Abstract
Influenza A virus (IAV) is a zoonotic pathogen threatening animal and public health; therefore, detection and monitoring of IAV in animal populations are critical components of a surveillance program. Swine are important hosts of IAV, wherein the virus can undergo rapid evolution. Several methods (i.e., nasal swabs, nasal wipes, and oral fluids) have been used to collect samples from swine for IAV surveillance. We utilized nasal wipes made from cotton gauze and multiple, polyester or mixed polyester fabrics to compare performance in the molecular detection and isolation of IAV. In vitro experiments revealed that no polyester or mixed polyester fabric was superior to cotton gauze for molecular IAV detection; however, 3 polyester or mixed polyester fabrics yielded significantly more viable IAV than cotton. In a field trial, both cotton gauze and the polyester or mixed polyester fabric yielded similar proportions of IAV isolates from swine. The results indicate that cotton gauze remains a practical and useful material for swine nasal wipes.
Influenza A virus (IAV) causes disease in a variety of host species, with widespread implications for human health, including hospitalization and death. IAV outbreaks in swine populations not only result in loss of productivity, but can generate novel reassortant IAV strains with pandemic potential in humans.6,8 The detection of swine lineage IAV in people, termed variant IAV, continues to demonstrate the importance of improving surveillance and laboratory testing for IAV in swine to help protect public health. 2 Optimizing sampling methods for IAV surveillance in swine populations enables enhanced monitoring of IAV evolution and spread. Veterinarians, human health officials, agricultural professionals, and animal producers rely on the data generated during surveillance of animal populations to make decisions aimed at protecting public and animal health.
Several methods are used for the collection of antemortem samples for IAV testing of swine: nasal swabs, oral fluids, and nasal wipes. 4 Collection of synthetic-tipped nasal swabs is the preferred method because it permits sampling within the nasal cavity where active IAV shedding occurs, and thus is the gold standard collection method for individual pig testing. 9 However, nasal swabs are not always ideal for quick, noninvasive sample collection because pigs must be snared for safe handling and proper sample collection. Snaring can be difficult to perform and unpleasant to observe because pigs resist and vocalize. 4
Noninvasive collection of oral fluids is useful for detecting IAV in groups of pigs. However, oral fluid samples are most useful for molecular detection of IAV because recovery of live virus from swine oral fluids can be difficult despite testing positive for IAV using reverse-transcription PCR (RT-PCR). 3 In contrast, cotton gauze has been used successfully in the field as substrate for nasal wipes to sample swine for IAV at agricultural exhibitions, allowing detection with molecular testing and recovery of infectious virus in cell culture. 4 Cotton gauze is very flexible, absorbent, commercially available in sterile packaging, and can be used to wipe the inside of the nostrils of a pig with relative ease and no restraint. However, the lower sensitivity of cotton gauze nasal wipes versus synthetic-fiber nasal swabs for IAV detection 4 may be a result of chemical residues left after bleaching fibers in the production of medical-grade cotton.5,9
We attempted to identify additional nasal wipe substrates that may yield better detection and recovery of IAV from swine. Given that the gold standard nasal swab sampling method utilizes synthetic-tipped swabs, we investigated polyester fabrics as possible nasal wipe substrates. We tested 6 polyester and mixed polyester fabrics and compared them to cotton gauze by determining viable IAV recovery in cell culture and detection using reverse-transcription quantitative PCR (RT-qPCR). The fabrics were composed of polyester or various polyester and co-polyester blends with various physical properties such as thickness, absorbency, and softness.
Six fabrics (M140-calendar, 6194, M137, 5889, 6198, and 93T8) were kindly provided by Polymer Group (Charlotte, NC). All fabrics were composed of 100% polyester except for M140-calendar and M137, which were composed of a 93% polyester and 7% co-polyester blend. All novel fabrics were cut into 5 × 5 cm squares to replicate the size of cotton gauze and sterilized by gamma irradiation (25–50 kGy) prior to inoculation. Sterile, pre-packaged, 5 × 5 cm medical-grade cotton gauze (Covidien, Minneapolis, MN), the currently used sampling material, was included for comparison. Testing was performed in triplicate for each fabric. The fabrics were placed in sterile petri dishes and inoculated with 0.2 mL of stock cell culture supernatant containing a total inoculum of 4.04 × 107 tissue culture infectious dose 50% (TCID50) of A/swine/Ohio/12TOSU447/2012(H3N2). Thirty seconds following inoculation, the fabric squares were manually picked up by a corner, allowing unabsorbed inoculum to run off. Squares were then placed in 15-mL sterile specimen cryogenic vials containing 5 mL of brain–heart infusion (BHI) broth supplemented with 10,000 U/mL penicillin and 10 mg/mL streptomycin. Sterile gloves were worn during procedures and replaced with new gloves between individual samples. The same volume of viral inoculum was added directly into 3 vials of BHI broth with no fabric substrate for positive controls. No treatment was added to the 3 negative control vials containing BHI broth. All vials were stored at −80°C for at least 24 h before virus isolation and molecular testing.
Vials containing inoculated fabric samples were rapidly thawed in a dry-bead bath at 37°C for 5 min. RNA was extracted from the samples (Mag-Bind viral DNA/RNA 96 kit, Omega Bio-Tek, Norcross, GA; MagMAX Express-96 magnetic particle processor, Life Technologies, Carlsbad, CA) according to the manufacturers’ instructions, but reagents were scaled to complement a sample volume of 0.1 mL. The VetMAX Gold SIV detection kit (Life Technologies) was used in conjunction with the 7500 fast thermocycler (Applied Biosystems, Thermo Fisher Scientific, Waltham, MA) for RT-qPCR, according to the manufacturer’s instructions. A standard curve was produced using a positive control supplied by the manufacturer. Log-transformed copy numbers were compared for each fabric using the Kruskal–Wallis test (Stata v.14.2, StataCorp, College Station, TX) at a confidence level of 95%. Fabric 6194 delivered the best RT-qPCR results of all of the fabrics tested, with a mean copy number of 8 × 106, but was not statistically different from cotton (Fig. 1). In vitro, none of the fabrics yielded RT-qPCR viral copy numbers that were significantly higher than cotton gauze, but 5889 only yielded 4 × 105 copies, which was significantly lower than cotton (p = 0.049).
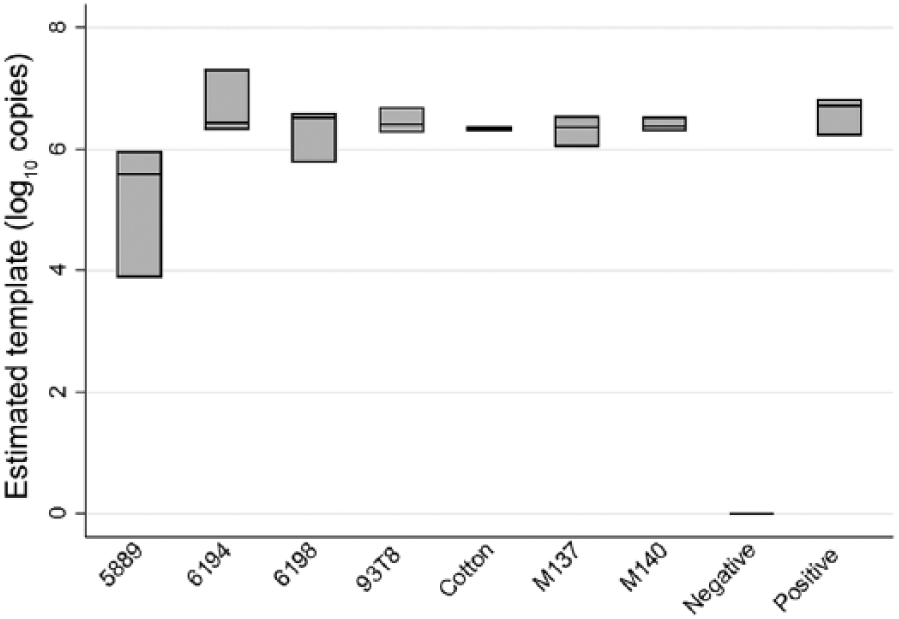
Comparison of influenza A virus detection based on reverse-transcription quantitative PCR. All fabrics were composed of 100% polyester except for M140 and M137, which were a 93% polyester and 7% co-polyester blend, and cotton, which was 100% cotton. Each box represents 3 samples for each fabric with log10-transformed estimated copy numbers shown; bar in each box is the mean.
A previous protocol was modified to allow viral quantification in 96-well assay plates containing Madin–Darby canine kidney (MDCK) cells. 1 Cells were washed with viral growth medium (VGM) and inoculated with 0.1 mL of each 10-fold serial dilution. Plates were incubated at 37°C and 5% CO2 for 1 h, then inoculum was overlaid with 0.1 mL of VGM. The plates were checked daily for cytopathogenic effects (CPEs), and all wells positive for CPE were recorded at 72 h. TCID50 values were calculated 7 and then log transformed. Statistical comparisons were performed using the Kruskal–Wallis test (Stata 14.2). Fabrics M140, 6194, and 93T8 all performed significantly better than cotton for the detection of viable IAV (Fig. 2). The 3 samples with cotton gauze substrate had a mean TCID50/mL of 1.2 × 106, whereas fabric M140 had a mean TCID50/mL value of 4.5 × 107. These 2 TCID50/mL values were statistically different from each other (p = 0.046). Fabric 6194 and 93T8 had mean TCID50/mL values of 2.7 × 107, which were statistically different from cotton (p = 0.049). Fabric M140 was chosen for subsequent field trials after yielding more viable IAV than 6194 and significantly more than 93T8 (p = 0.046).
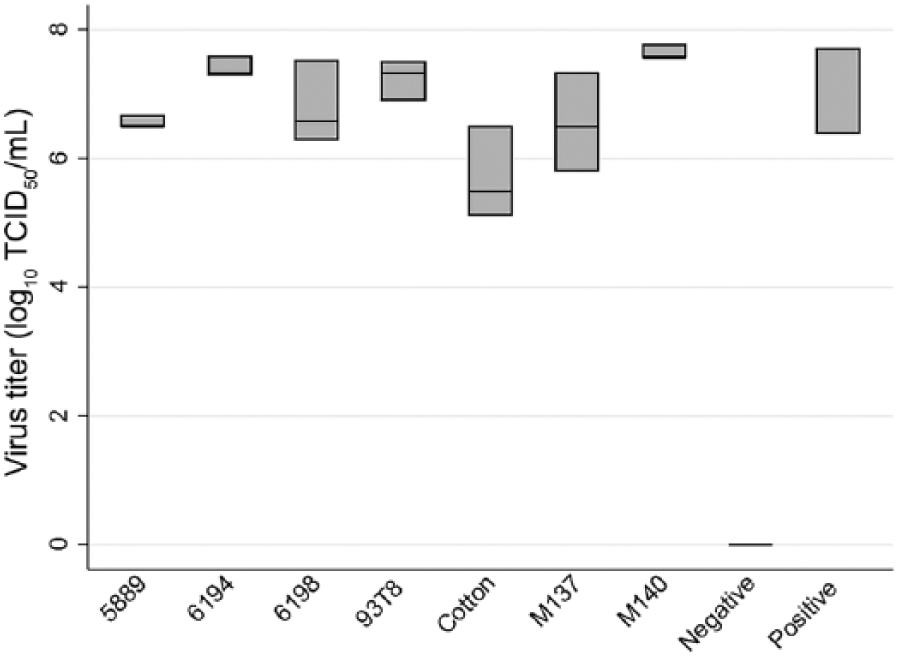
Comparison of influenza A virus growth for each fabric. All fabrics were composed of 100% polyester except for M140 and M137, which were a 93% polyester and 7% co-polyester blend, and cotton, which was 100% cotton. Log10-transformed TCID50/mL inoculation results are shown, with each box representing 3 samples for each fabric; bar in each box is the mean.
Polyester nasal swabs, cotton gauze nasal wipes, and fabric M140-calendar (M140) nasal wipes (given its performance with viral isolation) were used to sample pigs during active IAV outbreaks at 3 Ohio agricultural fairs. Samples were collected under Animal Use Protocol 2009A0134 approved by The Ohio State University’s Institutional Animal Care and Use Committee. At 2 fairs, 20 pigs were sampled by nasal swab, 20 different pigs were sampled with cotton gauze wipes, and an additional 20 pigs were sampled with M140 wipes. At the final fair, 20 pigs were sampled by nasal swab, 16 different pigs were sampled with cotton gauze wipes, and 16 additional pigs were sampled with M140 wipes. Fewer pigs were tested by gauze and M140 at the third fair because of limited supplies of materials. A total of 60 nasal swab, 56 cotton gauze nasal wipe, and 56 M140 nasal wipe samples were taken from 172 randomly chosen pigs. No comparison testing was done on the same pig in order to avoid reduction in nasal secretions as a result of taking consecutive wipes from the same pig. Samples were tested using RT-qPCR as described above, and samples with a cycle threshold (Ct) value ⩽40 were considered positive. All RT-qPCR–positive samples were inoculated onto MDCK cells as described previously 1 in an attempt to isolate IAV.
The proportion of positive samples detected by RT-qPCR with each sample type was compared using the proportions test in Stata software. Comparisons were also performed for the proportion of samples that were positive by virus isolation in each sample type. The proportion of positive samples by RT-qPCR and virus isolation from each sample type was comparable. Nasal swabs provided the most IAV-positive samples by RT-qPCR, with 58 of 60 (97%) swabs positive. IAV was detected in 51 of 56 (91%) M140 wipe samples, and in 50 of 56 (89%) cotton gauze wipe samples. The proportion of samples testing positive with RT-qPCR was not statistically different between the 3 sample types.
The most IAV isolates were recovered from pigs with nasal swabs (53 of 60 samples, 88%). The same numbers of isolates were recovered using both cotton wipes and M140 wipes (42 of 56 samples, 75%). The proportion of nasal swabs that tested positive by virus isolation was not statistically significantly different from M140 wipes and cotton gauze wipes (p = 0.062 for both). M140 nasal wipes did not outperform cotton gauze nasal wipes in the detection and isolation of IAV from swine.
Through our in vitro study and field trial, no polyester or mixed polyester and co-polyester fabrics performed better than cotton gauze when used as nasal wipes to collect samples for IAV testing from swine. In a previous study, cotton gauze was chosen as the most useful substrate for nasal wipes because it performed better than rayon-polyester gauze and similarly to dry electrostatic cleaning cloths for the recovery of IAV. 4 However, dry electrostatic cleaning cloths were not chosen for a subsequent field trial because of cost, difficulty in cutting and sterilizing the material, and the proprietary composition of the material, which could cause issues with downstream testing methods. 4 More viable IAV was recovered from polyester-tipped swabs in the previous study, so we investigated nasal wipe materials composed of polyester or mixed polyester and co-polyester. 4 The mixed polyester and co-polyester M140 fabric was more difficult to use when collecting samples in the field because it was more rigid and had a slick texture compared to cotton gauze.
The estimated copy number detected by RT-qPCR was similar for all tested materials, indicating that cotton did not significantly inhibit molecular detection of IAV. Cotton gauze performed as well as the polyester fabrics when used as nasal wipes in our study. It is possible that there are other materials that might perform better than cotton gauze, but difficulty in preparing and sterilizing other materials for sampling swine makes cotton gauze more appealing.
Footnotes
Acknowledgements
We thank Jean Schelhorn and Pierre Grondin for their assistance obtaining the fabrics.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was funded in part by federal funds from Centers of Excellence for Influenza Research and Surveillance (CEIRS), National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services, under contract HHSN272201400006C. Student support was provided by National Institutes of Health under award T35OD010977.
