Abstract
Desmozoon lepeophtherii is a microsporidian associated with gill disease in farmed Atlantic salmon (Salmo salar). Detection of the parasite in histologic tissue sections is challenging using common histochemical stains given that the small, widely distributed parasite spores typically occur individually or in small clusters. We compared the ability of 4 histologic methods to detect D. lepeophtherii spores in serial sections of Atlantic salmon gill tissue: hematoxylin and eosin (H&E), Gram–Twort (GT), calcofluor white (CW), and immunohistochemistry (IHC). Using CW as a benchmark to calculate a relative ratio, IHC consistently detected more spores than CW (median: 1.3), followed by GT (median: 0.2) and H&E (median: 0.1). IHC detected significantly more spores than GT (p < 0.05) and H&E (p < 0.05), and GT more than H&E (p < 0.05). We found significant underestimation of numbers of microsporidia spores in gill disease in Atlantic salmon using conventional histochemical stains and recommend the use of CW or IHC to detect the parasite in tissue sections.
Microsporidia are spore-forming parasites that infect all major animal groups. 3 The microsporidian Desmozoon lepeophtherii (syn. Paranucleospora theridion) was discovered initially in sea lice (syn. salmon louse; Lepeophtheirus salmonis Krøyer, 1838) 5 and described later in farmed Atlantic salmon (Salmo salar). 12 In salmon, the microsporidian forms 2 types of spores: auto-infective and environmental. The auto-infective spores (~1 µm diameter) develop in the cytoplasm of the gill epithelial cells, endothelial cells of blood vessels, macrophages, and polymorphonuclear leukocytes. 12 The environmental spores (~2.5 × 2.0 μm) develop within the nuclei of gill and skin epithelial cells 12 and gastrointestinal epithelium. 19 D. lepeophtherii is highly prevalent in the gill tissue of farmed salmon when screened by molecular methods,4,6 but parasite loads tend to be greater in fish with gill disease.6,15 The lesions associated with the presence of the parasite in gills include hyperplasia, hypertrophy, and necrosis of lamellar epithelial cells, and infiltration by phagocytic cells of degenerate hypertrophic epithelium.10,19 However, in outbreaks of complex gill disease of multiple causes, 7 the specific lesions caused by individual agents can be challenging to discern, including those caused by D. lepeophtherii. Additionally, the small size of the spores and the wide distribution of the more frequently occurring single spores or small clusters in salmon gills makes this organism difficult to detect using conventional histologic techniques.10,13 We compared the ability of 4 histologic techniques to detect D. lepeophtherii spores in sections of infected Atlantic salmon gill tissue.
Atlantic salmon gill tissue samples (n = 14), taken previously for diagnostic purposes and fixed in formal saline, were processed routinely for histologic examination. Samples were selected based on the presence of proliferative changes and the presence of D. lepeophtherii spores.10,19 Three of the selected fish were positive for D. lepeophtherii by reverse-transcription real-time PCR (RT-rtPCR), performed as described previously from gill tissue biopsies. 12 No RT-rtPCR results were available for the remainder of the fish. Five of the selected samples had amoebic gill disease, 1 and 8 had lesions suggestive of “Candidatus Branchiomonas cysticola” epitheliocysts. 17 Samples were from 4 different marine farm sites on the west coast of Scotland collected between late August to late October 2012 and 2014. Water temperature was 12–14°C in all farms during sampling, and salinities were 27–32%.
Samples were sectioned (3 μm) sequentially and stained with hematoxylin and eosin (H&E), Gram–Twort (GT), 10 and calcofluor white (CW; Fluka, Buchs, Switzerland) according to the manufacturer’s instructions. Briefly, one drop of KOH (10% w/v) and 1 drop of CW reagent were added to the tissue sections and slides were incubated for 1 min prior to application of a coverslip and examination under ultraviolet light (excitation range: 300–440 nm). All sections were examined microscopically (Olympus BX51 microscope; KeyMed, Southend-on-Sea, UK), photographed (Olympus DP70 digital camera system; KeyMed), and photographs analyzed (analySiS software; Soft Imaging System, Munster, Germany).
Rabbit polyclonal antiserum raised against spores of the closely related Enterocytozoon bieneusi 14 was used as the primary antibody for immunohistochemistry (IHC). Tissue sections mounted on slides (Superfrost Plus; Menzel-Gläser, Braunschweig, Germany) were dewaxed in 2 changes of xylene, rehydrated through graded alcohols, then placed in phosphate-buffered saline (PBS; pH 7.6) for 5 min. Heat-induced epitope retrieval (HIER) was performed in an autoclave at 121°C for 10 min in 1 mM of ethylenediamine tetra-acetic acid (EDTA) buffer (pH 8), containing 0.05% Tween 20. After HIER, slides were cooled for 20 min at 21°C, washed with PBS (5 min), and the tissue encircled (wax ImmEdge pen; Vector Laboratories, Burlingame, CA). Endogenous tissue peroxidase activity was quenched (Dual endogenous enzyme block; Dako, Glostrup, Denmark) for 10 min, as per manufacturer’s instructions, and then rinsed with and placed in fresh PBS (5 min). Nonspecific antibody binding was blocked by application of 25% normal goat serum (NGS) in PBS for 30 min at 21°C followed by washing for 5 min in PBS. Slides were drained, and the primary antibody was applied diluted 1/400 in 25% NGS in PBS and incubated for 1 h at 21°C. Slides were subsequently rinsed and then washed with PBS (3 × 5 min) and the bound antibody visualized (EnVision+ system; Dako) as per manufacturer’s instructions. Slides were washed again in PBS (5 min), incubated with the chromogen (3,3’-diaminobenzidine, DAB; Dako) for 10 min at 21°C, and the reaction stopped by washing with tap water. Slides were counterstained with 10% Giemsa solution (MilliporeSigma, St. Louis, MO) for 2 min, to enable melanin to be distinguished from DAB (Miller RT. Technical immunohistochemistry: achieving reliability and reproducibility of immunostains. Soc Appl Immunohistochem Ann Meeting; Sept 2001; Flushing, NY. Available from: https://pdfs.semanticscholar.org/7b7b/0ff695a5d8ee05a07bd4b4631b92a29aa9b8.pdf?_ga=2.246256323.695093924.1571253532-1669730724.1570566725), washed in tap water, counterstained with hematoxylin QS (Vector Laboratories) for 20 sec, rinsed in tap water, dehydrated through graded alcohols, cleared in xylene, and mounted (VectaMount; Vector Laboratories). Fish 10, which was observed to have a high number of spores by Gram stain, was used as a positive control. For the methodology negative control, the primary antibody was replaced with normal rabbit serum diluted identically. For the tissue negative control, a gill sample processed identically as the positive cases, and negative for D. lepeophtherii by RT-rtPCR, was used.
For quantification of the parasite, 3–6 filaments per gill and 40 interlamellar spaces (IS) per filament were selected randomly (a maximum of 240 IS/gill, or a minimum of 120 if the tissue sample was small) and examined. The total number of IS containing spores was counted and the mean number of positive IS per filament calculated. Serial sections allowed the same filaments to be used for each of the 4 detection methods. The mean number of positive IS per filament detected by CW staining was selected as the benchmark for the quantification studies, and the number of positive IS detected using the other techniques expressed as a ratio to CW to determine the relative spore load. Statistical analyses were performed (R software v.3.5.3; https://www.r-project.org/). The data were not normally distributed (Shapiro–Wilk test), and the Friedman test was performed to determine whether there were significant differences between techniques with respect to detection of relative spore load. Pairwise comparisons between techniques were subjected to the Wilcoxon signed-ranked test using the MASS package. 18 The significance threshold was set at p ≤ 0.05.
Spores stained with H&E and GT were spherical-to-oval, ~1 μm diameter, and present within the cytoplasm of degenerate gill epithelial cells (Fig. 1). Spores stained with H&E were eosinophilic, had a slightly birefringent wall, and were only detected when present as aggregates and using 1,000× magnification. Spores stained with GT were gram-positive and appeared singly or in aggregates of 2–8 (Fig. 2). Other smaller structures (0.7 μm diameter), usually present in clusters, had variable staining and appeared either weakly gram-positive or gram-negative and mostly in direct contact with cell debris. These structures were not considered to be spores, and therefore not included in the quantification to avoid potentially false-positive material.
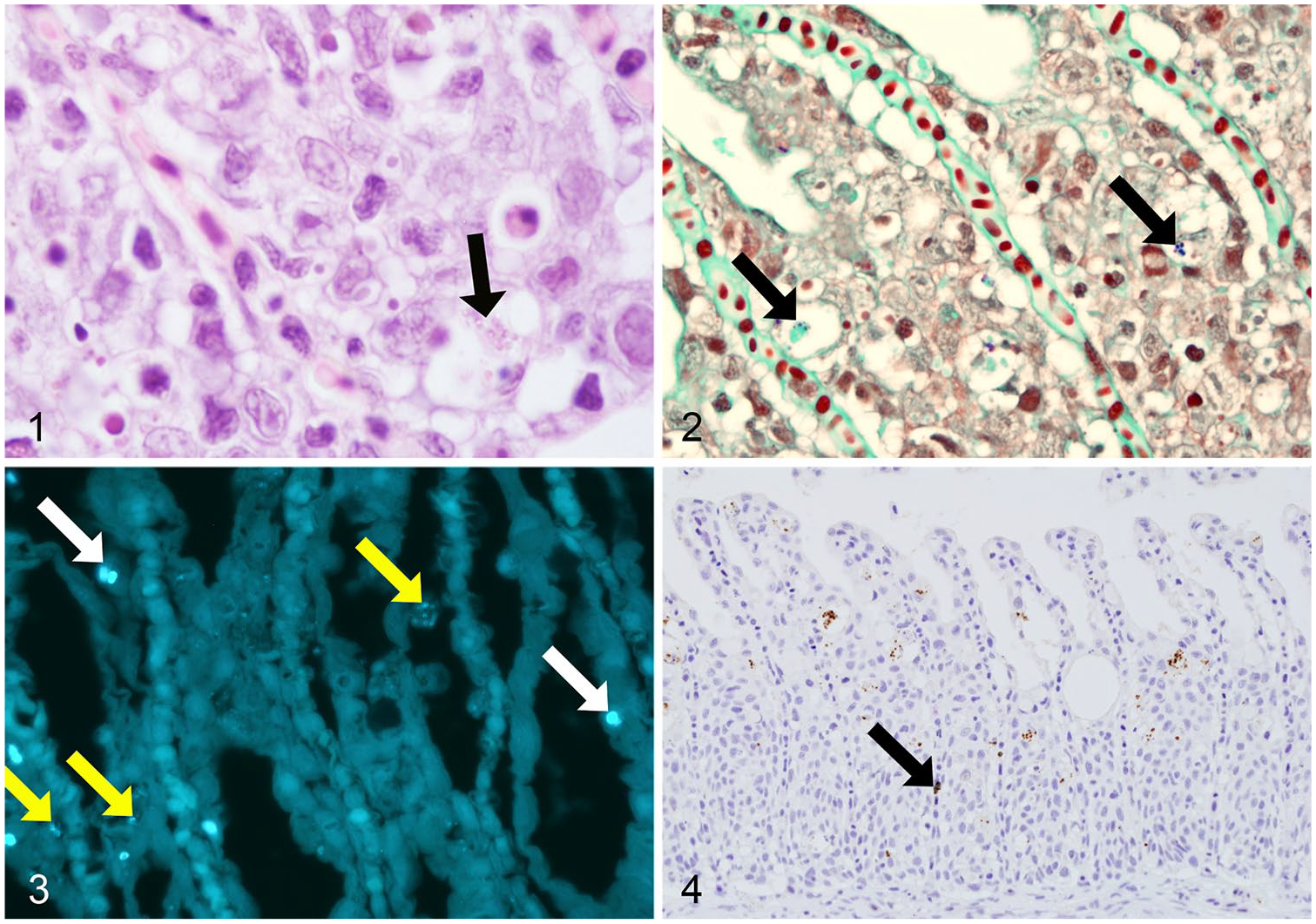
Histologic sections of gills from farmed Atlantic salmon with microsporidian spores of D. lepeophtherii.
CW stained 2 types of spores (Fig. 3). One was smaller (1.1 μm length), ellipsoidal, and present in the cytoplasm of cells along the lamellae, but the specific cell types were difficult to distinguish because of the absence of tissue counterstaining with this technique. These smaller spores were present in aggregates of 3–20 and, when not in aggregates, 600× magnification was required to identify them given their low fluorescence. Larger, oval, spore-like structures were clearly visible at 200× magnification given their stronger fluorescence and larger size (2.5 μm length). These larger structures were present singly or in pairs, typically within cells but it was unclear if they were located within the nucleus or the cytoplasm of the cells.
Spores were present in gills from all fish when stained with CW, including 4 fish in which no other staining method could detect spores (only the larger spores were detected). With the IHC technique, abundant spherical structures, ~1 μm diameter, labeled intensely within the cytoplasm of degenerate and nondegenerate gill lamellar epithelial cells (Fig. 4) and within macrophages present in the hypertrophied and necrotic epithelium. Melanin stained dark green, clearly differentiating it from positive immunolabeling (Fig. 4). All negative control preparations were devoid of labeling.
The medians (interquartile range) of the relative spore load detected with H&E, GT, and IHC were 0.1 (0.2–0), 0.2 (1.1–0), and 1.3 (1.9–0.1), respectively (Supplementary Fig. 1), and the differences between techniques were significant (p ≤ 0.05). The Wilcoxon signed-rank test showed significant differences between pairwise comparisons; IHC detected significantly more spores than GT (p ≤ 0.05) and H&E (p ≤ 0.05), and GT more than H&E (p ≤ 0.05).
Complex gill disease is an important problem for the farmed Atlantic salmon industry. 7 Although D. lepeophtherii has been associated with gill lesions and is found regularly in gills by molecular methods, detection of the microsporidian in tissue sections, and therefore in association with specific lesions, is generally difficult using routine H&E-stained sections. We used an antibody raised against E. bieneusi spores, 14 which shares 82% partial small subunit ribosomal RNA sequence identity with D. lepeophtherii, 12 which fortuitously cross-reacted with the auto-infective spores of D. lepeophtherii present in tissue sections, resulting in intense labeling and greater signal than the 3 other staining methods examined. Cross-reactivity between microsporidian spores using monoclonal and polyclonal antibodies has been reported previously, more commonly between species with higher phylogenetic relatedness, and attributed to the presence of shared antigenic epitopes. 8 The polyclonal antibody that we used did not cross-react with Encephalitozoon spp. spores when tested, 14 but did label spores of D. lepeophtherii. However, IHC identified only the smaller auto-infective spores of D. lepeophtherii, not the larger environmental spores detected by CW. The expression of common antigens between D. lepeophtherii and E. bieneusi auto-infective spores may be a result of their close phylogenetic relationship and common characteristics in their developmental cycles. 12
As a benchmark, the fluorochrome CW, shown to stain D. lepeophtherii spores successfully in previous studies,7,19 was selected given its high specificity for binding to the chitin present on the inner surface of microsporidian spores. 2 CW detected spores in all samples examined, including the 2 sizes of spores, in agreement with the life cycle of D. lepeophtherii in Atlantic salmon described previously. 12 However, the small spores (1.1 μm) were less intensely fluorescent, which may be a result of their thinner spore wall containing less chitin.
Compared with CW, GT stained less than half the number of spores per filament. This may be a result of counting only the strongly gram-positive spores and not including structures with a weak gram-positive stain. Variable Gram staining has been reported by other modified Gram techniques (e.g., Brown–Brenn) for microsporidian spores, 9 and previous studies 11 found sporoblasts and immature microsporidian spores stained gram-intermediate or gram-negative in contrast to mature forms that were mostly gram-positive. However, variable Gram staining can be the result of over-differentiation when performing this technique. 11
The H&E stain demonstrated the lowest number of D. lepeophtherii spores, and this was in agreement with previous studies in which different staining methods were compared to detect other species of microsporidian spores. 13 However, the birefringence of the wall, a characteristic that results from the presence of chitin in the endospore layer, does aid distinguishing spores from cell debris. 16 Although all of the methods investigated detected D. lepeophtherii spores, we were unable to definitively detect any developmental stages of the parasite.
We showed that the number of D. lepeophtherii spores present in tissue sections is severely underestimated using routine GT and H&E histochemical staining techniques. These techniques are useful for general screening and are possibly sufficient for routine complex gill disease diagnosis when the number of D. lepeophtherii spores is very high. However, the use of both IHC and CW methods is preferable for more detailed assessment of complex gill disease because they allow clear identification of the spores, the main characteristic for identification of D. lepeophtherii. An RNA in situ hybridization method has been reported that is capable of detecting developmental stages of D. lepeophtherii. 19 Although excellent for research purposes, the development of more practical approaches, such as a DNA-based oligoprobe in situ hybridization technique capable of detecting all stages of the parasite, would be more suitable for routine diagnostic purposes and complement the techniques described in our study.
Supplemental Material
Supplemental_material – Supplemental material for Comparison of histologic methods for the detection of Desmozoon lepeophtherii spores in the gills of Atlantic salmon
Supplemental material, Supplemental_material for Comparison of histologic methods for the detection of Desmozoon lepeophtherii spores in the gills of Atlantic salmon by Ana Herrero, Francesc Padrós, Sara Pflaum, Chris Matthews, Jorge del-Pozo, Hamish D. Rodger, Mark P. Dagleish and Kim D. Thompson in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
The rabbit polyclonal antiserum raised against Enterocytozoon bieneusi was kindly provided by Prof. Abhineet S. Sheoran (Tufts University, Grafton, MA).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
A Herrero was funded by Moredun Scientific and Fish Vet Group. MP Dagleish was funded by the Rural and Environment Science and Analytical Services (RESAS) division of the Scottish Government.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
