Abstract
Bovine herpesvirus 1 (BoHV-1) causes several clinical syndromes in cattle worldwide. There are 3 subtypes of BoHV-1: 1.1, 1.2a, and 1.2b. Several molecular methods are commonly used in the detection and characterization of BoHV-1. Among them, restriction endonuclease analysis (REA) and single-nucleotide polymorphism (SNP) analysis of the complete viral genome allow classification of BoHV-1 into different subtypes. However, developing countries need simpler and cheaper screening assays for routine testing. We designed a standard multiplex PCR followed by a REA assay allowing straightforward subclassification of all BoHV-1 isolates tested into 1.1, 1.2a, and 1.2b subtypes based on the analysis of fragment length polymorphism. Our standard multiplex PCR-REA was used to analyze 33 field strains of BoHV-1 isolated from various tissues. The assay confirmed the subtype identified previously by REA. In addition, non-polymorphic or undigested fragments were sequenced in order to confirm the mutation affecting the RE HindIII site. Our PCR-REA method is an affordable and rapid test that will subtype all BoHV-1 strains.
Bovine herpesvirus 1 (BoHV-1; Varicellovirus, Bovine alphaherpesvirus 1) is an important cattle pathogen causing infectious bovine rhinotracheitis (IBR), infectious pustular vulvovaginitis/balanoposthitis (IPV/IPB), and abortions, as well as systemic illness in young calves.13,29 Like other herpesviruses, BoHV-1 is able to establish life-long latent infection after primary infection, which can be reactivated under natural or experimental conditions.27,28 All BoHV-1 strains isolated belong to one viral species and are classified into 3 subtypes: 1.1, 1.2a, and 1.2b. 25
Several countries, and some counties within countries, are recognized as IBR-free. The European Commission has approved eradication programs in some countries and regions to which the additional guarantees for IBR apply (2011/674/EU. Available at: http://data.europa.eu/eli/dec_impl/2011/674/oj). In addition, IBR has become a limiting factor in livestock trade (64/432/EEC Article 9. Available at: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=DD:I:1963-1964:TOC:EN). 29 BoHV-1 is the cause of several clinical syndromes in cattle worldwide, resulting in its classification as an OIE-listed disease agent (https://www.oie.int/en/animal-health-in-the-world/oie-listed-diseases-2019/).
In Argentina, the first outbreak of bovine herpesviral disease was described in 1982, from which both BoHV-1 and BoHV-5b reference strain A663 were isolated. BoHV-5b was proposed to be a recombinant of BoHV-5 and -1. 20 Our group recently reported the circulation of all subtypes of BoHV-1 in Argentina. 21 Although most BoHV-1.1 strains have been isolated from respiratory tract diseases or abortion cases, and BoHV-1.2 strains from genital organ lesions, the only reliable means of typing the isolates is viral DNA analysis by restriction endonuclease fingerprinting.23 –25 Several assays are available to detect BoHV-1, including immunoassays using monoclonal antibodies,14,23,32 standard PCR,4,10,30 nested PCR, 6 multiplex PCR,2,7,25 and real-time PCR. 22 Some of these assays allow differentiation of BoHV-1 from BoHV-5.
The HindIII restriction enzyme analysis REA (gold standard method) of the complete virus genome has been widely used to subtype BoHV-1 isolates. 8 It proved to be particularly useful for the differentiation of various ruminant alphaherpesviruses antigenically related to BoHV-1, which show distinct DNA fingerprints.3,15 However, it is a laborious and time-consuming assay.
Random amplification of polymorphic DNA (RAPD), 1 PCR sequencing assays, 9 complete genome sequencing, and single-nucleotide polymorphism pattern analysis 10 are used to genotype BoHV. These methods, although possessing high accuracy, are expensive for routine use in developing countries. 5 Therefore, our aim was to develop an economical, quick, and simple molecular technique that allows the detection and differentiation of all BoHV-1 subtypes in developing countries and areas in which BoHV-1 causes endemic infections.11,17,21
We obtained BoHV-1 field isolates, identified by PCR, 7 from samples of naturally infected animals, collected across Argentina (Table 1). All viruses were propagated in Madin-Darby bovine kidney (MDBK) cells grown in Eagle minimal essential medium supplemented with 2% fetal bovine serum. 19
Argentinean bovine herpesvirus 1 field isolates and reference strains grouped into subtypes according to the REA profile and multiplex PCR-REA.
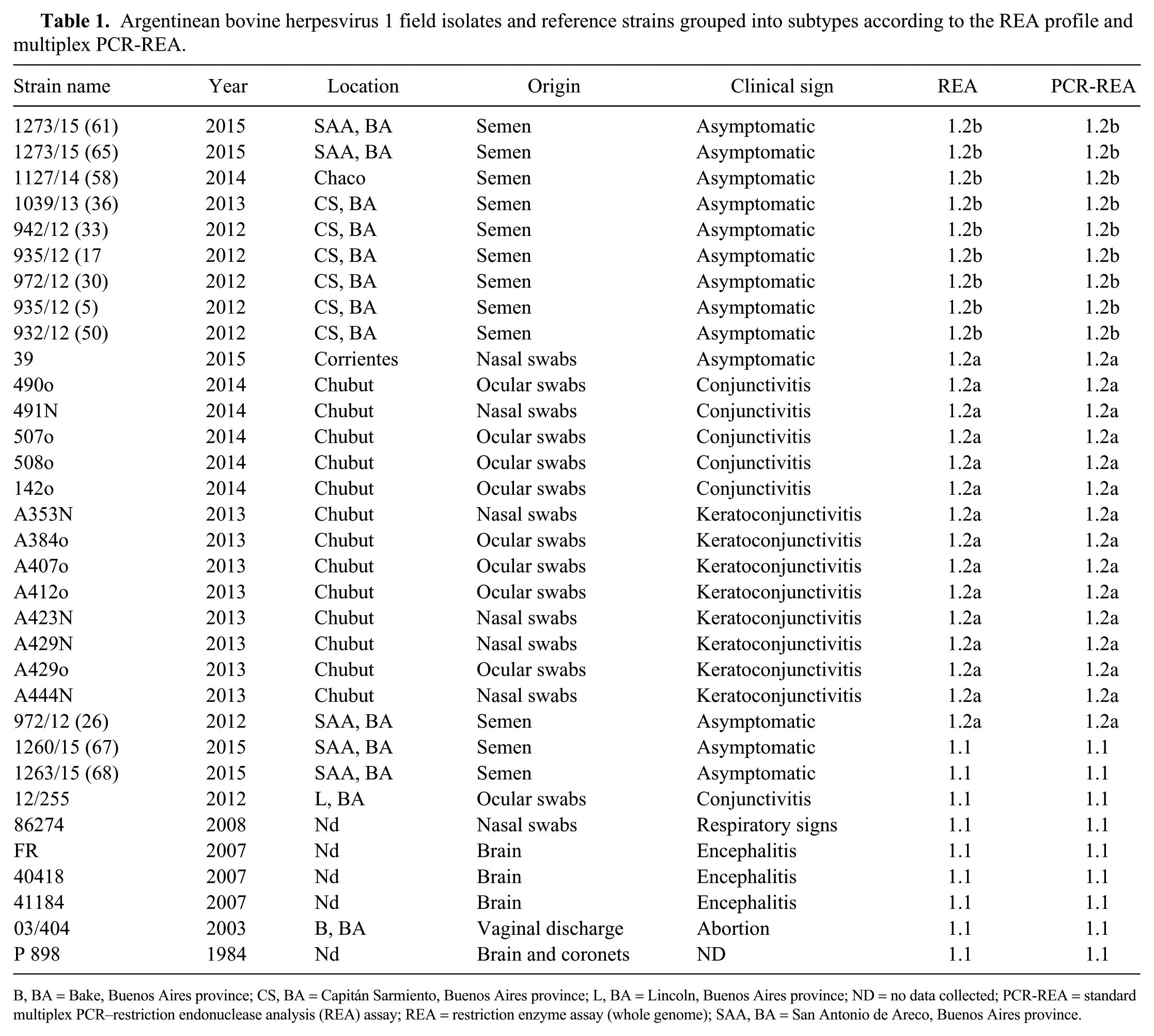
B, BA = Bake, Buenos Aires province; CS, BA = Capitán Sarmiento, Buenos Aires province; L, BA = Lincoln, Buenos Aires province; ND = no data collected; PCR-REA = standard multiplex PCR–restriction endonuclease analysis (REA) assay; REA = restriction enzyme assay (whole genome); SAA, BA = San Antonio de Areco, Buenos Aires province.
The assay consists of amplifying 2 BoHV-1 fragments by multiplex PCR using 2 sets of primers, followed by enzymatic digestion with HindIII and electrophoresis in agarose gel. The fragments to be amplified by the multiplex PCR were selected by an in-silico analysis using the complete genome of BoHV-1 (Cooper, K22 strains), available in GenBank (accessions KU198480.1 and KM258880). These sequences were compared with the REA of the strain ST (BoHV-1.2a) in agarose gels because the complete genomic sequence of this strain is yet to be published. 16 Two fragments that carry one HindIII restriction site (sequence AAGCTT), defined as RS1 and RS2, were found in the UL39 open reading frame and the US3 upstream intergenic region. The genital K22 subtype genome was used as reference strain to detect a mutation that produces different patterns for BoHV-1.1, -1.2a, and -1.2b subtypes. Primers were designed to amplify RS1, 493-bp (UL39, F: 5′-TCGTCGAAGAGCGTCCACACA-3′, and R: 5′-ACCGCGCTGTACCGGCAGCT-3′) and RS2, 700-bp (US3, F: 5′-TACAAATCGGCGGCGCCAAA-3′ and R: 5′-TTGTTGACGGCCAAGTATAA-3′; Vector NTI Suite v.8.0; InforMax-Invitrogen, Merelbeke, Belgium; Fig. 1).
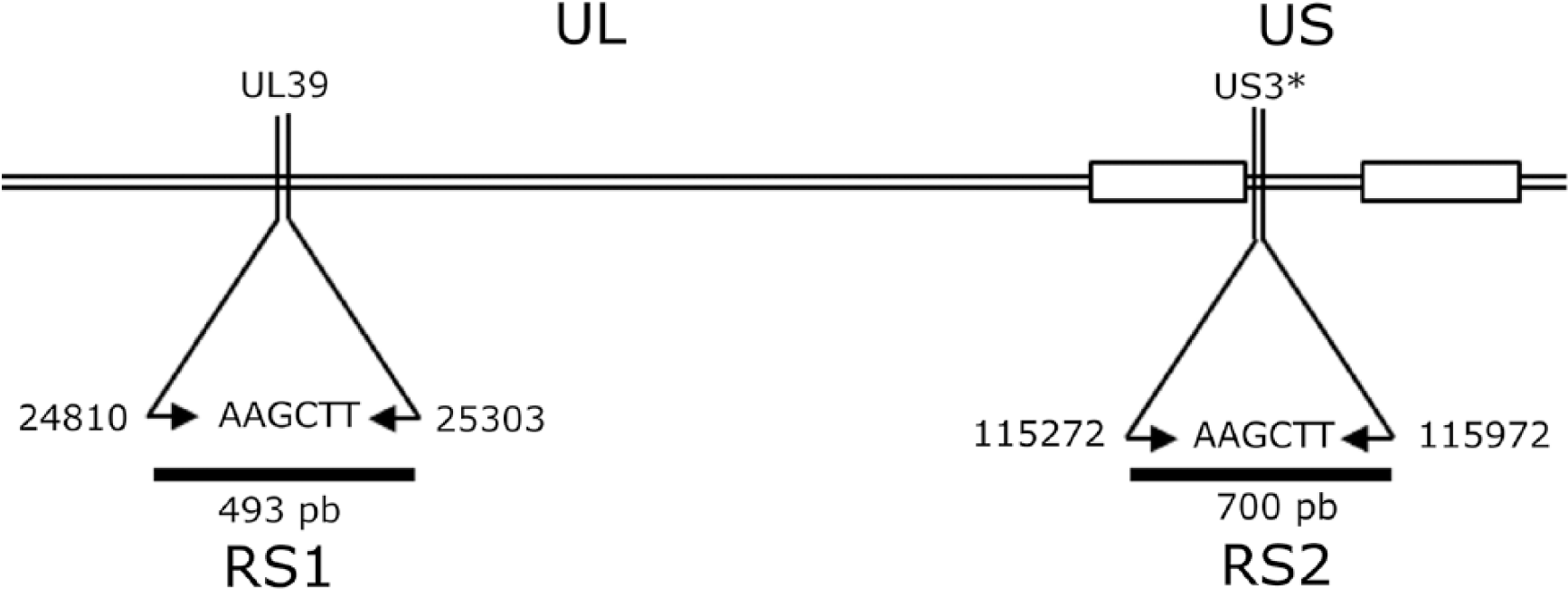

Organization of the bovine herpesvirus genome. The lines show the position and size of both RS1 and RS2 regions amplified by multiplex PCR. An asterisk (*) indicates the upstream intergenic region. The position numbers are according to the K22 reference strain (accession KM258880). UL = unique long region; US = unique short region.
Viral DNA was extracted from 200-µL suspensions of culture fluid and tissue extracts (QIAamp viral DNA kit; Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The multiplex PCR was carried out using a standard procedure. Briefly, the PCR reaction contained DNA (5 ng) of both RS1 and RS2 sense and antisense primers (0.3 µM each), PCR buffer (1×), dNTP (0.2 mM each), MgCl2 (2 mM), DMSO (6%), 1.5 U Taq DNA polymerase (Promega, Madison, WI), and ultrapure water up to 50 µL. Annealing temperature (54°C) was optimized for the simultaneous amplification of both RS1 and RS2 fragments. The PCR product was divided into 2 aliquots (25 μL each): 1 aliquot was incubated with HindIII enzyme for 1 h at 37°C, and the other aliquot was used as an undigested control. Given that both PCR and HindIII buffer reagents are compatible, the digestion of the PCR products can be done without a purification step.
Los Angeles (LA; American Type Culture Collection, VR-188), ST, 16 and K22 33 prototype strains of BoHV-1.1, BoHV-1.2a, and BoHV-1.2b subtypes, respectively, were used as reference viruses. Thirty-three field samples previously characterized by REA 21 (9 BoHV-1.1, 15 BoHV-1.2a, and 9 BoHV-1.2b field isolates) were analyzed (Table 1). In order to determine the amplicon sequence, both PCR products (RS1 and RS2) of 21 field isolate strains were sequenced twice in both directions using forward and reverse primers at the Sequencing Service of the Biotechnology Institute (INTA, Buenos Aires, Argentina). The sequences were visualized and edited using BioEdit. 12 Nucleotide and deduced amino acid sequences were aligned with ClustalX. 12 The RS1 and RS2 GenBank accessions are MN239798-MN239839.
The sensitivity of the multiplex PCR was evaluated by 10-fold serial dilutions carried out from cell culture DNA virus. 18 The correlation between the techniques was analyzed by χ2 test.
All field isolates were tested by PCR-REA analysis (Table 1, Supplementary Fig. 1). BoHV (1.1, 1.2a, 1.2b, 5a, 5b, 5c) and bubaline herpesvirus 1 (BuHV-1; Varicellovirus, Bubaline alphaherpesvirus 1) reference strains were tested. All BoHV-1 strains showed amplification of both RS1 and RS2 fragments, whereas all BoHV-5 strains and BuHV-1 showed amplification of only the RS1 fragment. Furthermore, BoHV-1.1 showed the 2 RS1 and RS2 fragments without cleavage. Subtype 1.2a had 1 fragment without cleavage (RS1); the RS2 fragment was cleaved into 2 sub-fragments. Finally, subtype 1.2b had both RS1 and RS2 fragments cleaved into 2 sub-fragments each (Fig. 2).
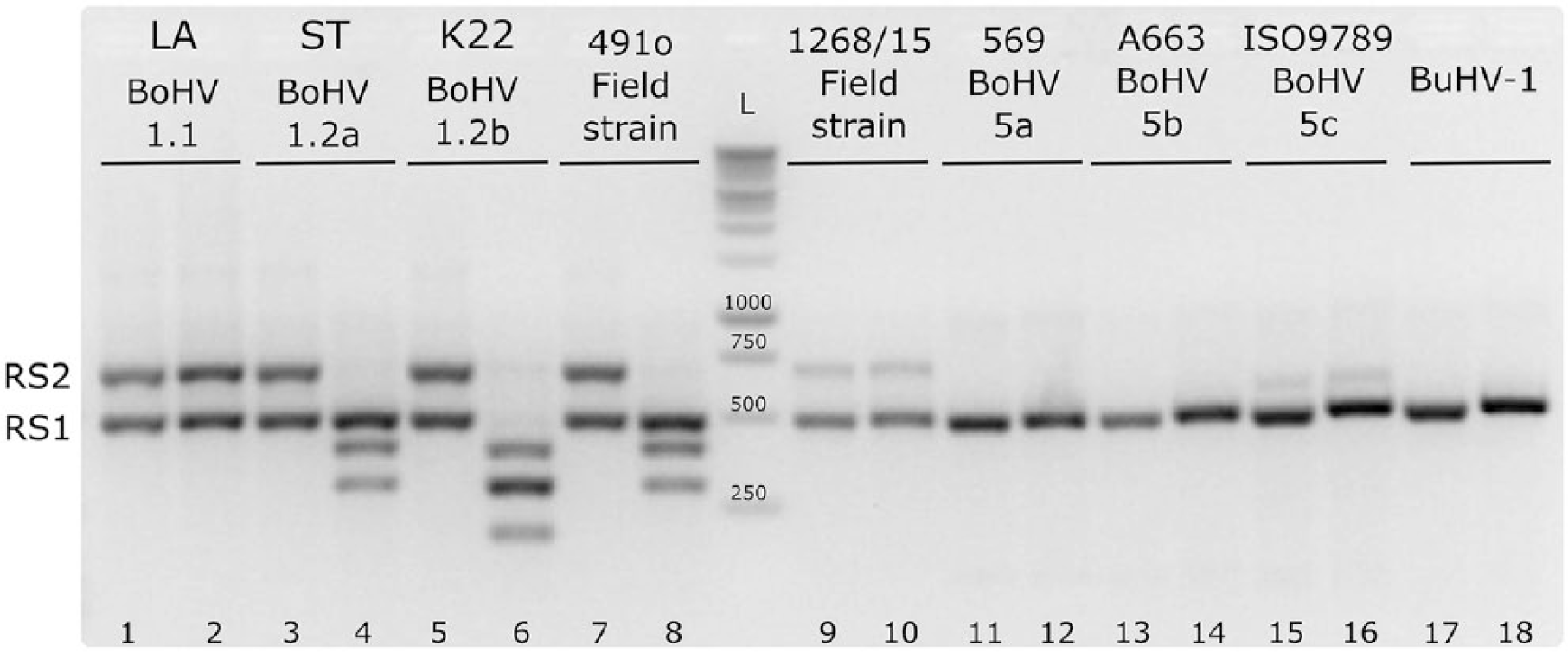
The agarose gel shows the pattern of BoHV-1 subtypes 1.1, 1.2a, and 1.2b, as well as the patterns of BoHV-5 subtypes 5a, 5b, and 5c, and the BuHV-1 references strains. The odd lines show the undigested PCR amplicons and the even lines the digested amplicons. L = ladder; RS1 = amplicon of 493 bp; RS2 = amplicon of 700 bp.
The sequences of the RS1 and RS2 fragments of 21 field strains were obtained. All of the BoHV-1.1 and -1.2a strains had a mutation T/C (AAGCT
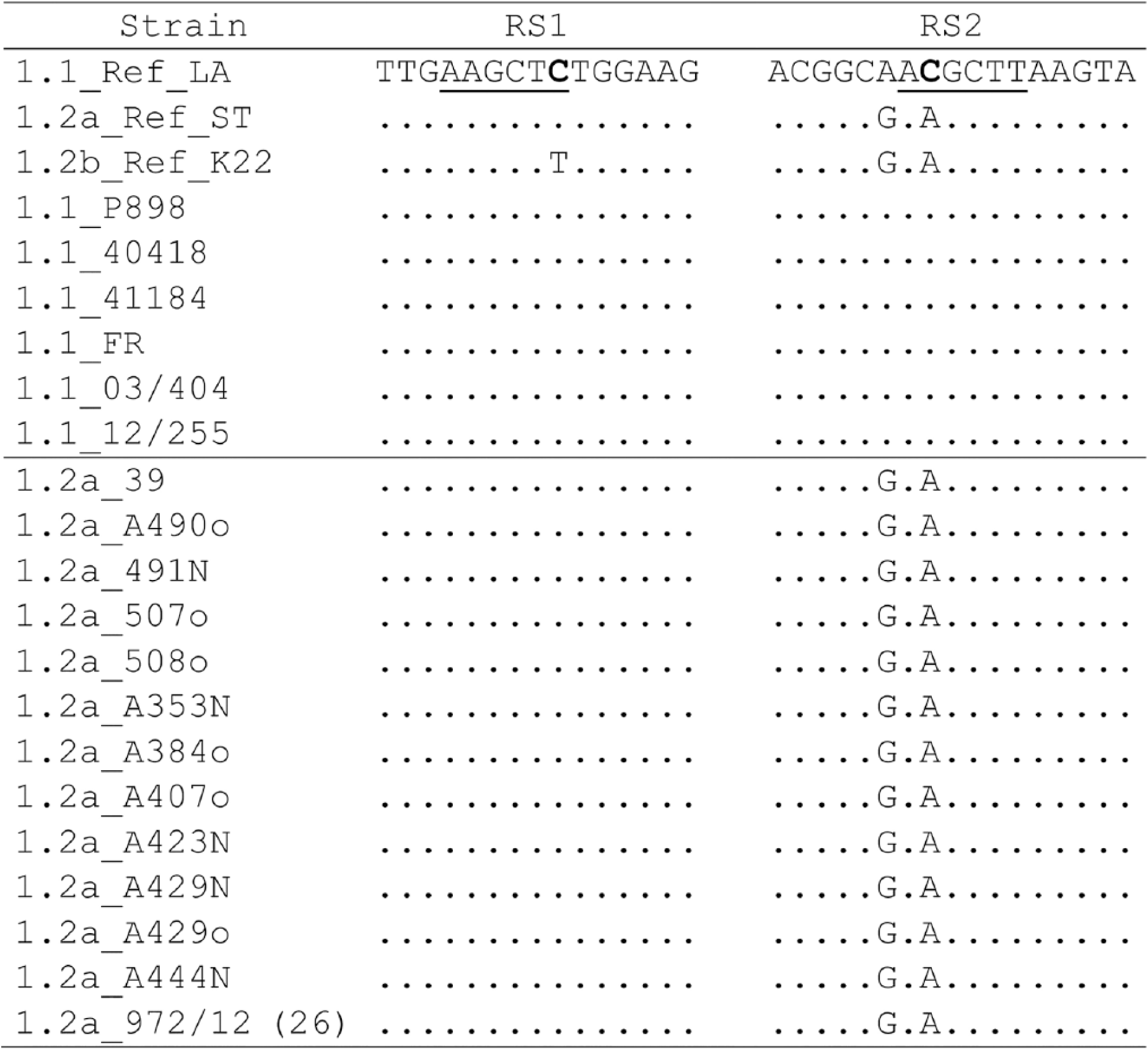
Multiple nucleotide sequence alignment of RS1 and RS2 fragments that shows the changes in the HindIII restriction sites (underlined). The nucleotide change is shown in bold.
The detection limit of the multiplex PCR assay was estimated at 160-260 genome copies/mL in DNA extracted from infected cell culture.
BoHV-1 and BoHV-5 can be isolated from cattle with similar clinical signs. 31 All viruses isolated from semen samples in our study, including BoHV-1.2b strains, were present in asymptomatic animals. In contrast, BoHV-1.2a and -1.1 strains were isolated mainly from nasal or ocular swabs from animals with clinical signs. Our standard multiplex PCR-REA assay could be an alternative to the currently available assays to detect and characterize BoHV-1 in various clinical samples.10,26
The sequencing of RS1 and RS2 fragments of all field strains allowed determination of the point mutation responsible for the change in the HindIII restriction site that generates the different profiles. In subtype 1.1, a nucleotide substitution was observed in both RS1 and RS2 fragments, resulting in the loss of the HindIII restriction site compared with referent K22 strain, generating the 2 bands observed with and without cleavage. In the 1.2a subtype, a nucleotide substitution was observed only in the UL39 gene, which produces the loss of the HindIII restriction site of RS1. It is noteworthy that the mutation observed in the UL39 fragment is the same in both of the subtypes (1.1 and 1.2a). Both for the field and for the reference strains, the mutation does not produce an amino acid change in the encoded protein. Finally, in subtype 1.2b, the HindIII restriction site is present in both RS1 and RS2 fragments, producing the cleavage of both PCR fragments.
Our standard multiplex PCR-REA assay was able to detect as few as 160 genome copies of BoHV-1 from cell culture supernatant. In addition, BoHV-1 DNA was detected from infected tissues without the need for enrichment using cell culture, but with lower sensitivity (data not shown). The identification and characterization of BoHV-1 into subtypes will allow us to study the epidemiology and distribution of BoHV-1. Furthermore, it may allow us to establish a pattern of distribution with the possibility of predicting changes of prevalent subtypes associated with livestock movements, type of production (extensive vs. feedlot), and management changes focused on increasing production, which could be useful information for the industry. Finally, given that the control programs in Argentina are based on vaccination with killed vaccine containing BoHV-1.1, this method could permit the typing of all detected viruses, data that could be used to alter the vaccine program.
Supplemental Material
Supplemental_material – Supplemental material for A new molecular method for the rapid subtyping of bovine herpesvirus 1 field isolates
Supplemental material, Supplemental_material for A new molecular method for the rapid subtyping of bovine herpesvirus 1 field isolates by Silvina S. Maidana, Samuel Miño, Romina M. Apostolo, Gabriel A. De Stefano and Sonia A. Romera in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. Juan Pablo Jaworski for the English revision.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by the Agencia Nacional de Promoción Científica y Tecnológica (ANPyCT), Ministerio de Ciencia, Tecnología e Innovación Productiva, Argentina (BID-PICT 1460), and Instituto Nacional de Tecnología Agropecuaria (PNSA 1115054 and 1115055).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
