial culture; DPP = dual-path platform; ELISA = enzyme-linked immunosorbent assay; DSe = diagnostic sensitivity; DSp = diagnostic specificity; FPA = fluorescence polarization assay; IMS-IFD = immunochromatographic lateral flow device; IMS-qPCR = immunomagnetic separation quantitative PCR; IRA = gamma-interferon release assay; LST = lymphocyte stimulation test; MAPIA = multi-antigen print immunoassay; PPD, bPPD = purified protein derivative, bovine purified protein derivative; rapid test = a lateral-flow rapid test; SICCT = single intradermal comparative cervical test; Stat-Pak = lateral flow immunoassay rapid test. For interpretation of convenience and opportunistic sampling, refer to the Results.
Summary of selected test validation studies (n = 30) for diseases other than Mycobacterium bovis in free-range, captive, and farmed wild mammals published between 2008 and 2017.
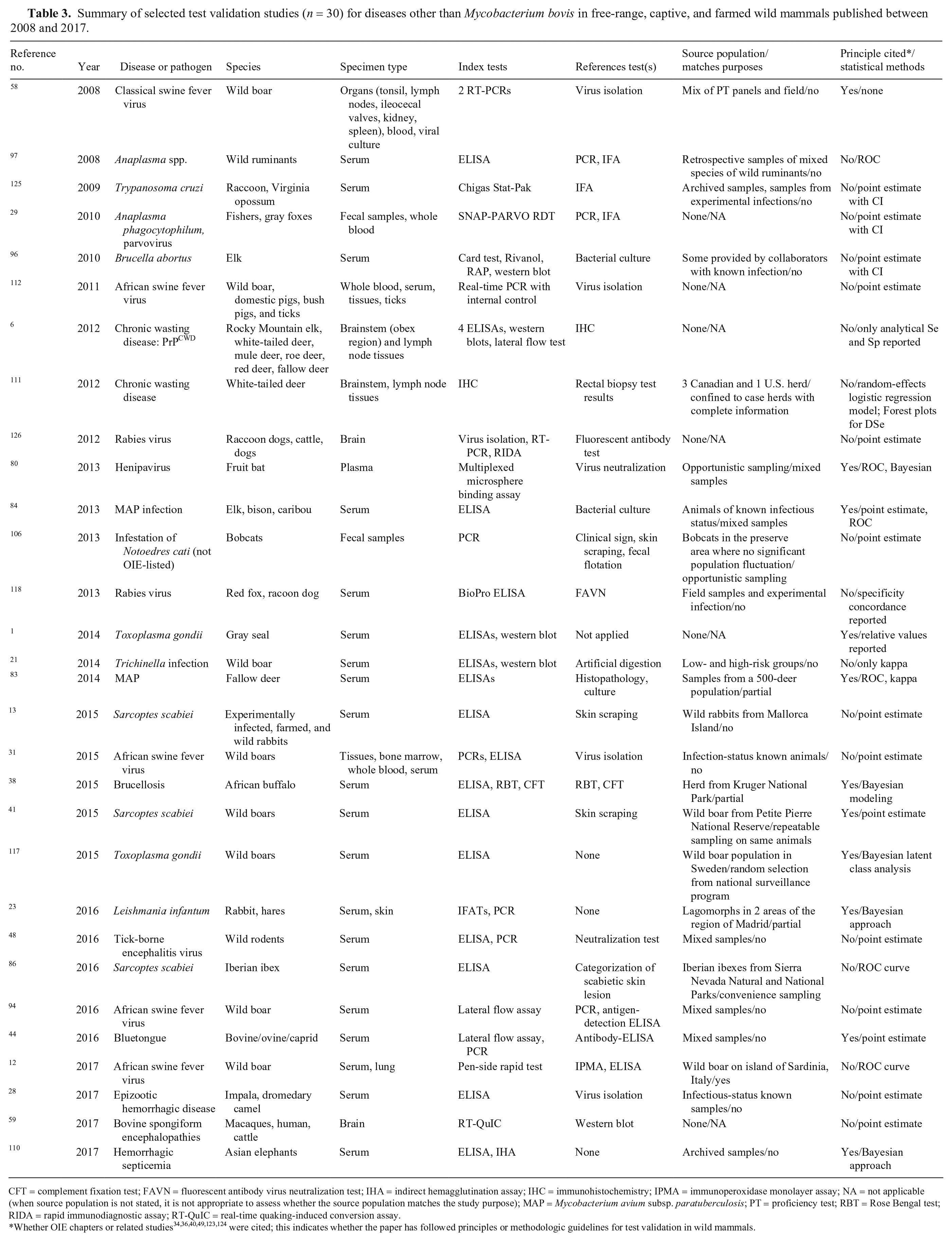
CFT = complement fixation test; FAVN = fluorescent antibody virus neutralization test; IHA = indirect hemagglutination assay; IHC = immunohistochemistry; IPMA = immunoperoxidase monolayer assay; NA = not applicable (when source population is not stated, it is not appropriate to assess whether the source population matches the study purpose); MAP = Mycobacterium avium subsp. paratuberculosis; PT = proficiency test; RBT = Rose Bengal test; RIDA = rapid immunodiagnostic assay; RT-QuIC = real-time quaking-induced conversion assay.
Limitations of published validation studies
The quality of test validation studies in wild mammals continues to improve, but many tests originally developed for humans and domestic animals have often not been validated adequately to include assessment of whether the test protocols or the designated cutoff value are appropriate for wild mammals. 120 The inconsistent performance of tests applied in phylogenetically related species or geographical locations remains a challenge for the management and control of infectious diseases in wild mammal populations. 30 We provide an overview of major limitations in the study design and results and inferences based on our full-text reading of papers, including source populations, sampling scheme, and interpretation of results.
OIE test validation chapters and related studies34,36,40,49,121,122 were cited in only 2 of 15 M. bovis papers and 10 of 30 non–M. bovis papers. These sources of information are best used during the design stage to ensure that the important elements that affect methodologic quality are not overlooked. Adherence to these principles will influence the quality of studies in terms of sampling, statistical methods, and inference.
Study design
Lack of description of the source population
Only 9 of 30 non–M. bovis studies provided information about the source population, compared with 9 of 15 M. bovis studies. For many test validation studies in wild mammals, the lack of source population information was not addressed, raising questions about their internal validity. When source population information was missing, it was not easy to assess whether studied animals could be used to make inferences about target populations (in which tests will be used in future) or whether sampled animals matched the target population defined in the study design. For example, calculation of DSe and DSp from clinically affected animals would be of minimal value if the assay was proposed to be used for certification of pathogen freedom in populations with no evidence of clinical diseases.
Lack of representativeness of the source population
Uncertainty about the temporal dynamics of wild mammal populations and the baseline frequency or threshold of infectious diseases in these populations influences the representativeness of the source population compared with the target population. Only a few non–M. bovis studies discussed the representativeness of source populations, including the cross-boundary collaboration between Canada and the United States to address chronic wasting disease (CWD) in white-tailed deer 111 and the validation of tests for Trichinella infections in wild boar in Sweden. 21
Additional examples of lack of representativeness between source and target populations can be also be found in other studies, 76 such as the use of a mix of experimental and field infection data, and for analysis of pooled data from wild and farmed mammals. In these instances, it is difficult to address how well the sampled animals matched the target population defined in the research question, which is a metric of internal validity of the study. 95 For test validation studies using multiple species, estimates of DSe and DSp were sometimes not reported separately for each species.28,97
Sampling scheme: sample selection, quality, and quantity
Sampling methods used in wild mammal populations are usually non-probability methods, such as convenience 76 or opportunistic sampling 90 because animals are often darted or trapped for sample acquisition. For example, the potential omission of follow-up samples for validation testing may result in over- or underestimation of DSe because of a lack of willingness of owners to allow an autopsy of M. bovis screening test–positive elk in a free-ranging herd. 76 The estimated DSp might not be representative for disease-free populations if DSp was only based on test-negative animals in infected populations. 90
Sufficient sample size can be a challenge for providing robust estimates of DSe and DSp. 19 Few studies justified their selected sample size, although most reported 95% confidence intervals (CIs) for DSe and DSp. Selection of an appropriate sample size for estimation of these parameters depends on their expected values, the desired error margin (e.g., ± 5%), and whether specimens are of known or unknown infection status. 121 Limited numbers of known positive samples are a common limitation that influences the precision of DSe, and this issue is not unique to wild mammals. 18 Logistics is a widely recognized factor for limited longitudinal follow-up of test positives, especially when multiple populations are studied. Repeated measurements to demonstrate temporal variation in serum antibody responses might not be realistic given the difficulty of resampling the same animals.
Sample quality often negatively impacts test accuracy for both direct detection assays (e.g., culture and PCR) and indirect assays (e.g., serologic and gamma-interferon tests). Storage, transportation, and handling of samples can also adversely affect test performance. Processing (freeze–thaw) and transportation were reported with possible decreased viability of the organism in bronchoalveolar lavage samples for culture of M. bovis from African lions (Panthera leo), especially those specimens collected from latently infected animals with a low initial count of bacilli. Low DSe for diagnosing individual wild mammals shedding M. bovis can happen with the combined conditions of small volume sample and temperature fluctuation during storage and transport. 26 Use of FTA cards to ensure long-term sample quality is described below in our study. None of the 45 papers provided data about effects of sample quality on DSe and DSp estimates or addressed methods to improve sample quality. Acceptance criteria for sample suitability facilitate a minimum level of quality and are a requirement in accredited laboratories (see also recommendations at the end of our study).
Results and inferences
Insufficient discussion of bias induced by heterogeneity of the source population
Given the long latent period of chronic pathogens, such as M. bovis, it is logistically difficult to track relevant information related to the population structure and temporal dynamics of infection, which is best characterized if animals can be individually identified, which is rarely done. 39 Furthermore, understanding basic population structure and dynamics is not possible in most cases, and if possible, would require a long-term commitment of substantial resources to allow repeated testing of surviving animals.
Insufficient consideration of impact of infection stage, disease progression, and cross-reacting infectious agents on test results
Data interpretation and reporting of DSe and DSp need to consider that spatial-temporal dynamics of susceptible and infected wildlife vary among geographic areas. 105 Furthermore, in addition to confounding factors such as age, sex, pregnancy, vaccination, stress, and nutritional status, test accuracy can be affected by the interaction between the pathogen of interest and the diversity of cross-reacting infectious agents in the host species. 14 None of the 15 M. bovis papers considered comorbidity issues that may lead to misclassification bias because of cross-reactions from coinfecting Mycobacterium spp. or reduced immunologic response associated with viral infections such as feline immunodeficiency virus (FIV). FIV coinfections negatively affect M. bovis test performance in African lions54,108 because of effects on both antibody and cell-mediated immune responses.82,88 Test validation results should be interpreted with respect to how missing information, including data on coinfections, may influence the proportion of positive and negative results.8,99
General progress in test validation for wild mammal diseases
Based on the full-text reading of the 45 manuscripts in our literature search, we describe how researchers have made progress embracing challenges in the validation of fit-for-purpose assays for wild mammal species, especially in the areas of study design and statistical analysis (Table 1).
Study design
Samples from reference animals of known infection status
Given the logistical challenges of collecting samples from wild mammals, strategies are needed to acquire sufficient numbers of positive and negative reference samples for estimation of DSe and DSp. This is a challenge for laboratory test evaluation in most wildlife diseases, especially for non–M. bovis studies. Utilization of samples of known infection (archived) status was done in 20% of studies for diseases including African swine fever, Brucella abortus, epizootic hemorrhagic disease, hemorrhagic septicemia, Johne’s disease, and Trypanosoma cruzi. However, only 3 M. bovis studies67,76,107 and 3 non–M. bovis studies84,96,110 used field samples for validation of novel or existing tests in addition to the use of samples from reference animals or experimentally infected wild mammals. The latter sample types often result in overestimation of DSe and DSp.
Statistical analysis
Most selected papers incorporated estimates of uncertainty in test performance. For example, studies of M. bovis reported CIs for both DSe and DSp, with a few exceptions because of the sample sizes available.8,76 Postmortem methods were shown to have DSp close to 100% with narrower CIs but low DSe with wide CIs (Suppl. Figs. 1, 2; Suppl. Data 1, 2). Some studies provided predictive values for a range of expected prevalence of infection in the target population.39,45,96,97,99,106
Bayesian latent class model (LCM) has been used increasingly for estimation of DSe and DSp for wild mammal diseases to account for imperfect reference tests. The use of LCM has been acknowledged by the OIE as a valid method for analysis of test accuracy data (stage 2 of the OIE pathway) in the context of assessing a test’s fitness for a defined purpose (e.g., surveillance, confirmation of infection or disease, and prevalence studies).121,122 Two M. bovis studies used LCM for the estimation of DSe and DSp with 95% probability intervals.26,99 Four non–M. bovis studies reported LCM as the statistical analysis method.23,38,80,117 A cost-effective ELISA for screening for brucellosis in African buffalo (Syncerus caffer) was developed with determination of cutoff value based on the use of a Bayesian LCM. 38 Similar methods were also applied in henipavirus assays in African bats. 80
The quality of papers using Bayesian LCM for the period from 2008 to 2017 is variable. For M. bovis, a 3-test in 1-population model was used for comparison of DSe and DSp of the caudal-fold test (CFT) and 2 serologic tests, a lateral flow rapid test (RT) and fluorescent polarization test (FPA), in 212 free-ranging wild wood bison (Bison bison athabascae), with the latter tests assumed to be conditionally dependent. 15 Given that this model is non-identifiable (i.e., there is no unique set of values for DSe and DSp given the joint test results), prior information on at least 2 parameters was necessary. The authors used prior information on 5 parameters, which should have ensured model identifiability, if at least 1 test was conditionally independent of the other 2 tests. Model results for the FPA indicated that the assay was less accurate than tossing a coin (i.e., worthless). OpenBUGS code was not provided in the paper, which would have allowed readers to identify possible reasons why such results were obtained. In another study, the accuracy of 2 new serologic tests was compared, multi-antigen print immunoassay (MAPIA) and a lateral-flow rapid test (RT), with results of culture of tracheal washes in 110 meerkats. 26 Three approaches for the statistical analysis, including a Bayesian LCM, were used. The joint counts for all test result combinations of positive and negative results were presented in Table 2 of their paper, and the Bayesian estimates were realistic, with culture being the most specific but least sensitive test. Code and convergence testing were not reported. In a follow-up study, the accuracy of mycobacterial culture, gamma-interferon assay, and a commercial lateral-flow immunoassay (Stat-Pak; DPP VetTB assay for cervids, DPP VetTB assay for elephants; Chembio Diagnostic Systems, Medford, NY) were compared in 305 badgers (Meles meles). 27 A Bayesian model of 3 conditionally independent tests in 1 population was used. Adequate sensitivity analysis was done, and sources of prior information (expert opinion and published studies) were well justified. However, a code was not provided, therefore it is unclear whether the values for use of tests in parallel were done within the Bayesian code and whether uncertainty in the estimates was captured. In conclusion, all 3 papers would have benefited from adherence to current STARD-BLCM (Bayesian LCM) standards. 56
Reporting and discussion
Use of multiple tests for interpretation of results
Combinations of tests that are run simultaneously or sequentially and interpreted in series or in parallel are used for detection of chronic bacterial infections caused by M. bovis, M. paratuberculosis, or Brucella abortus. Measuring different aspects of immunity (cellular- vs. antibody-based) for M. bovis will likely result in a higher combined DSe for detecting infection than 2 tests that measure the same immune response. 68 Even with antemortem tests, several researchers noted limitations of the validation results and challenges in detecting M. bovis–infected animals. Seroreactive animals were not culture-positive for M. bovis. 114 Animals testing negative by rapid tests (e.g., Stat-Pak) were only confirmed by autopsy if they underwent slaughter inspection. 8 Inconsistencies among serologic tests existed, and those negatives were shown to be culture or histopathology positive for tuberculosis, which indicated that those antemortem tests cannot definitively detect M. bovis infection but in return are likely to be highly specific. 45 In some studies, postmortem descriptions of sampled animals were also provided by incorporating other information (e.g., descriptions of gross lesions and histopathology) and antemortem test results (e.g., mycobacterial culture) from other animals in the source population. 68
Combinations of tests have been used for early detection of highly contagious diseases, such as African swine fever (ASF). Compared with other serologic tests, the immunoperoxidase test (IPT) has higher DSe for antibody detection earlier in the serologic response and can thus be used to test tissues of dead wild boars to determine whether the animals were exposed and/or infected with ASF virus (ASFV). The combination of an antibody test (IPT) and antigen-detection test (Universal Probe Library, UPL-PCR) was reported as the most reliable method for detecting ASFV during epidemics in Europe in 2014. 31 In another study, pen-side antigen and antibody tests used in combination were proposed for cost-effective monitoring of ASF. 12 ASFV-specific nucleic acid was found reliably using enrichment procedures in seized pork products in Australian airports (pers. comm., Jianning Wang, 9 Nov 2019, https://www.weeklytimesnow.com.au/agribusiness/african-swine-fever-found-in-pork-products-at-australian-airports-and-in-mail/news-story/d78eb71cdd6119a5e97d28d069549e39).
Another example is the detection of Mycobacterium avium subsp. paratuberculosis, the causative agent of paratuberculosis (Johne’s disease). Serum ELISAs for paratuberculosis detection in fallow deer (Dama dama) were evaluated, and their predictive values were interpreted together with the presence or absence of histologic lesions. 83 In the same study, ELISA-based results were compared pairwise to results from other testing techniques, including Ziehl–Neelsen staining, quantitative PCR, and bacterial culture. These multi-test comparisons for paratuberculosis indicated that pathogen exposure likely triggered antibody production without progressive infection. 83 Thus, ELISA can offer a low-cost screening tool in paratuberculosis surveillance to segregate fallow deer populations into infected and non-infected groups. 83
Improved reporting of pooled data from multiple species or multiple target populations
In an attempt to match the target population, results of testing from subpopulations were reported together with those of pooled data (i.e., mixed species samples from a source population with different infection histories).8,79 In addition to the pooled estimate, separate estimates for each species were provided to mitigate concern about differences in susceptibility, spectrum of disease, age and sex ratio, or phylogenetic distance from the domestic species for which the index test was developed. 80 For example, rapid tests developed for detecting Anaplasma phagocytophilum and parvovirus were validated and reported for fishers (Martes pennanti) and gray foxes (Urocyon cinereoargenteus), respectively. 125 Additional examples can be found for M. bovis papers, including serologic assays for detection of M. bovis in wild ruminants, 76 and ELISA for detection of the organism in cattle and deer. 116 Another study addressed the use of multiple species to establish cutoff values for free-ranging bison and caribou when no reference samples were available and infection status of the tested animals was not known. 84
Case studies: test validation of endemic diseases in wild mammals
Before proposing general strategies to address the aforementioned challenges, we describe specific problems of test validation for 2 endemic diseases in North America for purposes of disease management. These examples demonstrate that validation is an on-going process as pathogens change, testing technology evolves, and understanding of host–pathogen–environmental interactions in wild mammals improves. The second case of white-nose syndrome (WNS) in hibernating bats may set a precedent and establish an approach for broader harmonization and professionalization of laboratory testing for wildlife diseases.
Hemorrhagic disease in deer
Hemorrhagic disease (HD) is an important mortality factor affecting white-tailed deer (WTD) in North America and is caused by both bluetongue virus (BTV) and epizootic hemorrhagic disease virus (EHDV); mortality associated with BTV and EHDV infections has also been documented in other wild ungulate species, including mule deer, pronghorn antelope, and elk. 46 The interpretation of tests as applied to field-acquired HD is complicated by multiple and genetically related viruses (BTV and EHDV), multiple serotypes of viruses (at least 6 BTV and 3 EHDV serotypes in North America), diversity of ungulate species that are susceptible to this disease, and extreme variability in clinical outcome. Tests to detect BTV and EHDV include virus isolation in embryonated chicken eggs and cell culture, and various reverse-transcription PCR (RT-PCR) and RT-qPCR protocols.46,55,123 In addition, serologic techniques, including virus neutralization and ELISA, are available to detect prior or current exposure to the viruses and are most often used in surveillance or other epidemiologic studies. When these assays are assessed under controlled experimental conditions in young naïve WTD, it is not uncommon to achieve 100% estimates for both DSe and DSp depending on stage of infection. 85 However, currently, there are no reliable estimates of either DSe or DSp when applied to North American ungulates naturally infected with either BTV or EHDV.
Test performance of virus detection techniques may vary over the course of infection. Both BTV and EHDV can be associated with a long duration (up to 50 d) of viremia, but duration is highly variable between strains and possibly serotypes.46,85 Although estimates of BTV or EHDV RNA persistence are not available for wild ungulates, it is likely that this also is prolonged as found with BTV in cattle (up to 160 d) and sheep (up to 89 d). 53 Long-lasting naturally occurring reductions in antibody titer occur over time, and cross-reactive antibodies between EHDV and BTV serotypes may cause false-positive results. 53 The extent of this cross-reactivity may broaden with time as a result of exposure to multiple BTV and EHDV serotypes and possibly to a wider range of microorganisms and other nonspecific substances. 53 Hence, cross-reactivity may vary greatly between age cohorts within the study population, and geographically, between populations in BTV- and EHDV-endemic and epidemic areas.91,103,104
With cases submitted from the field, sample quality also is an important consideration given that most submissions are initiated by the public and occur in late summer and early fall when environmental temperatures are often high. To date, there are no estimates available related to potential loss of infectivity or loss of detectable RNA associated with sample quality. Similar issues also arise with sera collected from hunter-killed deer because blood collected from body cavities may be of questionable quality.
With HD, as with many wildlife diseases, DSe or DSp estimates are complicated by the complexity associated with multiple host species, multiple viruses, and highly variable clinical and epidemiologic patterns over the range of these hosts and viruses. The duration and levels of infectious virus, viral RNA, and antibodies will affect DSe and DSp, but unfortunately, current experimental data on the extent of virus and host variability is inadequate to accurately estimate potential effects on test accuracy.
White-nose syndrome in hibernating bats
WNS caused the most precipitous decline of a North American wild mammal population in recorded history. 4 Spread of the disease in the northeastern United States is described elsewhere. 7 The white-and-gray powdery substance observed on muzzles, wings, and tail membranes of affected bats had the gross appearance of a fungus, which was subsequently confirmed by direct microscopy and histopathology. 7 Culture conditions (initially 4°C) that replicated skin temperatures of hibernating bats were needed for successful isolation of the causative agent, a novel psychrophilic (cold-loving) fungus that was ultimately named Pseudogymnoascus destructans.37,72 The fungus has been shown to be the primary causative agent of WNS through fulfillment of Koch postulates. 64
The initial laboratory-based assessment of skin samples from 117 bats showed a strong association between the presence of P. destructans and histologic skin lesions. 7 Subsequently, to facilitate consistent analyses across multiple diagnostic laboratories, histologic criteria for WNS were established. 7 Also, to enable more rapid detection of the fungus, a conventional PCR-based method was developed to detect P. destructans based upon amplification of a 624-nt fragment of the internal transcribed spacer (ITS) region of the ribosomal RNA (rRNA) gene. 63 This test was validated using skin samples from bat carcasses that were previously identified as either positive (n = 48) or negative (n = 32) for WNS by histology and had 100% DSp and 96% DSe for detection of P. destructans. However, when this method was subsequently used to analyze environmental samples, such as soil or sediment collected from bat hibernation sites, the test had lower DSp than when previously validated using a subset of skin samples from infected bats that harbored a less complex background of fungal species and that was effectively enriched for P. destructans.
Sequence-based analyses of the PCR amplicons from environmental samples revealed a previously unknown diversity of closely related fungal species that cross-reacted with conventional PCR primers, 63 but those were collectively still distinguishable from P. destructans by single nucleotide polymorphisms following DNA sequence analysis. However, as genetic differences between P. destructans and closely related fungal species are often subtle, sequence-based comparisons must be conducted precisely and require a higher-level understanding of the nuances of GenBank and BLAST. Thus, efforts were undertaken to develop an improved PCR test, based on TaqMan procedures, which would more robustly distinguish P. destructans from near-neighbor fungal species without the need for follow-up sequence-based analyses of PCR amplicons.
To address this problem, it was necessary to characterize a less conserved molecular marker, the rRNA gene intergenic spacer (IGS) region, from P. destructans and closely related fungal species, to facilitate consistent identification of P. destructans without yielding false-positive results. 65 Following analysis of the resulting IGS region sequence data from 172 fungal isolates, a real-time TaqMan PCR test was developed and validated based on analysis of skin samples from bat carcasses that were previously identified as either positive (n = 49) or negative (n = 42) for WNS by histology. 73 This assay was shown to have perfect DSe and DSp for P. destructans, additionally exhibited no cross-reactivity when tested for amplification of nucleic acid extracts from 54 fungal isolates designated as near neighbors of P. destructans, and was considered the most accurate PCR-based assay for detecting the organism. With the availability of robust techniques to culture P. destructans, 63 to identify the presence of fungal DNA by real-time PCR 73 and to identify lesions diagnostic for WNS, 71 the final step was to develop a case definition for WNS based on test results combined with field observations to define infection by P. destructans and WNS (see case definitions for WNS). 113
The approach for developing test procedures in response to the emergence of WNS has been a multi-step process completed with necessary urgency. The steps have included identification and cultivation of the causative fungus, establishment of histologic criteria for WNS, development of a robust real-time PCR test, and creation of a case definition for infection by P. destructans and for WNS. Despite these advances, challenges remain. For example, there are now several PCR tests for P. destructans described in the peer-reviewed literature that differ in DSe and DSp,16,63,73,98 and which may be applied without discrimination by independent laboratories unfamiliar with strengths and weaknesses of each published test. This presents challenges for standardization or harmonization of testing outcomes from independent laboratories and increases the chances for reporting false-negative or false-positive results. Efforts are now underway to promote harmonization of molecular testing among a voluntary network of laboratories performing PCR-based assays for P. destructans.
Strategies at different stages of test validation to improve study design, sample integrity, test methods, and statistical approaches
In the following sections, we elaborate on how test validation has been improved for infectious diseases of wild mammals. We address the use of experimentally- versus field-infected animals, sampling methods for improving long-term sample quality, and latent-class statistical methods for analysis of test-accuracy data from populations of unknown infection or disease status.
Study design: trade-off between experimental infection studies and samples from naturally occurring infections
One of the challenges described in the introduction of our study is the use of samples from naturally occurring infections. Experimental infection studies may address this issue when infection is induced in the wildlife species of interest or in a suitable model species (e.g., ferrets), which is then monitored to acquire relevant clinical data and samples. Strengths of these studies include: 1) suitable controls that can be incorporated in the design; 2) frequent, even continuous, monitoring to support improved clinical information for collection of samples; and 3) the potential to determine incubation periods.
Most experimental infection studies are designed to understand pathogenesis and can be useful to determine the host–pathogen biology of a hypothetical host species. However, appropriate experimental designs also allow estimation of DSe in anticipation of, and in preparation for, pathogen incursion events. 5 The added value of including estimation of assay performance in infection studies allows for the ability to adequately sample animals to provide quality samples for evaluation of the accuracy of multiple tests, the opportunity to obtain standardized positive-control samples for future field studies, and the ability to evaluate DSe and DSp within a defined timeline.
Host–pathogen biological factors that may impact test accuracy, and that can be determined from infection studies, include the organ and tissue types that support infection (predilection sites), routes of pathogen shedding, kinetics and timing of the pathogen replication cycle, nature and magnitude of immune responses, and presence of related pathogens within the population. These data can also be determined from the study of natural field cases, although such studies may not have the same precision because of the lack of case numbers, quality of samples, and quality of related case information. Nevertheless, such field case information is critical to interpreting results from host–pathogen studies because it helps to verify that experimental infection studies are representative of natural infections. For further details about sample size considerations, readers are referred to epidemiology texts. 25
There are different ways in which data on host–pathogen biology can be acquired within animal infection studies. These include clinical monitoring, autopsy and histologic examination, and other pathology assays. Such data help to define each infection case, allowing characterization of norms and variations of the infected captive population, and comparison with natural cases. Test accuracy metrics, such as DSe and DSp, should be interpreted in light of these findings. For example, pteropid bats, which are known to be maintenance hosts for henipaviruses, usually shed only limited virus and develop low levels of antibody following experimental infection with henipaviruses, 43 and this may be the result of a limitation of sampling and monitoring protocols in these poorly understood host–pathogen systems. 100
However, experimental infection studies may also have limitations. The acquisition of wildlife is usually complex, resource-intensive, and sometimes unsafe for the animals and their handlers. The most favorable option is to raise animals in captivity within breeding colonies, therefore providing opportunities for acclimatization and training to the conditions and needs of the study. However, setting up these colonies is generally not practical, is always resource-intensive, and rarely can it be justified solely for purposes of studying a disease of interest. Additionally, captive-bred animals will not always be representative of the natural populations and the range of cross-reacting agents to which free-ranging animals may have been exposed.
On the other hand, using captive animals from the wild may be a short-term option, but needs to account for the stress and safety of capturing animals and in removing them from their natural environmental and social settings. Stresses related to capture and captivity may alter pathophysiologic responses11,24,89 and may, in turn, influence disease presentation and sample composition. This will affect their representation of the natural disease process. Humane endpoints in animal infection studies may exclude disease stages that occur under field conditions. Each species is unique in how it responds to capture and confinement, as well as sedation and anesthesia regimens and responses to handling by its caretakers. These species-specific characteristics need to be built into study designs to accurately generate representative data based on high-quality samples.
Methods to improve sample quality for molecular-based assays
Specimen quality depends on timely acquisition, adequate cold chain maintenance, and target stabilization. All these requirements are difficult to manage with wildlife studies. To address the needs for stabilizing and transporting biological samples, various studies have been performed, including the use of chemically treated filter paper cards (e.g., Whatman FTA cards; Flinders Technology Associates, GE Healthcare Bio-Sciences, Pittsburgh, PA).74,93 Samples collected onto cards are chemically stabilized, do not require immediate refrigeration or freezing, and the cards are lightweight and easily transported.22,75,87,101
Nucleic acids suitable for PCR testing have been recovered from wild mammals, including herpesvirus DNA from Asian elephants (Elephas maximus), 62 DNA from trypanosomes from non-human primates 2 and African zebu (Bos taurus indicus), 20 RNA from rabies virus isolates, 81 and DNA for PCR and restriction-fragment length polymorphism (RFLP)-based genetic profiling of wildlife in general. 101 In addition to specific research involving sampling from wildlife, the literature includes studies demonstrating stabilization of RNA and DNA from a range of pathogens that can infect wildlife.60,61 The quality of a sample preserved on FTA cards has been addressed in studies focused on genetic sequencing of the collected sample. Diagnostic and surveillance testing that require sequence analysis of the sample are more prone to failure given that quality (i.e., stabilization of the genetic materials, requirement for large segments of intact genome) in the sample diminishes compared to testing by conventional PCR and real-time PCR, which are more tolerant of shorter nucleic acid fragments in the test sample. 115 Multiple reports have demonstrated effective use of samples collected onto FTA cards for sequence analysis, including genome sequencing of opossums, rodents, and bats 9 ; microbiome sequencing from feces of spider monkeys (Ateles geoffroyi) 42 and dogs 102 ; genotyping of rotaviruses 109 ; and automated high-throughput genotyping of cattle using whole blood and nasal samples. 69
Statistical perspectives: use of LCMs for analysis of test accuracy data from wild mammals of unknown infection status
Bayesian LCMs are appealing to wildlife health specialists because they can be used to evaluate test accuracy in naturally occurring infectious diseases, the sample sizes are often small, and the use of non-invasive methods is greatly preferred for testing species of conservation concern especially when it is prohibited or perhaps unethical to autopsy and collect specimens to establish a definitive diagnosis. Inherent in a Bayesian analysis are the underlying model assumptions of conditional independence of tests, 33 constant sensitivity and specificity across populations, 38 and distinct prevalence when applied to scenarios with at least 2 tests and at least 2 populations 47 and further developed in a Bayesian framework. 50 For the LCM with 3 tests in a single population, only the assumption of conditional independence is needed for model identifiability. This assumption is most likely correct when the assays target different analytes (e.g., virus isolation, PCR, and serology). A crucial aspect of any Bayesian analysis is proper justification of the prior distributions provided by subject-matter experts. Beta priors, which are commonly used for DSe and DSp, are based on expert guesses of the most likely [modal] value (e.g., DSp = 0.99) and a lower limit (e.g., DSp = 0.95) that the expert is 95% or 99% sure that the true value exceeds. Use of priors centered on the correct parameter values, but which are diffuse enough to not be overly influential, is especially important in non-identifiable models 52 because prior information about parameters may substantially influence posterior inferences about DSe and DSp. A sensitivity analysis using different priors is considered an essential prerequisite for reporting of a BLCM. 56 Construction of biologically reasonable Bayesian models that are also statistically sound requires input from biostatisticians, epidemiologists, disease specialists, laboratory diagnosticians, and others who use the tests for clinical decision-making, routine diagnosis, disease surveillance, or research. Bayesian LCMs are readily implemented in freeware platforms such as OpenBUGS 66 and R (https://www.r-project.org/), and their use is facilitated because authors of many studies share code through websites (e.g., https://cadms.vetmed.ucdavis.edu/diagnostic-tests).
For study planning purposes, the most common design is to have 2 geographically distinct and unconnected populations in which 2 tests are applied to all sampled animals. Sample sizes depend primarily on estimated disease prevalence in respective populations, whether prior information is available for model parameters, and how close DSe and DSp are to 100%. These factors are described in other papers.36,51 In many situations with wild mammal studies, logistical issues constrain the maximum sample size that is obtainable.
Recommendations for future test validation studies and interpretation of test results
Design including statistical analysis
There is no single set of design guidelines for test accuracy studies in wild mammals, but purpose and case definition usually define whether the study design will be based on samples and test results from naturally occurring disease or infection events in populations, samples from animals of known infection status (infected or not), samples from experimental challenge studies, or combinations of 2 or more sources. Each sample type has advantages and disadvantages, and the design needs to balance the potential actions of wildlife health management and practical issues, such as costs and logistics.32,49
For chronic and progressive infections in wild mammals, several components should be considered specifically at the initial planning stage: pathogenesis of the infectious disease in the targeted population, analytical principles of the tests under evaluation, alternative tests and their strengths and limitations, and context (e.g., target population) in which the validation data will be used. 77 Internal validity (freedom from bias) and external validity (generalizability) of study results are important methodologic qualities that warrant careful consideration by test users. 119
The design of the study informs choice of the method of statistical analysis of test results. Guidance on the latter is available elsewhere, 124 and additional information on LCMs is included earlier in our study and elsewhere.36,51 Statistical advice, including the calculation of appropriate sample size, is most useful if done during the planning phase rather than after the data are collected. Although DSe and DSp are the most common metrics of test accuracy, consideration can also be given to alternative measures (e.g., likelihood ratios) and cutoff-independent measures of accuracy (e.g., area under the ROC curve).
Reporting
The reporting of test accuracy studies in wild mammals should follow STARD guidelines 10 given that the only modifications for animals are Standards for Reporting of Animal Diagnostic Accuracy Studies (STRADAS)-aquatic for aquatic animals 35 and STRADAS-paratuberculosis, 34 which is targeted to a single disease of wild and captive ruminants. STARD assumes that there is a near-perfect reference standard (based on one or more tests) to which results of the test(s) under evaluation are compared. As previously indicated, the assumption of a perfect reference standard is rarely correct, and hence, LCMs are appropriate methods of analysis for many test accuracy studies. STARD-BLCM reporting standards have been published to address key reporting issues in diagnostic accuracy studies that are specific to Bayesian analysis regardless of the species in which tests were evaluated. 56
Interpretation
We make the following recommendations about the interpretation of laboratory test results from validation studies and application of the tests in future studies:
Have clearly defined questions (purposes and case definition), use the appropriate tests to answer them, and be aware of their limitations. For example, the best serologic tests would be inappropriate to determine the cause of mortality but may be very appropriate for evaluation of prior exposure to pathogens.
Clearly define how tests will be interpreted and justify any adjustments. For example, it may be valid to reduce a minimum positive titer threshold for a serologic test if sampling a population well after a disease event or if the intention is to determine the longevity of the antibody response. Analyzing the distribution of data and a graphical display in a histogram or dot diagram may be helpful to compare patterns of recent, mid- and long-term infection in populations.
Use multiple testing approaches. For example, detection of BTV and EHDV serotypes can be much more reliable if supported by spatially and temporally matched virologic data even if collected independently of the study. Likewise, the combination of virus isolation, RT-PCR, and next-generation sequencing may provide a broader perspective related to diagnosing a cause of death or clinical outcome.
Pay attention to sample quality, develop guidelines, and adopt easily standardized sample collection and preservation technologies, such as FTA cards. Quality control guidelines may be subjective, such as discarding samples with obvious hemolysis or contamination, or could be very specific, such as maintaining a well-defined cold chain. In either case, these considerations are vital to the diagnostic process and should be included in a standard operating procedure. It also may be of value to future researchers to provide some data and analyses supporting potential guidelines for quality assurance.
The study population and how it was sampled needs to be defined in detail. This is especially true if seasonal transmission patterns are not considered.
Validate and scrutinize laboratory tests, if possible. Simple experimental infections may be possible, or multiple samples with known positive or negative status may be available for inclusion as controls. It also may be helpful to verify results with additional repeat testing and to validate testing using different test platforms, by sequencing, or by testing through independent laboratories.
Be cautious with all interpretations. Low detection frequencies of positive results especially should be approached with caution and should be interpreted with the knowledge that positive predictive values will be much reduced at low prevalence. Additionally, DSp estimates do not account for unknown infective agents to which a wild animal may be exposed. In North America, for example, there are numerous exotic BTV and EHDV that may not be currently represented in existing test formats such as virus neutralization, which would lead to false-negative results. 92
Supplemental Material
Supplemental_material – Supplemental material for Validation of laboratory tests for infectious diseases in wild mammals: review and recommendations
Supplemental material, Supplemental_material for Validation of laboratory tests for infectious diseases in wild mammals: review and recommendations by Beibei Jia, Axel Colling, David E. Stallknecht, David Blehert, John Bingham, Beate Crossley, Debbie Eagles and Ian A. Gardner in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
The use of trade, product, or firm names is for descriptive purposes only and does not imply endorsement by the U.S. government.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was funded in part through the OIE Collaborating Centre for Diagnostic Test Validation Science in the Asia-Pacific Region.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

