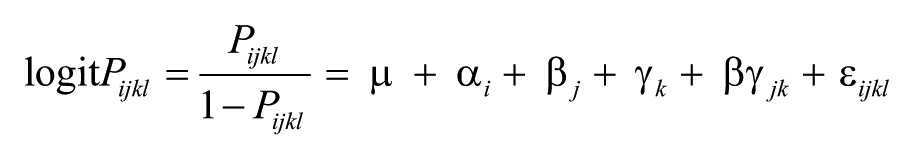Abstract
We evaluated effects of handling procedures on detection of porcine reproductive and respiratory syndrome virus (PRRSV) in oral fluids (OFs) by reverse-transcription real-time PCR (RT-rtPCR). The experiments were conducted using a composite sample of PRRSV-positive OF collected from 5-wk-old pigs vaccinated 15 d earlier with a modified-live PRRSV vaccine. Five pre-extraction sample-handling steps and all combinations thereof were evaluated: 1) thaw temperature (4°C or 25°C); 2) sample diluent (1:1 dilution with nuclease-free water or guanidinium thiocyanate–phenol); 3a) sonication of the sample (yes or no); 3b) temperature (4°C or 25°C) at which step 3a was conducted; and 4) temperature at which the sample was maintained after step 3b and until RNA extraction was initiated (4°C or 25°C). All combinations of the 5 sample-handling steps (i.e., 32 unique treatments) were tested in a completely randomized factorial design with 4 replicates and 1 negative control for each treatment. The entire experiment was repeated on 5 separate days to produce a total of 800 PRRSV RT-rtPCR results. Binary (positive or negative) data were analyzed by logistic regression and results (Ct) were analyzed using a generalized linear model. Overall, 1 false-positive result was observed among 160 negative controls (99.4% specificity), and 85 false-negative results were observed among the 640 known-positive samples (86.7% sensitivity). The most significant factor affecting test outcome was thaw temperature (4°C or 25°C); samples thawed at 4°C had higher positivity rate (94% vs. 80%, p < 0.0001) and lower Ct (36.2 vs. 37.5, p < 0.0001).
Introduction
Serum samples are the traditional antemortem sample in veterinary medicine, but swine oral fluid (OF) samples are increasingly used to surveil for a variety of pathogens using either antibody or nucleic acid–based assays. Thus, the cumulative number of tests performed on OF samples at the veterinary diagnostic laboratories at Iowa State University, University of Minnesota, and South Dakota State University increased from ~21,000 in 2010 to ~370,000 in 2016. 2 The increased use of OF samples by swine producers and veterinarians reflects the fact that such samples are collected without stress to pigs or people and, when used with assays optimized for OF, provide a higher population-level probability of target analyte detection than individual pig serum samples.12,16
The detection of porcine reproductive and respiratory syndrome virus (PRRSV) in swine OF was reported in 2008.13,14 Using reverse-transcription real-time PCR (RT-rtPCR), PRRSV was detected in serum for ~5 wk and in OF for ~4 wk in pigs inoculated with PRRSV-2 under experimental conditions. 14 Independent testing of OF samples at 2 laboratories showed 88% agreement in RT-rtPCR results, thereby affirming the validity of the OF-based approach for PRRSV detection while simultaneously suggesting the need for improved assay reproducibility. Later work reported that the detection of PRRSV in OFs by RT-rtPCR varied among extraction and PCR protocols and suggested that further research was required to optimize PCR assays for the OF matrix. 5 Research focused on technical aspects is crucial and will lead to improvements in assay performance, but factors external to nucleic acid extraction and PCR performance procedures may also affect test outcomes. Therefore, we examined the effect of specific pre-testing procedures on the detection of PRRSV RNA in swine OFs by RT-rtPCR.
Materials and methods
Experimental design
Five pre-extraction sample-handling steps (1, 2, 3a, 3b, 4) and all possible combinations (n = 32) were evaluated for their effect on PRRSV RT-rtPCR results: 1) thaw temperature (4°C or 25°C); 2) sample diluent (i.e., 1:1 dilution with nuclease-free water [NFW]; UltraPure DNase/RNase-free distilled water, Life Technologies, Carlsbad, CA) or guanidinium thiocyanate–phenol (GTP) reagent (Trizol LS, Life Technologies); 3a) sonication of the sample (yes or no); 3b) temperature at step 3a (4°C or 25°C); and 4) temperature at which the sample was maintained after 3b and until RNA extraction was initiated (4°C or 25°C; Fig. 1). The time between the end of step 1 (thaw) and the initiation of RNA extraction was always <4 h.
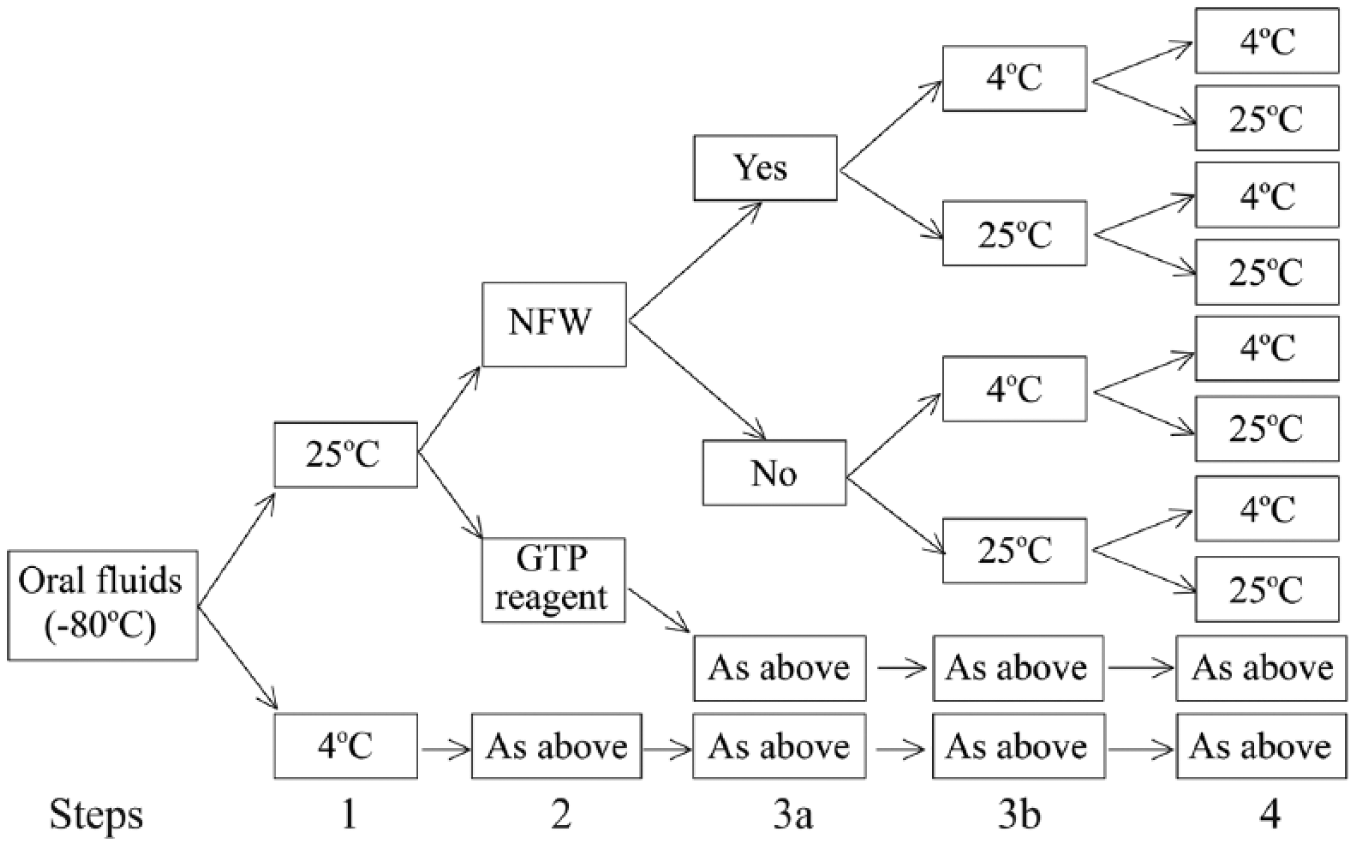
Process chart for evaluation of the effect of specific sample-handling steps on the detection of porcine reproductive and respiratory syndrome virus RNA in oral fluid specimens:
All combinations of the 5 steps produced 32 unique treatments, which were tested in a completely randomized factorial design with 4 PRRSV-positive OF samples and 1 negative control (NFW or GTP reagent) randomly assigned to each treatment. The entire experiment was repeated 5 times (once each week for 5 wk) to produce 800 PRRSV RT-rtPCR results. The effect of sample-handling treatments on PRRSV RT-rtPCR results was analyzed using logistic regression for binary outcomes (positive or negative), and a generalized linear model for numeric results (cycle threshold [Ct]).
Oral fluid
A composite sample (2.2 L) of PRRSV-positive OF was collected from ~250 five-wk-old pigs vaccinated 15 d earlier with a modified-live PRRSV vaccine (Ingelvac PRRS MLV, Boehringer Ingelheim Vetmedica, St. Joseph, MO). OFs were collected as described previously using 1.6-cm (5/8 in), 3-ply, 100% cotton rope (Web Rigging Supply, Lake Barrington, IL) suspended to pig shoulder height.13,17 The procedure was approved by the Iowa State University Office for Responsible Research (IACUC 3-14-7763-S). Following collection, the composite sample was dispensed into 25-mL tubes that were stored at ⩽ –70°C.
Sample-handling procedures
Each time the experiment was repeated, 4 aliquots of OF and 1 negative control (NFW or GTP reagent) were tested in each treatment (n = 32 treatments; 32 × 5 samples per treatment = 160 samples). The experiment was repeated 5 times to produce 800 RT-rtPCR results (i.e., 640 from PRRSV-positive aliquots of OF and 160 from negative controls).
Prior to performing the experiment, 5-mL polypropylene tubes (BD Falcon, BD Biosciences, San Jose, CA) were labeled with a treatment ID (1–32) and a random number (1–160; Randomness and Integrity Services, Dublin, Ireland). Treatment ID was used to track tubes through the 5 steps of their assigned treatment, and their random number was used to randomly order the tubes after treatment had been completed and prior to PRRSV RT-rtPCR testing. In step 1, four 25-mL tubes of OF were moved from storage (⩽ –70°C); 2 were placed in a 4°C environment (Caron Products, Marietta, OH) and the other 2 in a 25°C environmental chamber (Caron). After 24 h, 0.75 mL was aliquoted from the 25-mL tubes into the pre-labeled 5-mL polypropylene tubes (BD Falcon). In step 2, aliquots were diluted 1:1 with either NFW or GTP reagent to a total volume of 1.5 mL and inverted 3 times at room temperature. One negative control per treatment (1.5 mL of the diluent used in the treatment) was created at this step. In steps 3a and 3b, aliquots were sonicated for 10 min at 4°C or 25°C in a 40-kHz water bath sonicator (Sonicator B-series 2510, Branson Ultrasonics, Emerson Industrial Automation, Danbury, CT). If not sonicated, samples were simply held for 10 min at 4°C or 25°C. In step 4, aliquots were maintained at 4°C or 25°C until the viral RNA extraction procedure was initiated (1–4 h). Temperature was monitored throughout the process using NIST-certified thermometers (Traceable waterproof thermometer, Thermo Fisher Scientific, Waltham, MA).
Nucleic acid extraction protocol
Testing was conducted at the Iowa State University–Veterinary Diagnostic Laboratory (ISU-VDL) using procedures performed routinely for OF specimens. Total RNA extraction was performed using a single lot of a magnetic bead–based commercial kit (MagMAX viral RNA isolation kit, Life Technologies). Two positive controls and one negative control were included with each extraction.
Samples were processed using the “high-volume” extraction and “high-throughput method” lysate preparation (MagMAX viral RNA extraction, ISU-VDL SOP 9.459). The modified lysis/binding solution for the extraction protocol was prepared using 45 mL of lysis/binding solution with 200 µL of carrier RNA without the addition of isopropanol. For the lysis step, 450 µL of modified lysis/binding solution followed by 300 µL of sample was added to the wells of a 96 deep-well plate (Polypropylene microtiter microplates, Thermo Fisher Scientific). Following the addition of the sample, the plates were sealed, mixed (Titer plate shaker, Thermo Fisher Scientific) for 3 min, and then centrifuged at 2,500 × g for 6 min. A volume of 600 µL of lysate was transferred to the wells of a plate containing 300 µL of isopropanol and 20 µL of magnetic bead mix. The plates containing the samples were loaded onto a semi-automated nucleic acid purification system (KingFisher 96, Thermo Fisher Scientific) along with 2 plates of 300 µL of wash solution 1, 2 plates of 450 µL of wash solution 2, and 1 plate containing 90 µL of elution buffer. Extraction was then completed on the semi-automated system by using program AM1836_DW_HV_v3 (Applied Biosystems, Foster City, CA). Thereafter, the elution plate was sealed using a plate sealer until assayed by PRRSV RT-rtPCR.
PCR protocol
PRRSV RT-rtPCR was performed using a single lot of a commercial kit (EZ-PRRSV MPX 4.0 assay, Tetracore, Rockville, MD). PCR reactions were performed in 96-well fast plates (MicroAmp Fast optical 96-well reaction plate with barcode, Applied Biosystems) with component volumes per well including 17.25 µL of reagent (including buffer, primer, and probes), 0.75 µL of enzyme blend, and 7 µL of extracted OF sample. Extraction controls (negative and positive), internal control (EZ-PRRSV inhibition control, Tetracore), and amplification controls (negative, PRRSV-1 and -2) were run on each RT-rtPCR plate. Plates were shaken on a vortex mixer for 10 s and then centrifuged before being loaded onto the thermocycler (7500 Fast real-time PCR system, Applied Biosystems). The assay covered 2 target regions of the PRRSV-1 and -2 genes with primers and probes. The reaction was completed using the following thermocycling conditions: 1 cycle at 48°C for 15 min, 1 cycle at 95°C for 2 min, 45 cycles of 95°C for 5 s, and 60°C for 40 s. PRRSV-2 was detected via 6-carboxyfluorescein (FAM) as a reporter dye, and carboxy tetramethylrhodamine (TAMRA) alternative as a reporter dye for PRRSV-1. Cyanine 5 alternative reporter dye was utilized for the detection of the extraction/inhibition control. To ensure the validity and repeatability of RT-rtPCR results, positive control Ct values were continuously monitored using statistical process control charting. Plates with positive control Ct values outside the acceptable range were considered invalid and the samples re-tested. On valid plates, samples with Ct values <40 for either genotype were considered positive.
Statistical analysis
Data were analyzed (SAS v.9.3, SAS Institute, Cary, NC). The experimental design allowed for direct comparison of the effect of all steps and treatments on PRRSV RT-rtPCR performance. The effect of repetition (day), replication (within treatment), treatment, and their interactions on PRRSV RT-rtPCR performance were considered in the analyses. Nonsignificant effects (p > 0.05) were excluded from the final models.
For statistical analysis, only results of the known-positive samples were used. Binary test outcomes (positive, negative) were analyzed using logistic regression, and numeric outcomes (Ct values) were analyzed by a generalized linear model. Because Ct values were truncated (i.e., negative samples are represented as ⩾40), Ct values were only evaluated using known-positive OF samples with a positive result (i.e., Ct values <40).
Results
All PRRSV RT-rtPCR plates met quality control criteria, and the testing results were accepted. Defining a positive result as a Ct value <40, RT-rtPCR testing of 640 PRRSV-positive samples produced 85 false-negative results (86.7% sensitivity) and testing of 160 PRRSV-negative samples (controls) produced 1 false-positive result (99.4% specificity). Statistical comparison found no difference in the detection rate among repetitions (days).
Logistic regression analysis identified step 1 (thaw temperature) as the only step with a significant effect (p < 0.0001) on positivity, with more false-negative results produced when samples were thawed at 25°C than 4°C (80% vs. 94% positives; Table 1). The logistic regression model derived from the final model produced the following equation:
where Pijkl = probability of a PCR-positive result, μ = intercept, i = 2 levels of step 1, j = 2 levels of step 2, k = 2 levels of step 3b, and l = repeated observations.
Logistic regression analysis of effects of handling of 640 known-positive oral fluid samples on porcine reproductive and respiratory syndrome virus reverse-transcription real-time PCR qualitative outcomes (positive, negative).
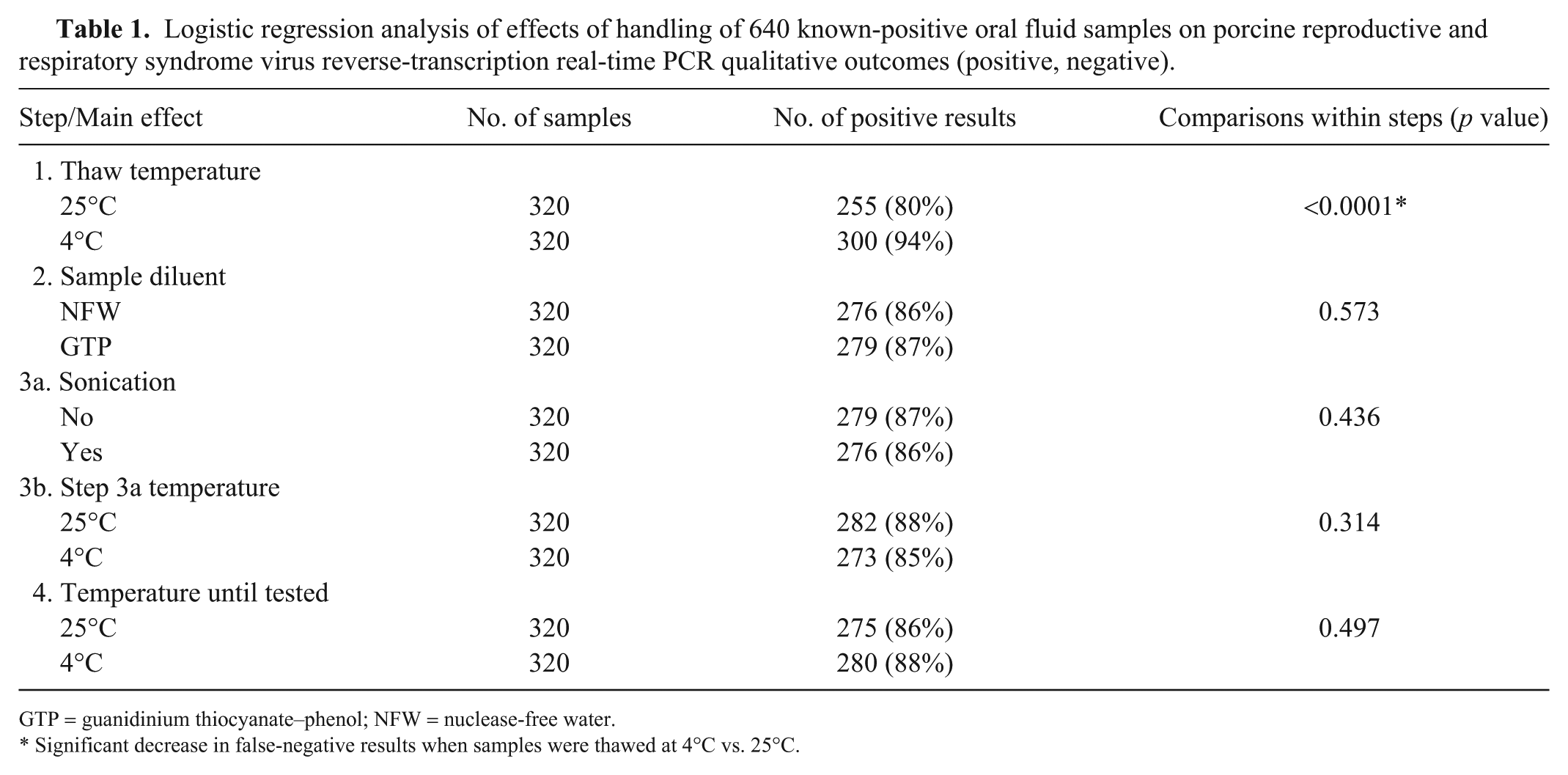
GTP = guanidinium thiocyanate–phenol; NFW = nuclease-free water.
Significant decrease in false-negative results when samples were thawed at 4°C vs. 25°C.
Although steps 2 (diluent) and 3b (4°C or 25°C at step 3a) were not significant individually, they were included in the final model because of a significant interaction between the 2 steps (p = 0.0428). Specifically, the combination of steps 2 and 3b produced the following detection results: NFW at 4°C (140 of 160 = 88%), NFW at 25°C (136 of 160 = 85%), GTP at 4°C (133 of 160 = 83%), and GTP at 25°C (146 of 160 = 91%).
Statistical analysis showed that step 1 (thaw temperature) impacted test performance (p < 0.0001), with lower Ct values in samples thawed at 4°C than 25°C (mean Ct 36.2 vs. 37.5, respectively; Table 2). Step 2 (diluent) was also significant (p = 0.0006), with lower Ct values observed in samples diluted with GTP than NFW (mean Ct 36.6 vs. 37.1, respectively). The linear model derived from the analysis of the outcomes (Ct results) produced the following equation:
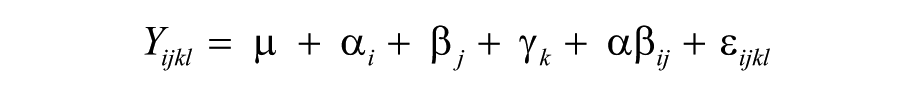
where Yijkl = PRRSV RT-rtPCR Ct value, μ = intercept, i = 2 levels of step 1, j = 2 levels of step 2, k = 2 levels of step 4, and l = repeated observations. As reflected in the final model, significant interactions were detected between steps 1 and 2 (p = 0.0165) and between steps 2 and 4 (p = 0.0271).
Effect of oral fluid sample-handling procedures on porcine reproductive and respiratory syndrome virus reverse-transcription real-time PCR outcomes (Ct values).
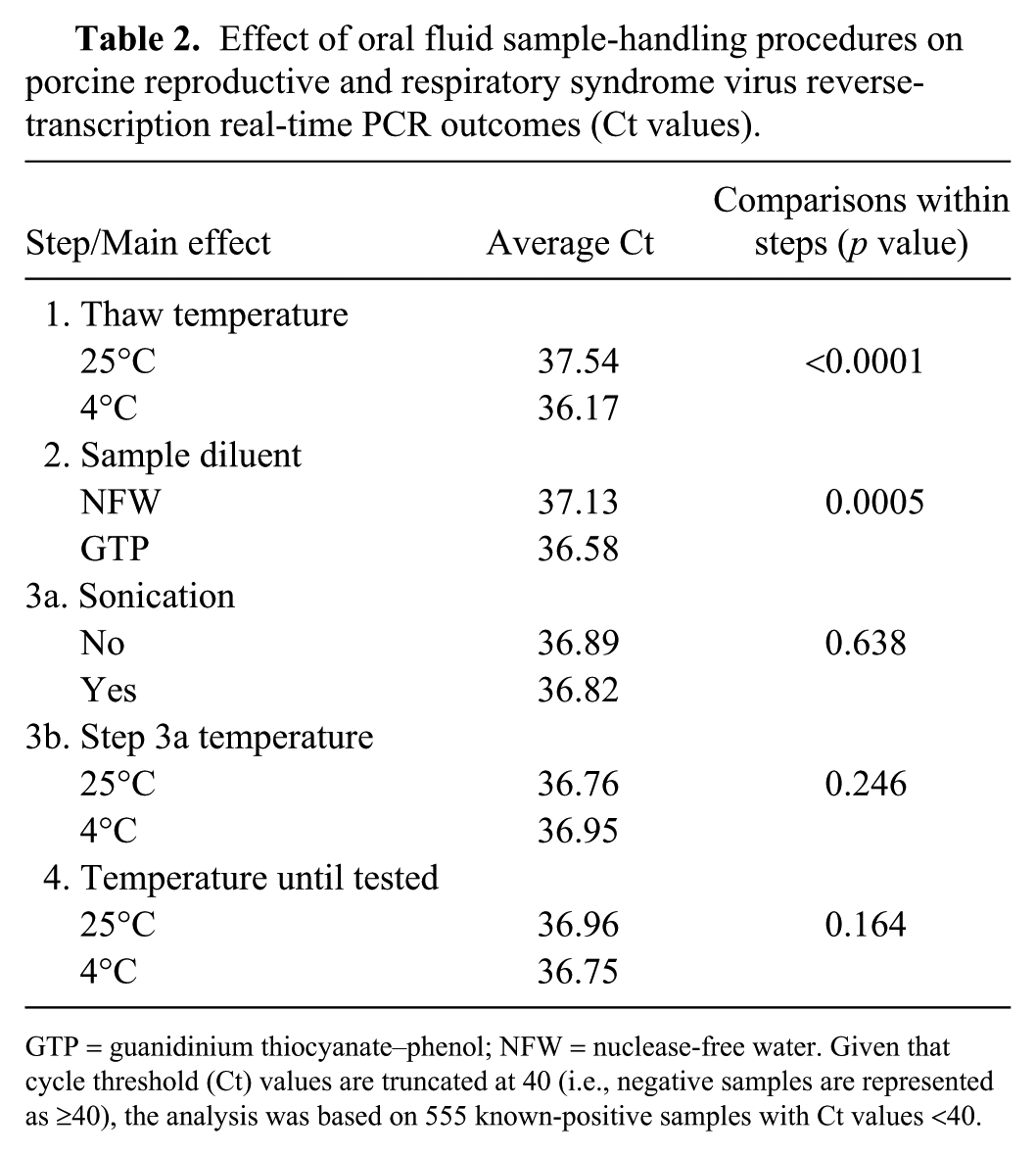
GTP = guanidinium thiocyanate–phenol; NFW = nuclease-free water. Given that cycle threshold (Ct) values are truncated at 40 (i.e., negative samples are represented as ⩾40), the analysis was based on 555 known-positive samples with Ct values <40.
Discussion
The physical effect of the freeze–thaw cycle on virus infectivity and/or nucleic acid detection has long been a topic of discussion, but published results from controlled studies are sparse. A 2017 study found that influenza RNA in Tris–EDTA buffer was detectable for up to 7 freeze–thaw cycles. 7 Similarly, 8 freeze–thaw cycles had minimal effects on hepatitis B virus DNA and hepatitis C virus RNA in serum specimens, leading to the conclusion that both DNA and RNA were resistant to degradation by freeze–thaw. 11 However, others have reported that freeze–thaw could produce false-negative PCR results, especially in serum specimens with low concentrations of RNA. 11 This could explain anecdotal reports of PCR-positive samples that subsequently re-test negative. Mechanisms for the degradation of nucleic acids by the thawing process are not known, but are postulated to involve microenvironmental changes in pH and/or oxidative damage catalyzed by ions in the sample. 3
The effect of ambient temperature on the stability of PRRSV RNA in OFs has been described.6,15 PRRSV RNA in OF has been reported to be stable at −20°C, 4°C, and 10°C for up to 12 d, whereas PRRSV RNA in OF samples stored at 20°C or 30°C declined significantly within 12 h. 15 Other workers evaluated the stability of PRRSV RNA in OF samples held at 4°C or 22°C and arrived at similar conclusions, noting that careful handling and storage procedures were especially important for samples with low concentration of RNA. 6 In our study, the temperature (4°C or 25°C) at which steps 3, 3a, and 4 were performed had no effect on RT-rtPCR performance. Presumably, the <4-h period between the end of step 1 and the initiation of RNA extraction did not provide sufficient time for detectable decay of RNA.
Among the factors evaluated, only step 1 had a significant effect on the detection of PRRSV RNA. In step 1, samples were moved from −70°C to 4°C or 25°C and held for 24 h, during which time the samples thawed and then equilibrated to ambient temperature. Unexpectedly, step 1 significantly affected test outcomes (i.e., samples at 4°C had a significantly higher rate of positivity compared to samples thawed at 25°C; p < 0.0001). Logically, this result should reflect the cumulative effect of thaw plus ambient temperature (discussed in the previous paragraph) on RNA degradation.
Several chemical additives (e.g., nucleic acid preservation buffers, ethanol, bacteriostatic compounds, and various commercial products) have been used to preserve RNA and DNA in clinical specimens.4,15 GTP, a chaotropic salt, is known to inhibit RNase, disrupt cell and organelle membranes, and denature proteins. Previous studies reported that GTP extended the infectivity and storage life of classical swine fever virus and foot-and-mouth disease virus RNA in harvested cell cultures and tissue samples. 8 Likewise, serum and peripheral blood mononuclear cell specimens frozen with Trizol reagent, a commercial formulation that contains GTP, showed significantly higher messenger RNA integrity when compared to specimens without Trizol. 9 However, GTP provided no benefit in the detection of PRRSV RNA in our study.
Sonication was included in the evaluation because sound waves have the potential to physically disrupt aggregates or particles in samples prior to extraction, thereby releasing target nucleotides. 10 For example, sonication of synovial fluid or periprosthetic tissue significantly improved the diagnostic sensitivity of a multiplex PCR for detecting Propionibacterium acnes and Staphylococcus spp. 1 In our study, the use of bath sonication provided no improvement in the detection of PRRSV RNA in OF samples.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This project was funded in part by the U.S. Department of Agriculture, National Institute of Food and Agriculture, Agriculture and Food Research Initiative Competitive Grants Program (project IOWV-ZIMM-2011-02975).