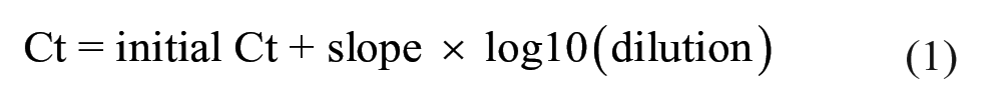Abstract
A sampling technique has been validated to monitor porcine reproductive and respiratory syndrome virus 2 (PRRSV-2) using the serosanguinous exudate known as processing fluids (PFs) that accumulate from tissues obtained during tail docking and castration. PFs are an aggregate sample of large numbers of piglets and litters. However, little is known about the effect of litter aggregation on the ability of PCR to correctly classify an aggregated PF sample as positive. We evaluated both the effect of litter aggregation and of PF pooling on PCR detection. We estimated that aggregation of at least 50 litters was possible when a pig with a Ct value of ~22 was present in the sample, and aggregation of up to 40 litters was possible when there was a sample with a Ct value of ~33. Pooling did not affect PCR detection when initial Ct values of 20 and 25 were assessed. However, in litters with initial Ct values of ≥30, the amount of pooling should be reduced. Our results provide producers and practitioners with a general framework to interpret more accurately the results of their PRRSV-2 surveillance programs using PF.
Keywords
Porcine reproductive and respiratory syndrome virus 2 (PRRSV-2; family Arteriviridae, subfamily Variarterivirinae, genus Betaarterivirus, subgenus Ampobartevirus, species Betaarterivirus suid 2) causes respiratory and reproductive disease in swine. 3 Producers who are controlling or eliminating PRRSV from breeding herds monitor virus presence through different approaches. A new sample type known as processing fluids (PFs), which includes the serosanguineous exudate that accumulates from tissues obtained during piglet processing activities (i.e., tail docking and castration), has been deemed suitable to monitor swine herds for PRRSV infection. 5 This sample type has become popular given the ease of collection and its high herd sensitivity at lower cost compared with blood sampling piglets individually at weaning. 2 PFs allow aggregation of large numbers of piglets and litters while still being able to detect the virus. 2 Collection of PFs involves collecting all tails and testes from all processed litters in a container bag that is sent as a laboratory submission. PFs from the bags can be tested individually, aggregated by day of processing, room, or as an aggregate weekly sample.
Pooled serum samples are commonly used to test for PRRSV in order to reduce the cost of testing while maintaining acceptable sensitivity. 4 Although by using PFs more animals can be sampled at a low cost, the effect on test sensitivity of aggregating large numbers of litters and pooling various PF samples is largely unknown. Therefore, our objectives were to investigate the limit of detection in aggregated PF samples (study 1), and to evaluate the dilution effect of pooling PF on initial cycle threshold (Ct) values (study 2).
As part of study 1 to evaluate the limit of detection of aggregating PF from multiple litters, 40 piglets from a recently infected breeding herd were sampled immediately after processing (~3–5-d-old) by placing docked tails and testes into individual round-bottom polystyrene tubes (Corning, Corning, NY) and stored at 4°C. Additionally, both the tail and scrotum incisions were swabbed using a sterile swab (Copan, Murrieta, CA) to assess the pig PRRSV status at processing by PRRSV reverse-transcription PCR (RT-PCR). Nucleic acid extraction was performed (MagMAX kit for whole blood samples; Applied Biosystems, Waltham, MA), and then RT-PCR was performed on nucleic acid extracts (VetMAX NA and EU reagents; Applied Biosystems). Three PRRSV-positive piglets were selected based on the results of the incision sample: 1 male with a Ct value <25 (low Ct group) and 2 (male and female) with a Ct value >25 (high Ct group). A female with a Ct value <25 was not found. The tail from the selected positive female and tails and testicles from the 2 selected positive males were placed in separate plastic bags containing processing tissues from 10 PRRSV-negative litters (~120 piglets) obtained from a PRRSV-free herd. These 3 bags were stored at 4°C for 24 h after mixing the tissues. After this time, 4 mL of PF were extracted for RT-PCR testing, and then the processing tissues from 10 additional PRRSV-negative litters were added to each bag. The process was repeated until each bag contained processing tissues from 50 litters (Fig. 1). The maximum number of litters was selected as an approximation of the maximum number of litters that are processed during a day in a breeding herd.
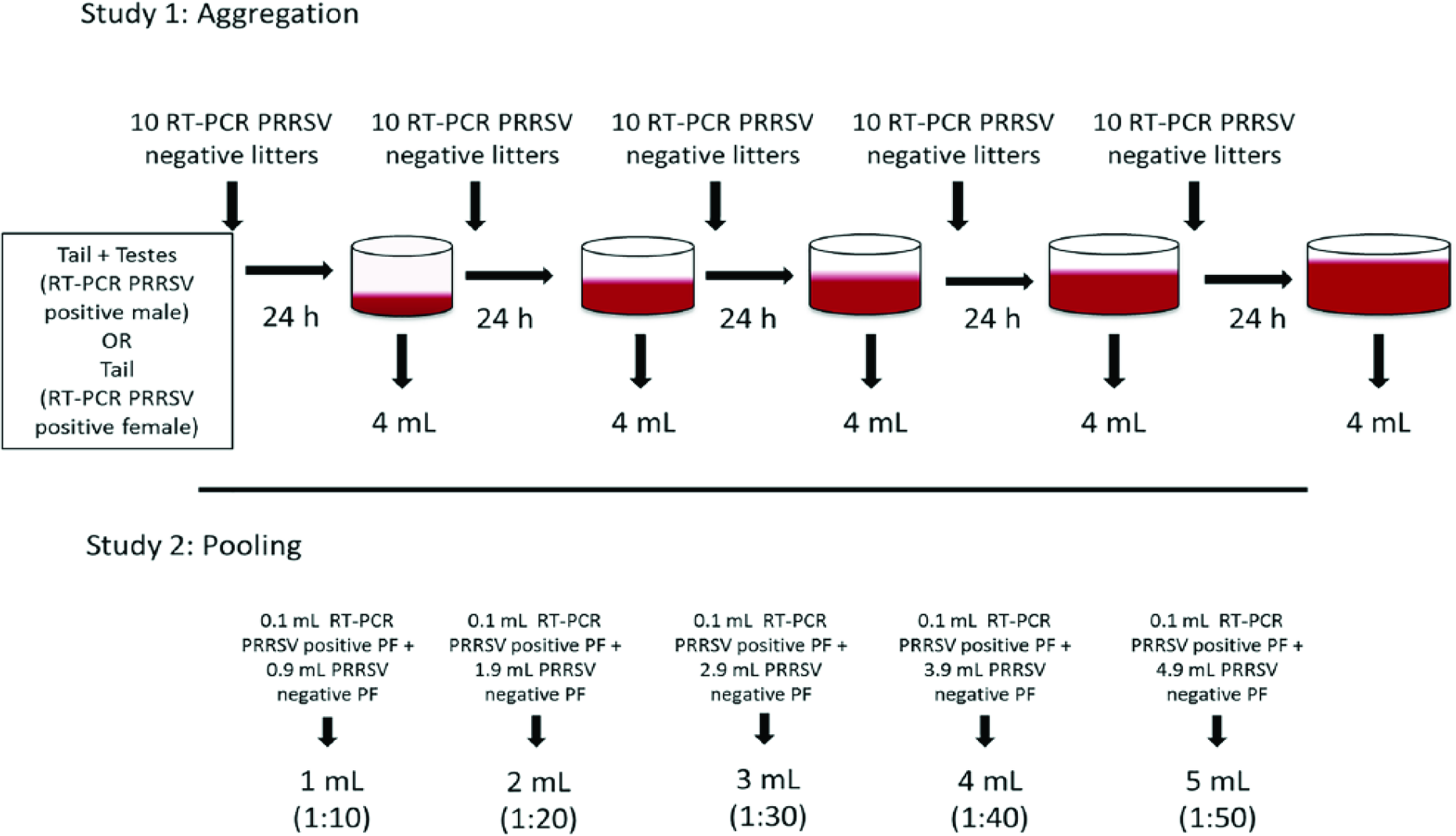
Diagram of studies evaluating the effect on detection of porcine reproductive and respiratory virus (PRRSV) of aggregation and pooling of processing fluid (PF) samples.
In order to evaluate the dilution effect of PF pooling on initial Ct values, 8 PRRSV RT-PCR–positive PF samples with Ct values of 15–30 were selected from a previous study. 5 Nucleic acid was extracted and analyzed as described in study 1 (MagMAX and VetMAX; Applied Biosystems). Each selected sample was then diluted 10, 20, 30, 40, and 50 times using PRRSV RT-PCR–negative PF samples, and sample dilutions were tested by RT-PCR (Fig. 1). The Mann–Whitney test was performed to assess whether the Ct value decrease at the highest dilution (1:50) was different between samples that prior to dilution had an initial Ct value <25 or an initial Ct value >25. A linear model was built for each sample to appreciate the individual dilution slopes. Additionally, a generalized linear mixed model was used to estimate the increase in Ct values (slope) by each unit of dilution increase in all samples. Dilutions (1:10, 1:20, 1:30, 1:40, 1:50) were converted to the base 10 logarithmic scale (1, 1.3, 1.47, 1.6, 1.69, respectively) prior to inclusion in the model. The upper 95% confidence interval (CI) of the dilution slope was used as a worst-case scenario of the dilution effect. Sample ID was included in the model as a random intercept and the dilution as a random slope. The worst-case scenario was used to estimate the increase in Ct value after diluting 10, 20, 30, 40, 50, 100, and 1,000 times an initial Ct value of 20, 25, 30, and 35 (Equation 1). Additionally, the maximum number of dilutions that can be done before reaching Ct values of 35 (suspect threshold) and 40 (negative threshold) (Equation 2) was estimated.
In study 1, PRRSV could be detected in PF when 1 PRRSV-positive pig was present in the aggregate sample of 50 litters or ~600 pigs (Table 1). However, PRRSV was only detected consistently when the Ct value of the positive pig (male) was low (<25), indicating more viral particles in the sample.
Summary of porcine reproductive and respiratory syndrome virus processing fluid RT-PCR cycle threshold results following tissue aggregation of 1 positive piglet with 10, 20, 30, 40, or 50 PRRSV-negative litters.
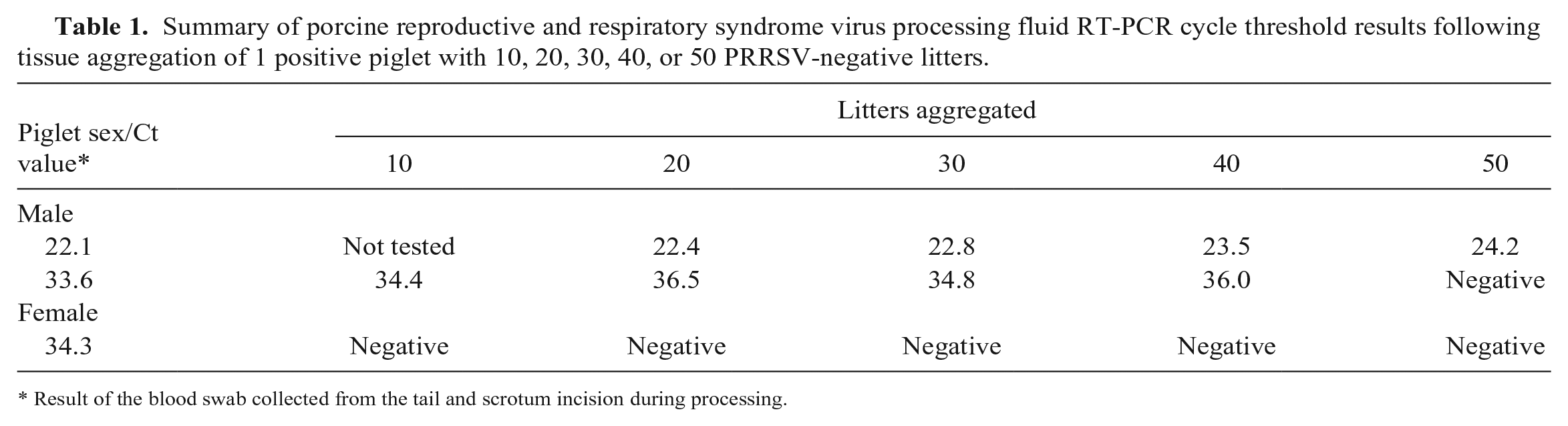
Result of the blood swab collected from the tail and scrotum incision during processing.
In study 2, an increase of 4.6 ± 0.7 Ct (mean ± SD) from their initial Ct value was observed after the initial 10-fold dilution. The mean increase between Ct values of the 1:10 and 1:50 dilution was 1.4 ± 0.9. Ct increase from the initial values to the ones of the last dilution (1:50) was not statistically different between groups with initial Ct values considered as low (<25) or high (≥25; Mann–Whitney p = 0.89). Individual slopes were 2.94–4.11 for the 9 samples used for pooling, and the calculated worst-case Ct value slope was 3.7 for each log10 unit of increase (Table 2, Fig. 2).
Linear regression values (slope and intercept), standard error (SE), and R2 for each set of samples and for the model (linear mixed effect model considering sample as a random effect).
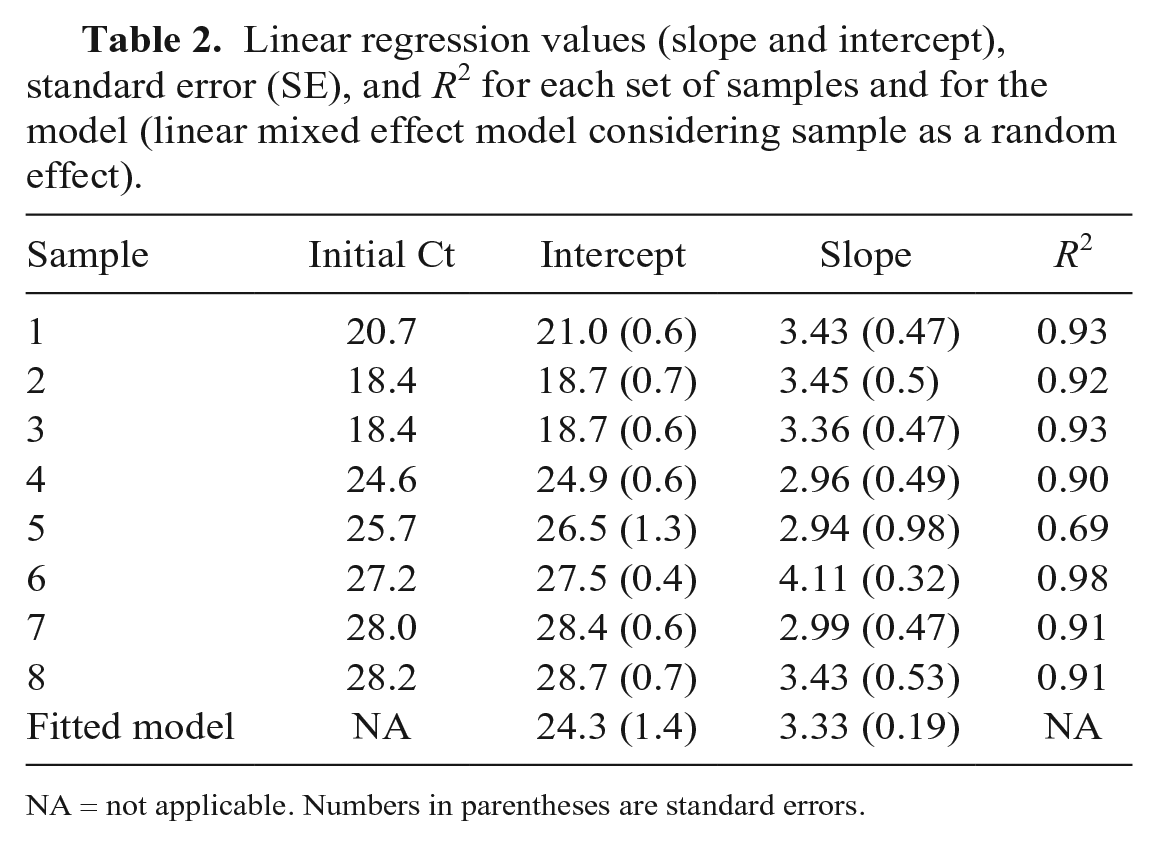
NA = not applicable. Numbers in parentheses are standard errors.
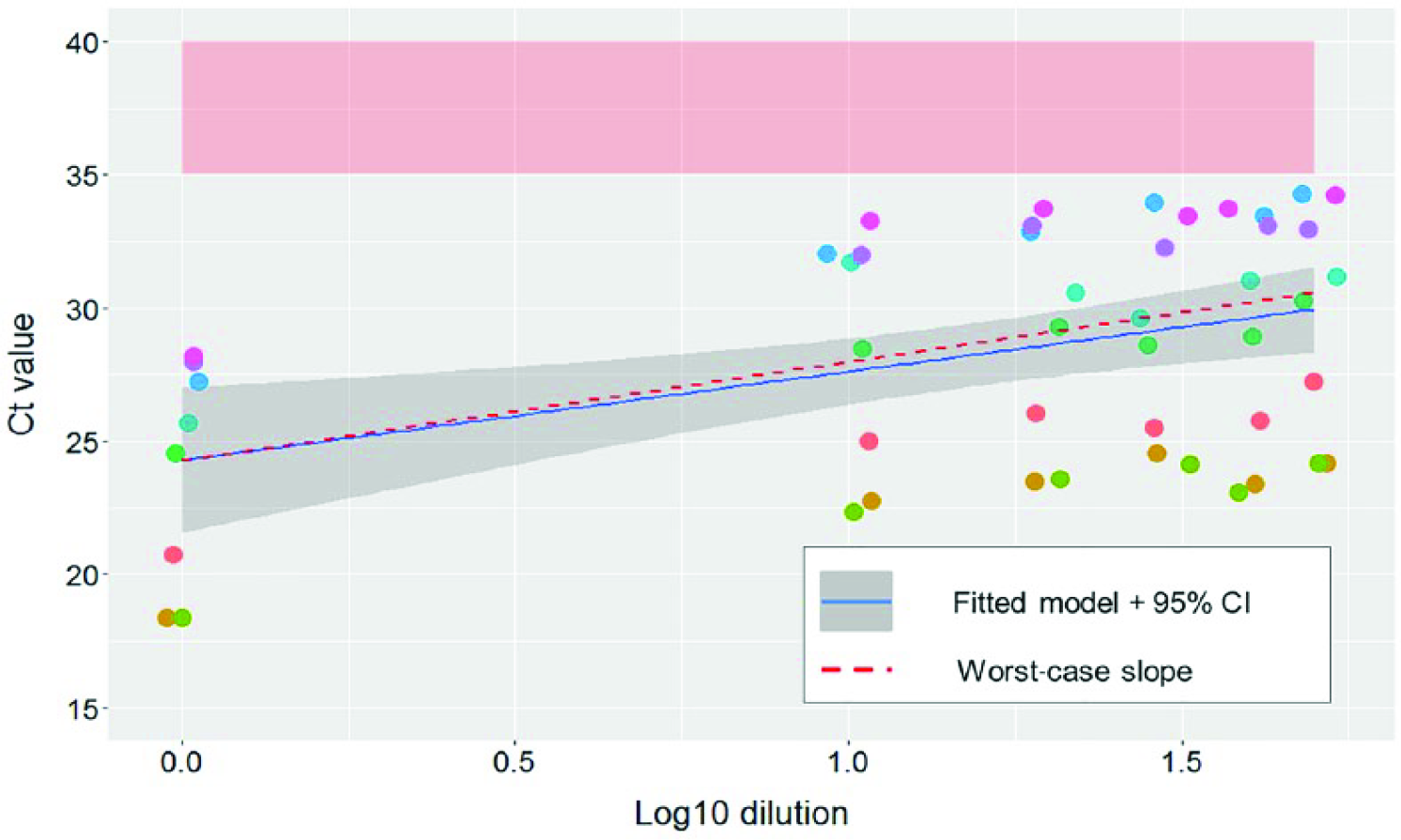
Reverse-transcription PCR (RT-PCR) cycle threshold (Ct) values of pooled processing fluid samples and predicted values based on best fit and worst-case scenarios. Each set of dilutions is shown in a different color. The RT-PCR suspect area (35–40 Ct) is shaded in light red. Fitted linear regression model and 95% confidence interval (CI; continuous blue line and gray area) and the worst-case slope (dashed red line) are also represented. Dilutions of 1:1. 1:10, 1:20, 1:30, 1:40, and 1:50 are represented in the log10 scale (0, 1, 1.3, 1.47, 1.6, and 1.69, respectively).
The maximum number of dilutions resulting in a RT-PCR–positive result (Ct ≤ 35) was highly variable according to the initial Ct value of the samples (Table 3). For samples with an initial Ct value of 20 or 25, a large number of dilutions was possible without losing sensitivity. Conversely, an initial Ct value of 30 would fall above the suspect threshold after 22 dilutions. Additionally, 40% of the time, initial Ct values of 30 will fall above the suspect threshold after 20 dilutions. For initial Ct values of 35, 95.9% of the time values will be under the Ct of 40 after 10 dilutions but only 60.5%, 25.3%, 11.5%, and 4% of the time after 20, 30, 40, and 50 dilutions, respectively.
Predicted average cycle threshold (Ct) values in different dilutions according to the initial Ct values using the estimated worst-case scenario slope.
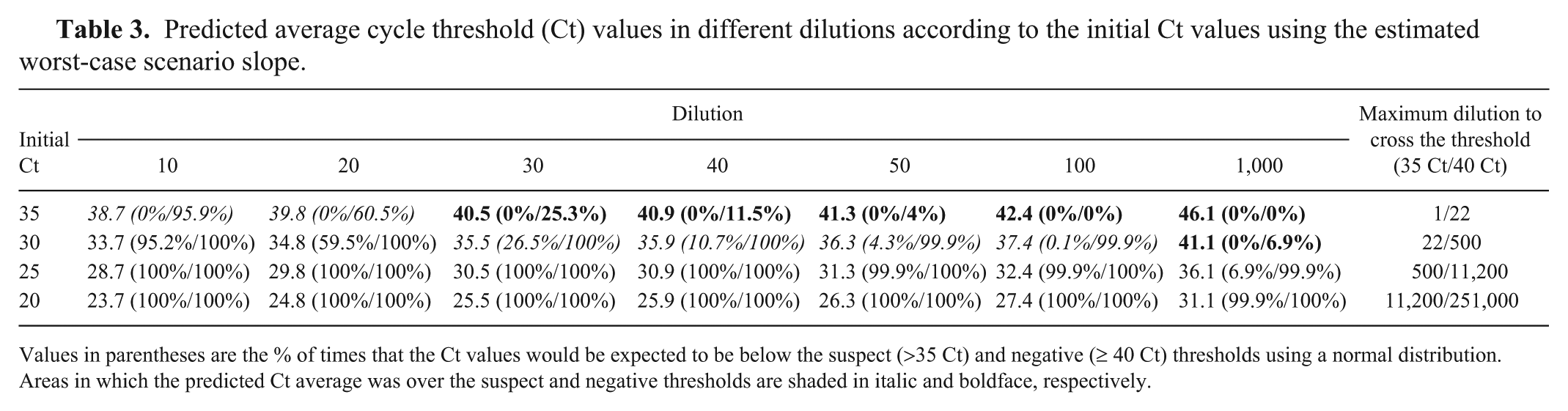
Values in parentheses are the % of times that the Ct values would be expected to be below the suspect (>35 Ct) and negative (≥ 40 Ct) thresholds using a normal distribution. Areas in which the predicted Ct average was over the suspect and negative thresholds are shaded in italic and boldface, respectively.
The use of PF to evaluate the PRRSV status of herds has been increasingly adopted among U.S. veterinarians and producers. Understanding the capabilities of this sampling methodology is vital for designing monitoring strategies that optimize virus detection while maintaining sensitivity at a low cost. Detection of PRRSV in aggregated samples is dependent on the viral load of the positive pigs contributing to the PF sample; thus, the higher the viral load (i.e., the lower the Ct) of an individual pig, the higher the probability of detection. The viral load in serum and PF samples is higher after a recent PRRSV outbreak and decreases over time, unless a subsequent PRRSV infection occurs. 5 Thus, it would seem reasonable to aggregate more litters during the initial stages of infection as the viral load per pig and the number of positive pigs is high, and reduce the aggregation a few months after the original PRRSV outbreak when it will become more difficult to detect the virus. In our study, it was possible to aggregate up to 50 litters (~600 piglets) when there was a single male pig with a Ct value of 22 in the sample. This is in agreement with the findings of a previous study that used a different methodology to collect the PF. 2 In that study, researchers were able to detect the virus in PF of an aggregated sample of 650 pigs 21 wk after a PRRSV outbreak had occurred. However, PRRSV was not detected when the processing tissue (i.e., only docked tail) of a female pig was used as the source of virus in the aggregate sample. A decrease in sensitivity was expected because tails contribute less fluid to the aggregate sample compared to testes.
Pooled serum samples have been used in the industry as a way to decrease testing costs. 4 In our study, the maximum number of dilutions possible before crossing the suspect or negative threshold varied depending on the initial Ct value, particularly for initial Ct values >30. For Ct values <30, a large number of dilutions is possible without losing PCR sensitivity. Similar results were found in another study in which detection was possible after five 10-fold dilutions when an initial sample with a 17 Ct value was used. 1
A limitation of our study is that it was conducted with only 2 strains of PRRSV-2 from 2 farms that were recently infected with PRRSV. Further studies should be done to compare the dilution effect on different strains or genotype (e.g., type 1).
Footnotes
Acknowledgements
We thank Aaron Rendahl, Albert Rovira, and My Yang for their valuable contribution in the methods and the analytical part of this manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was partially funded by the Awards Advancing Research in Respiratory Disease from Boehringer Ingelheim Vetmedica, and the Swine Disease Eradication Center of the University of Minnesota.