Abstract
Currently, a variety of tests are used to detect bovine viral diarrhea virus (BVDV) in persistently infected (PI) cattle. These tests include immunohistochemical staining (IHC), antigen capture enzyme-linked immunosorbent assay (ACE), virus isolation (VI), and reverse transcription-polymerase chain reaction (RT-PCR). However, a lack of methods standardization could compromise the ability to consistently identify animals infected with BVDV. This study evaluated the diagnostic proficiency of current methods for detecting BVDV in infected cattle using intra- and interlaboratory comparisons. Samples were collected from 4 animals more than 7 months of age (2 BVDV negative animals, a PI animal, and a PI animal that previously lacked detectable virus in serum as determined by VI). Samples were submitted to 23 participating diagnostic laboratories using the respective laboratory's standard submission protocol. Samples collected for submission included: 1) serum for ACE, RT-PCR, and VI; 2) whole blood for RT-PCR and VI; and 3) skin biopsies for ACE and IHC. The ACE performed on skin provided the greatest consistency in detecting positive samples and a perfect level of agreement among laboratories. Reverse transcription-polymerase chain reaction and IHC performed well by correctly identifying ≤85% of samples positive for BVDV. Virus isolation performed on serum yielded the lowest consistency in detecting positive samples and the lowest level of agreement. The level of agreement between laboratories for detecting BVDV in persistently infected cattle ranged from perfect to less than expected by chance. The variation between laboratories suggests a need for training opportunities in standardized laboratory protocols and proficiency testing.
Introduction
Bovine viral diarrhea virus (BVDV) is a significant pathogen associated with gastrointestinal, respiratory, and reproductive diseases of cattle worldwide. 7 Persistent infection with BVDV occurs when a conceptus is infected with a noncytopathic strain of BVDV before the development of a competent fetal immune system (about gestation day 125). Persistently infected (PI) cattle are capable of shedding large quantities of virus throughout their lives and are considered the primary reservoirs for BVDV. Evidence exists that some PI animals are capable of producing antibodies that neutralize the persisting or resident strain of BVDV. 1,6 Brock et al. (1998) described a PI cow with an absence of detectable virus in serum and spontaneous production of virus neutralizing antibody. 1 Although BVDV could not be detected in the serum, virus was isolated from white blood cells (WBC) lysates from of this “atypical” PI cow. 1
The identification and removal of PI animals that serve as the natural reservoirs is essential to preventing the spread of BVDV. Currently, a variety of tests are used for this purpose. These tests include virus isolation (VI) of WBC lysates, tissues, or whole blood in cell culture; reverse transcription-polymerase chain reaction (RT-PCR); antigen capture enzyme-linked immunosorbent assay (ACE), and immunohistochemical (IHC) staining. Isolation of BVDV from WBC lysates followed by identification of the viral isolate by immunofluorescence or immunoperoxidase monolayer assay is one of the most reliable diagnostic techniques. This assay is considered to be the “gold standard” for the detection of BVDV. 5 Pooling of serum, milk, and supernatant from skin biopsy samples for the highly sensitive RT-PCR, which detects viral RNA, is becoming a popular screening method for detection of PI cattle. When determining the size of the pool, the pooling protocol should use the fewest number of tests required to identify all animals PI with BVDV in a herd. A simulation model for determining the economically optimum sample size in populations with various prevalences of PI found that the economically optimum sample size is dependent upon the prevalence of true positives in the population. 12 For a PI prevalence of 0.5% to 1.0%, the optimum number of samples in an initial pool is 20 to 30, and as prevalence increases the least-cost initial pool size decreases. 12 If the pool size is too large, there is an increased risk that any single pool will test positive, requiring additional testing to identify the small number of viremic animals in the positive pool. If the samples are grouped in unnecessarily small pools, the cost benefit of pooling samples is lost to the large number of negative pools tested for each positive pool identified. 8,9,11,12 The ACE, a relatively new assay available as a commercial test kit, a uses monoclonal antibodies to capture viral antigen Erns (gp48) and detects antigen-antibody complexes with enzyme-conjugated antibody by spectrophotometer. 5 Because of reduced cost and ease of sample collection, IHC staining of formalin-fixed, paraffin-embedded skin biopsies is widely used for the detection of PI animals. 2 This IHC staining uses the 15C5 monoclonal antibody, which reacts with the Erns protein of BVDV for the detection of diverse isolates of the virus. 2 These tests exhibit high levels of sensitivity and specificity that are considered adequate for use in screening programs aimed at detecting BVDV in infected cattle. 3,10,15
Although many tests are available for the detection of BVDV in infected cattle, failure to validate these tests or maintain laboratory technician proficiency could lead to variable results among diagnostic laboratories and compromise the overall ability to accurately detect BVDV in persistently infected cattle. The purpose of this study was to investigate the diagnostic proficiency of various methods for detecting BVDV in infected cattle using intra- and interlaboratory comparisons.
Materials and methods
Animals. This study was approved by the Auburn University Institutional Animal Care and Use Committee (AUIACUC No. 2004–0746). Samples were collected from 4 hybrid beef calves more than 7 months of age. Two animals were negative for BVDV, 1 was PI with a type 1b genotype of BVDV, and 1 was an animal PI with a type 2 genotype of BVDV that previously lacked easily isolatable virus in serum by VI. The animal that was PI with a type 2 genotype did not have titers to type 1 genotypes of BVDV (I-23 and SD1). However, this same PI animal had a titer of 1:2 to type 2 BVDV (PA131) and a titer of 1:8 to the homologous strain of BVDV. Immunohistochemical staining of multiple skin biopsies (ear notches) and VI of multiple WBC lysates were initially performed to confirm that both the typical and atypical PI animals were PI with BVDV as well as at the time of sample collection for this study. Further testing of samples from the atypical PI revealed that this heifer was unusual because virus could be isolated by IPMA from buffy coat samples, but virus could not be isolated on several occasions by IPMA from serum. This heifer is referred to as atypical because of the lack of easily isolated virus in serum. Results from RT-PCR on both serum and buffy coat samples revealed weak positives for BVDV. Weak positives were samples with faint bands of RT-PCR product on agarose gel as determined by subjective, visual evaluation. The heifer that was PI with a type 2 genotype of BVDV that previously lacked easily isolatable virus in serum by VI was estimated to have 3.5 × 103 CCID50/ml. In addition, serum neutralization (SN) assays for detecting antibodies to the resident strain of virus in the atypical PI heifer revealed the presence of serum neutralizing antibodies. The 2 BVDV negative calves were confirmed to be free of virus and anti-BVDV antibodies as determined by VI on WBC lysates and SN assays, respectively. Persistently infected calves were maintained and samples collected at a BVDV isolation facility. Calves negative for BVDV were maintained and samples collected at a location separate from the BVDV isolation facility.
Sample collection. Calves were restrained in a squeeze chute and head gate for sample collection. Animals were administered butorphanol (0.1 mg/kg) intramuscularly in the neck for sedation and sulfadimethoxine sustained release antibiotic boluses (137.5 mg/kg) orally for prevention of secondary infections after collection of skin biopsies. Whole blood (collected in EDTA) and serum (collected in 25-ml microvette syringe tubes b ) were collected by jugular venipuncture. After blood collection, skin biopsies were harvested from the left and right paralumbar areas of each calf. The paralumbar area was cleaned with 10% povidone-iodine surgical scrub, c and local anesthesia was provided by performing an inverted-L block with 25 ml of 2% lidocaine. d A 10-mm sterile punch biopsy instrument e was then used to collect skin biopsy samples. Each biopsy site was then closed with either #1 chromic gut suture f or #1 supramid suture g in a simple interrupted pattern. Skin biopsies were placed into additive-free glass tubes that contained either 5 ml of 10% neutral buffered formalin h for IHC, 5 ml of Dulbecco phosphate-buffered saline i for ACE, or no buffer for ACE according to laboratory preferences.
Triplicate samples from animals PI with BVDV and duplicate samples from animals negative for BVDV were collected, processed, and shipped to participating laboratories within 10 hours. A total of 562 ml of blood and 50 skin biopsies were collected from each of the BVDV negative calves. A total of 843 ml of blood and 75 skin biopsies were collected from each animal PI with BVDV. Samples collected for submission included: 1) serum for ACE, RT-PCR, and VI; 2) whole blood for RT-PCR and VI; and 3) skin biopsies for ACE and IHC.
Sample preparation and submission. After collection, samples were placed in a cooler and taken to the laboratory for processing within 2 hr. Each sample was randomly assigned an identification number based on the sample type (serum, whole blood, or skin biopsy) and the animal type (calves negative for BVDV, animal PI with BVDV, and atypical PI animal) to blind laboratories to the identification of the samples. After collection, samples were prepared for submission to each of the 23 participating laboratories. Clotted blood samples were centrifuged at 600 × g for 30 min, and serum was aliquoted into 4-ml cryogenic tubes j according to volume requirements for each laboratory. Before shipment, serum and whole blood samples were stored at 4°C while skin biopsies were held at room temperature. Samples were placed into specimen shippers, k with or without ice packs, and shipped by overnight mail. Coded samples were packaged and submitted according to each laboratory's standard submission protocol.
Diagnostic laboratories. All laboratories voluntarily participated in the study and were informed of the nature of the research before the initiation of the study. Each of the 23 participating laboratories was asked to perform all tests offered for the detection of cattle PI with BVDV.
Summary of experimental design. Blinded samples were collected on the same day from 2 animals PI with BVDV and 2 animals negative for BVDV. Samples were then submitted to diagnostic laboratories according to each laboratory's preference. A total of 10 samples (6 samples from 2 animals PI with BVDV and 4 samples from 2 BVDV negative animals) were submitted for each test performed.
Statistical analysis. The percentage of samples correctly identified as positive [percentage correctly positive = true positives/(true positives + false negatives) × 100], the percentage of samples correctly identified as negative [percentage correctly negative = true negatives/(true negatives + false positives) × 100], positive predictive value [positive predictive value (PPV) = true positives/(true positives + false positives) × 100], and negative predictive value for detecting cattle infected with BVDV [negative predictive value (NPV) = true negatives/(true negatives + false negatives) × 100] were calculated for each of the following diagnostic tests: IHC on skin, ACE on serum, ACE on skin, VI on serum, VI on whole blood, PCR on serum, and PCR on whole blood. The kappa value (κ) was used to determine the level of agreement between any 2 laboratories for each diagnostic test. A negative K indicates that the agreement between laboratories is less than expected by random chance, a κ of 0 indicates that the agreement is the same as chance, κ ≤ 0.20 = poor agreement, 0.21 ≤ κ ≤ 0.40 = fair agreement, 0.41 ≤ κ ≤ 0.60 = moderate agreement, 0.61 ≤ κ ≤ 0.80 = substantial agreement, κ > 0.80 = good agreement, and κ = 1 indicates perfect agreement. Chi-square statistical comparison was performed to evaluate the number of samples from the typical and atypical PI animals identified positive by each prospective test.
Results
Of diagnostic laboratories involved in this research, 43.5% (10/23) accurately identified all coded samples submitted for detection of PI cattle. In addition, 30.4% (7/23) of laboratories correctly identified ≥90% of samples positive for BVDV, and 21.7% (5/23) of laboratories correctly identified between 71% and 79% of samples positive for BVDV. Only 1 of the 23 participating laboratories correctly identified <50% of samples positive for BVDV for all tests performed. The ACE performed on skin provided the greatest ability to identify samples positive for BVDV (100%) and a perfect level of agreement among laboratories.
The ACE performed on serum correctly identified 91% of samples positive for BVDV. Immunohistochemical staining of skin biopsies correctly identified 90% of samples positive for BVDV. Reverse transcription-polymerase chain reaction performed on serum and whole blood correctly identified 85% and 93% of samples positive for BVDV, respectively. Virus isolation performed on whole blood correctly identified 88% of samples positive for BVDV. Virus isolation performed on serum yielded the lowest ability to identify samples positive for BVDV (76%) and the lowest level of agreement. The agreement level between laboratories for detecting animals infected with BVDV ranged from perfect (κ = 1.0) to less than expected by random chance (κ = −0.2). Analysis of 890 test results for samples submitted to 23 participating diagnostic laboratories is shown in Tables 1 and 2. Chi-square statistical analysis revealed no significant difference in the ability to correctly identify samples positive for BVDV obtained from typical or atypical PI animals.
Discussion
In this study, the diagnostic proficiency of tests used to detect BVDV in PI cattle was compared between 23 participating laboratories using intra- and interlaboratory assessments. Many tests are currently available for the accurate detection of BVDV from a variety of samples, including IHC, ACE, VI, and RT-PCR. Samples were collected from only 2 PI animals to create consistency among samples submitted to the diagnostic laboratories. Because of the small sample size, the percentage of samples correctly identified as positive does not directly correlate to sensitivity nor does the percentage of samples correctly identified as negative directly correlate to specificity. However, the percentage of samples correctly identified as positive and the percentage of samples correctly identified as negative does enable evaluation of the proficiency of the laboratories and their ability to accurately detect BVDV in diagnostic samples.
Laboratories A through J accurately identified all samples submitted for the detection of cattle infected with BVDV. Laboratories K through Q also performed well and demonstrated an acceptable ability to correctly identify samples positive for BVDV at ≥ 90%. However, laboratories R through V (percentage of samples correctly identified as positive for BVDV between 71% and 79%) and laboratory W (percentage of samples correctly identified as positive of 43%) confirm the need for validating testing procedures for the detection of cattle infected with BVDV. Overall, 17 of the 23 participating laboratories (74%) performed at an acceptable detection level, with ≥ 90% of samples correctly identified as positive for BVDV. Only 6 laboratories (26%) reported an unacceptable detection level, with ≤79% of samples correctly identified as positive for BVDV.
Results of assays used to detect bovine viral diarrhea virus in diagnostic samples summarized by test and laboratory. Immunohistochemical (IHC) tests were performed on skin biopsies. Reverse transcription-nested PCR (RT-nPCR) and virus isolation (VI) assays were performed on whole blood (wb) and serum
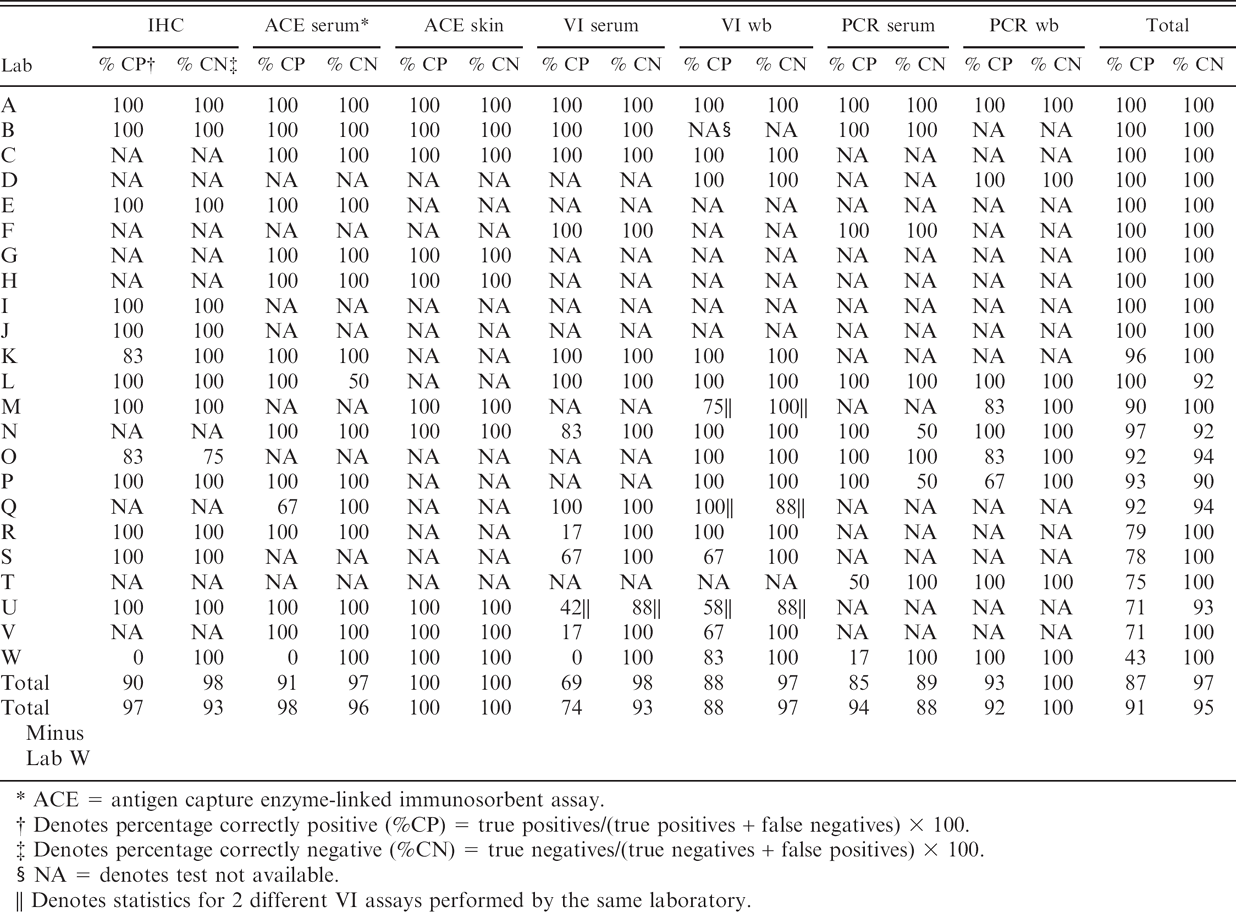
ACE = antigen capture enzyme-linked immunosorbent assay.
Denotes percentage correctly positive (%CP) = true positives/(true positives + false negatives) × 100.
Denotes percentage correctly negative (%CN) = true negatives/(true negatives + false positives) × 100.
NA = denotes test not available.
Denotes statistics for 2 different VI assays performed by the same laboratory.
Summary by test of 890 results for research samples submitted to 23 participating diagnostic laboratories*
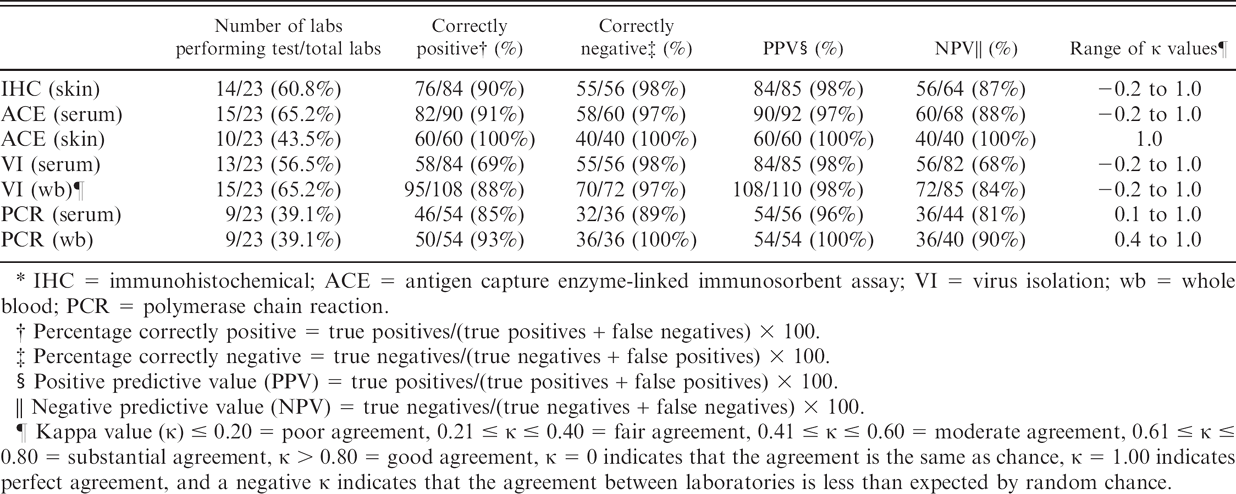
IHC = immunohistochemical; ACE = antigen capture enzyme-linked immunosorbent assay; VI = virus isolation; wb = whole blood; PCR = polymerase chain reaction.
Percentage correctly positive = true positives/(true positives + false negatives) × 100.
Percentage correctly negative = true negatives/(true negatives + false positives) × 100.
Positive predictive value (PPV) = true positives/(true positives + false positives) × 100.
Negative predictive value (NPV) = true negatives/(true negatives + false negatives) × 100.
Kappa value (κ) ≤ 0.20 = poor agreement, 0.21 ≤ κ ≤ 0.40 = fair agreement, 0.41 ≤ κ ≤ 0.60 = moderate agreement, 0.61 ≤ κ ≤ 0.80 = substantial agreement, κ > 0.80 = good agreement, κ = 0 indicates that the agreement is the same as chance, κ = 1.00 indicates perfect agreement, and a negative κ indicates that the agreement between laboratories is less than expected by random chance.
When comparing the results from the atypical and the typical PI animals for each assay, there was no significant difference in the ability to detect BVDV in samples from the 2 PI animals. Analysis of 890 test results for whole blood, serum, and skin biopsy samples revealed that the ACE performed on skin biopsy samples was the most repeatable test for detecting cattle infected with BVDV, which is consistent with previous studies. 3,15
Virus isolation performed on serum was the least repeatable at detecting PI animals and yielded the lowest ability to identify samples positive for BVDV and the lowest level of agreement between laboratories. The ability to correctly identify samples positive for BVDV for most tests (IHC, ACE from serum, VI from serum, PCR from serum) improved with the removal of laboratory W from the statistical calculations (Table 1). These results were unexpected because VI from WBC lysates has long been considered the most reliable method for the detection of viremic cattle and is the “gold standard” diagnostic technique. 4,14 Previous studies have demonstrated close agreement between IHC and VI. In one of these studies, specimens collected from 15 PI animals demonstrated complete agreement between IHC and VI while only 13 samples were positive by ACE. 16 Of the 104 cattle that tested negative by parallel tests, there was complete agreement among the tests, which indicates 100% sensitivity when comparing IHC to VI. 16 In another study, 97.5% agreement was reported between IHC on skin biopsies and VI on peripheral blood leukocytes, and all 45 animals that tested negative by VI were also negative by IHC. 13 Cornish et al. also reported complete agreement among IHC from skin, ACE from skin, VI from WBC lysates, and RT-PCR from WBC lysates for calves PI with BVDV. 3 The discrepancy between our comparisons of IHC, VI, and ACE and results from previous studies may be due to differences in the samples tested (WBC lysates vs. serum) or in VI protocols used by the laboratories participating in this study. In previous studies, VI was validated by a single laboratory to make positive comparisons between and among samples. 4,13,14,16 Some laboratories participating in this study may not routinely perform VI for the detection of BVDV and may not routinely validate their assays for VI of BVDV. The ability of different laboratories to accurately detect BVDV by VI may be influenced by susceptibility of their cell cultures to infection, the cell culture medium, the serum used to supplement media, the testing procedure, the method and reagents used for immunological labeling, laboratory contamination of samples or cell cultures, and familiarity with the assay protocol. 14
When choosing a diagnostic test for screening purposes it is important to consider the sensitivity, specificity, and the prevalence of disease in the population being tested. A PPV represents the percentage of animals with positive tests that are truly infected with BVDV. An NPV represents the percentage of animals with negative tests that are truly not infected with BVDV. The ACE performed on skin provided the greatest ability to accurately identify animals that were both infected with BVDV (PPV = 100%) and animals truly negative for BVDV (NPV = 100%). All of the diagnostic tests used in this study had a PPV ≥96%. Virus isolation performed on serum demonstrated the lowest NPV at 68%, but the remaining diagnostic tests had an NPV ≥81%. The ability to control BVDV is directly related to the ability of diagnostic tests to correctly identify animals that are infected with BVDV. With a likely decrease in the prevalence of BVDV within the cattle population because of the implementation of control and eradication programs, the ability to accurately identify animals that are truly free of disease will be greater (higher NPV). However, a decrease in BVDV prevalence will decrease the ability of a given test to correctly identify animals infected with BVDV (lower PPV). A lower PPV suggests more false positives in the testing population and would require follow-up testing to confirm positive results found in the initial screening of calves for persistent BVDV infection. When using a test for screening purposes in a control and eradication program, it is important to choose a test with a high sensitivity and high NPV to ensure that false negatives do not occur. Animals with false negative test results could seriously undermine efforts to control and eradicate BVDV. Thus, care must be exercised when selecting the appropriate diagnostic test for detecting cattle PI with BVDV.
Each method to detect BVDV has advantages, disadvantages, and applicability for different diagnostic situations. The results of this study indicate that IHC, ACE, VI, and RT-PCR can all be reliable tests for the detection of BVDV in PI cattle but that variation among assays and laboratories suggests a need for training opportunities in standardized laboratory protocols and proficiency testing. Before a voluntary BVDV eradication program can begin in the United States, diagnostic laboratories should complete and pass annual proficiency tests similar to those tests offered for other infectious diseases such as Johne disease and brucellosis in cattle and pseudorabies in swine.
Acknowledgements
YiJing Zhang, Ann Cochran, Pat Galik, Kay Riddell, George Fincher, and Joe Galik provided technical assistance with this project.
Footnotes
a.
HerdCheck, IDEXX Laboratories, Inc., Westbrook, ME.
b.
25 ml microvette syringe tube, Sarstedt Inc., Newton, NC.
c.
Povidone-iodine surgical scrub 10%, Farnam Companies Inc, Phoenix, AZ.
d.
Lidocaine injection 2%, Hospira, Inc., Lake Forest, IL.
e.
10 mm sterile biopsy punch, Acuderm Inc., Ft. Lauderdale, FL.
f.
#1 chromic gut suture, Ethicon, Inc., Somerville, NJ.
g.
#1 supramid suture, S. Jackson, Inc., Alexandria, VA.
h.
Neutral buffered formalin 10%, Fisher Scientific International, Inc., Pittsburgh, PA.
i.
Dulbecco phosphate-buffered saline, Gibco-Invitrogen, Grand Island, NY.
j.
4-ml cryogenic tube, Corning, Inc., Corning, NY.
k.
#STP 100 specimen shipper, Saf-T-Pak, Inc., Edmonton, AB, Canada.
