Abstract
We report herein the use of crude extracts obtained from samples of Taylorella equigenitalis–infected horses for the purpose of multi-locus sequence typing (MLST). Samples (n = 36) were collected from horses in South Africa from 1996 to 2017: 34 from genital swabs (stored at −20°C for 2–3 y) and 2 from cryopreserved raw semen aliquots (stored at −70°C for 18 y) prior to assay. The MLST assay showed a single sequence type (ST), designated ST4, that supported a point introduction and thus a common source for the South African outbreak of contagious equine metritis.
Taylorella equigenitalis, a gram-negative, microaerophilic, frequently pleomorphic bacterium, 18 is the causative agent of contagious equine metritis (CEM), a non-systemic, venereally transmitted disease of horses. T. equigenitalis is one of only 2 species of the Taylorella genus, a member of the Alcaligenaceae family. 9 This organism causes an inapparent carrier status without associated clinical signs in stallions and temporary infertility in affected mares characterized by mucopurulent vaginitis, cervicitis, or endometritis. 19 Although the majority of mares rid themselves of infection, 20–40% may become carriers. 11 T. equigenitalis is spread by natural mating or artificial insemination with contaminated semen and by fomite transmission.1,16,19 The effect on equine reproductive efficiency, especially in systems utilizing natural mating, as well as international trade restrictions, can result in substantial economic losses to the equine industry. These outcomes have resulted in CEM being one of the most regulated equine bacterial diseases in the world.1,19
The 2018 World Organisation for Animal Health (OIE) gold standard for identification of T. equigenitalis is bacterial culture (http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.05.02_CEM.pdf). However, given associated difficulties, including the fastidious transport and growth requirements required for isolating T. equigenitalis, various PCR-based methods have been developed to detect and additionally differentiate T. equigenitalis from the other member of the genus, T. asinigenitalis.5,12,20
Since the first reported outbreak in the United Kingdom in 1977, 3 CEM has attained worldwide distribution. The most recently reported outbreaks in non-endemic countries include a 2008 outbreak in Portugal, 15 a 2008–2011 outbreak in the United States, 6 a 2012 outbreak in the United Kingdom, 14 and a 2015 outbreak in South Korea. 10 South Africa was considered to be free of CEM until the first reported case in May 2011 (OIE. Immediate notification (09/05/2011) of contagious equine metritis, http://www.oie.int/wahis_2/public/wahid.php/Reviewreport/Review?reportid=10553). Since identification of the South African outbreak, development of molecular techniques such as MLST and genome sequencing have allowed further characterization of Taylorella spp.4,8,9 We report herein the application of an MLST assay to crude extracts from both genital swabs and cryopreserved semen samples in the absence of stored isolates.
T. equigenitalis–positive genital swab samples were collected from mares and male horses in South Africa from 2011 to 2017 (n = 33). This inventory originated from samples obtained during both a nationwide stallion screening program, legislated subsequent to outbreak recognition, and an epidemiologic traceback. 13 Heightened awareness of the CEM-associated risk in South Africa prompted additional voluntary submissions of cryopreserved semen straws derived from ejaculates collected and processed from stallions prior to outbreak identification in 2011. In 2015, this submission of cryopreserved semen identified an additional T. equigenitalis–positive case linked to a stallion that had semen collected and cryopreserved in 2008. The stallion was subsequently gelded and on follow-up was found to be positive 7 y later (case 34).
The traceback exercise had furthermore identified a subpopulation focus at the South African Lipizzaner Centre and, based on this finding, archived semen samples (n = 2; cases 1 and 2) were accessed in August 2014. These samples had been collected from Lipizzaner stallion residents of this Centre during an equine viral arteritis investigation 7 in 1996 and were included in the MLST analysis.
Swabs were agitated for 5 s in 0.4 mL of 0.1 M phosphate-buffered saline (PBS; pH 7.4) in separate 1.5-mL microfuge tubes. In the case of semen samples, after thawing, 0.1 mL was added to 0.4 mL of 0.1 M PBS as described above. All samples were then pelleted by centrifugation (18,000 × g, 60 s), and the supernatant was aspirated. The resultant pellet was resuspended in 0.1 mL of nuclease-free water (MilliporeSigma, St. Louis, MO) and then heated at 95°C for 15 min. Insoluble material was pelleted by centrifugation (18,000 × g, 60 s) and the supernatant stored at −20°C.
Lysates were amplified using specific primers for 7 loci (gltA, citrate synthase; gyrB, gyrase subunit B; fh, putative hydratase; shmt, serine hydroxymethyltransferase; tyrB, tyrosine aminotransferase; adk, adenylate kinase; txn, thioredoxin) as described previously. 4 Primers for the PCR 2 were standardized with universal tail C and tail D sequences (Integrated DNA Technology, Whitehead Scientific, Cape Town, South Africa). PCR amplification (KAPA SYBR FAST qPCR master mix, ABI Prism; Kapa Biosystems, Cape Town, South Africa) following the manufacturer’s instructions on a PCR machine (StepOnePlus real-time; Applied Biosystems, Thermo Fisher Scientific, Johannesburg, South Africa) was performed as follows: 95°C for 5 min followed by 35 cycles of 95°C for 30 s and 60°C for 60 s. We visualized the PCR outcome using commercial software (StepOne software v.2.3; Thermo Fisher Scientific). PCR products with a cycle threshold value <25 were purified (illustra ExoProStar 1-step; GE Healthcare, Little Chalfort, Buckinghamshire, UK) and diluted in half with Tris-EDTA buffer (MilliporeSigma) before sequencing. A commercial sequencing kit (ABI Prism BigDye terminator v.3.1 cycle; Applied Biosystems) was used for the sequencing reactions with C and D tail primers using a quarter of the recommended concentration of BigDye terminator v.3.1. Sequencing products were purified with an ethanol precipitation and analyzed (3130xl genetic analyzer; Applied Biosystems).
Geneious v.8.0.4 (https://www.geneious.com) was used to assemble forward and reverse sequences, and the resulting consensus sequences were uploaded to the relevant PubMLST database (http://pubmlst.org/perl/bigsdb/bigsdb.pl?db=pubmlst_taylorella_seqdef&page=batchSequenceQuery) for strain type identification. No ambiguous sequencing results were observed.
A single sequence type (ST4) was identified from all South African samples, with 34 of 36 (94%) obtained from male and 2 of 36 (6%) from female horses (Table 1). Sample distribution by breed was as follows: Lipizzaner (23), Warmblood (10), Holsteiner (1), Andalusian (1), and Thoroughbred (1).
Taylorella equigenitalis isolates (n = 36) obtained from crude extract of samples (genital swabs, n = 34; frozen semen samples, n = 2) collected in South Africa in 1996–2017. All samples had an identical allelic profile* (1-2-5-3-2-2-3) and strain type (ST4).
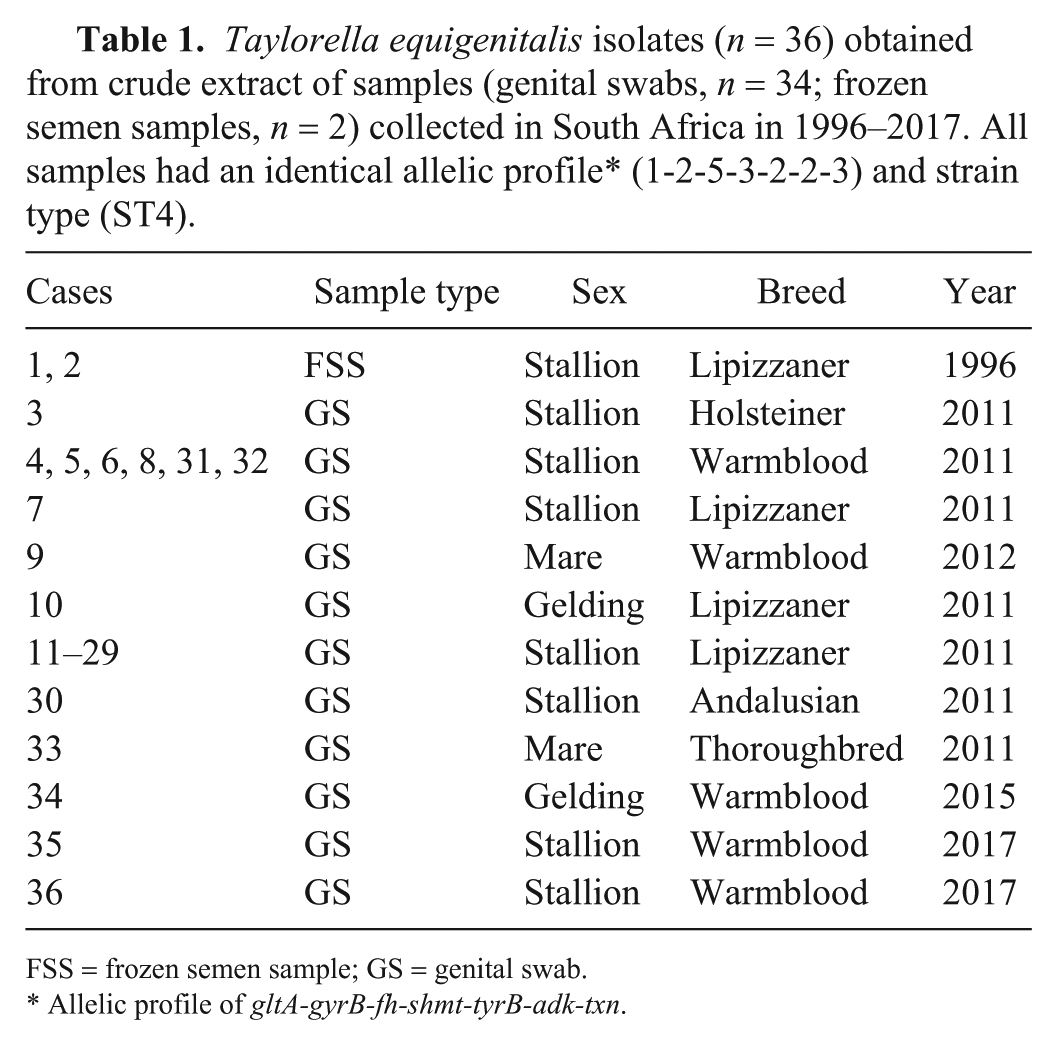
FSS = frozen semen sample; GS = genital swab.
Allelic profile of gltA-gyrB-fh-shmt-tyrB-adk-txn.
The successful outcome of our study using MLST analysis for T. equigenitalis supported the utilization of crude extracts direct from clinical samples in cases in which bacterial isolates are unavailable. The samples examined in our study were from 14 of 15 properties at which T. equigenitalis was identified. However, evidence of the point introduction, the epidemiologic link between cases, and an identical ST of samples isolated from all other cases, strongly suggested that this remaining property would most likely have been associated with the same ST4.
A study reporting genotyping of German and Austrian T. equigenitalis isolates using repetitive extragenic palindromic (REP) PCR and pulsed-field gel electrophoresis (PGFE) included a South African Holsteiner stallion isolate, which was the index case (case 3) in our study and which was allocated the REP/PGFE genotype rep-E1/TE-A5. 17 In that report, 17 rep-E1/TE-A5 was present exclusively in the majority of Austrian Lipizzaners (n = 38) and additionally was identical to the TE011 genotype reported from 4 stallions imported from Austria and Germany as part of the U.S. outbreak from 1978 to 2010. 1 These data show that ST4, rep-E1/TE-A5, and TE011 represent a single T. equigenitalis type. These findings clearly support both a common source and breed of origin associated with the South African outbreak.
The epidemiologic findings during the initial outbreak investigation suggested a point introduction of T. equigenitalis into South Africa prior to outbreak identification in May 2011. 13 It also indicated that current stringent pre- and post-entry CEM importation screening procedures of all equids entering the country, implemented in the early 1980s, have to date proved to be an effective barrier against the reintroduction of T. equigenitalis into South Africa.
We have shown that MLST assays for T. equigenitalis can be successfully performed on a crude extract originally prepared for PCR assays, negating the need for prior culture. This finding has important potential benefits for future large-scale investigation of CEM outbreaks in which organism identification is time-sensitive or isolates are not available. The analysis of MLST results from South Africa over a 21-y period showed the presence of a single ST, namely ST4, strongly suggesting a common, as yet unconfirmed, point introduction.
Footnotes
Acknowledgements
We thank Susan Miller and the staff of the Veterinary Genetics Laboratory, Faculty of Veterinary Science, Onderstepoort, for assistance with data analysis.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
