Abstract
Peaton virus (PEAV; family Peribunyaviridae, genus Orthobunyavirus) appears to be capable of producing congenital malformations in ruminants; however, its pathogenicity remains unknown given its relatively low incidence. We evaluated the relationship between congenital abnormalities of calves and PEAV infection by serologic, epidemiologic, pathologic, and virologic investigations using specimens from 31 malformed calves in the years 1996–2016 in Japan. Antibody testing was carried out for known teratogenic viruses, including Akabane, Aino, Chuzan, and bovine viral diarrhea viruses, in the precolostral sera of these abnormal calves, but all results were negative. However, all 31 malformed calves were positive for antibodies against PEAV. A PEAV-specific gene was amplified from central nervous system tissues from a stillborn calf delivered in April 2007, and its nucleotide sequence was identical with that of PEAV isolated from healthy sentinel cattle in September 2006. These findings indicate that PEAV can cause bovine congenital anomalies.
Introduction
Fetal death and congenital malformations caused by arthropod-borne viruses (arboviruses), such as Akabane virus (AKAV, species Akabane orthobunyavirus) and Aino virus (AINOV, species Shuni orthobunyavirus; order Bunyavirales, family Peribunyaviridae, genus Orthobunyavirus) occasionally have impeded the production of ruminant livestock in the temperate zones of Asia, the Middle East, and Australia.4,5,9,16,29 AKAV and AINOV have been experimentally verified as teratogenic pathogens in ruminants.18,19,24,31 Studies on field clinical cases have indicated the teratogenicity of other orthobunyaviruses, such as Schmallenberg virus (SBV) and Shamonda virus (SHAV, species Shamonda orthobunyavirus), in domestic ruminants. An epidemic in 2011–2013 of fetal deformities caused by SBV resulted in large economic losses in the European livestock industry. 1 Naturally occurring intrauterine SHAV infections were likely involved in bovine congenital malformations in an epizootic in Japan in 2015–2016. 11 Based on complement fixation testing, the above-mentioned orthobunyaviruses have been included in the Simbu serogroup. 15 Although this serologic classification has not been certified by the International Committee on Taxonomy of Viruses (ICTV), the designation “Simbu serogroup” is often used for convenience. The relation of SBV to this serogroup was also indicated by phylogenetic studies.10,12
Peaton virus (PEAV) was initially isolated from Culicoides brevitarsis and bovine blood in Australia in the 1970s and has been recognized as a member of the Simbu serogroup. 28 In the 2017 ICTV classification, PEAV is regarded as a variant of species Shamonda orthobunyavirus. 8 However, genetic analyses showed that SHAV and PEAV are remotely related and thus reclassification is needed.10,35 As in other orthobunyaviruses, PEAV possesses 3 distinct linear, negative-stranded RNA segments: the S RNA segment encodes the nucleocapsid protein and a nonstructural protein (NSs); the M RNA segment encodes the precursor polyprotein, which is cleaved to form 2 viral surface glycoproteins Gc and Gn, and a nonstructural protein NSm; and the L RNA segment encodes the RNA-dependent RNA polymerase.8,10,36 Experimental infection of a pregnant ewe with PEAV induced congenital defects in its lamb. 25 However, the etiologic role of PEAV in nature has not yet been evaluated, to our knowledge. PEAV has been isolated repeatedly from Culicoides biting midges and sentinel calves since 1987 in southern Japan. 34 After the spread of PEAV, antibodies against PEAV were detected from calves with arthrogryposis and gross anatomic lesions in the central nervous system (CNS), suggesting its teratogenicity to fetal calves. 21 We examined the possible role of PEAV in congenital abnormalities of calves using clinical samples.
Materials and methods
Abnormal calves
Thirty-one newborn calves with abnormalities consistent with an arbovirus infection, but neither AKAV nor AINOV, were subjected to serologic, epidemiologic, pathologic, and virologic examinations (Table 1). Virus neutralization (VN) titers ⩾2 from malformed calves were considered positive for neutralizing antibody to the virus.
Characteristics of bovine congenital anomalies associated with Peaton virus in Japan, 1996–2016.
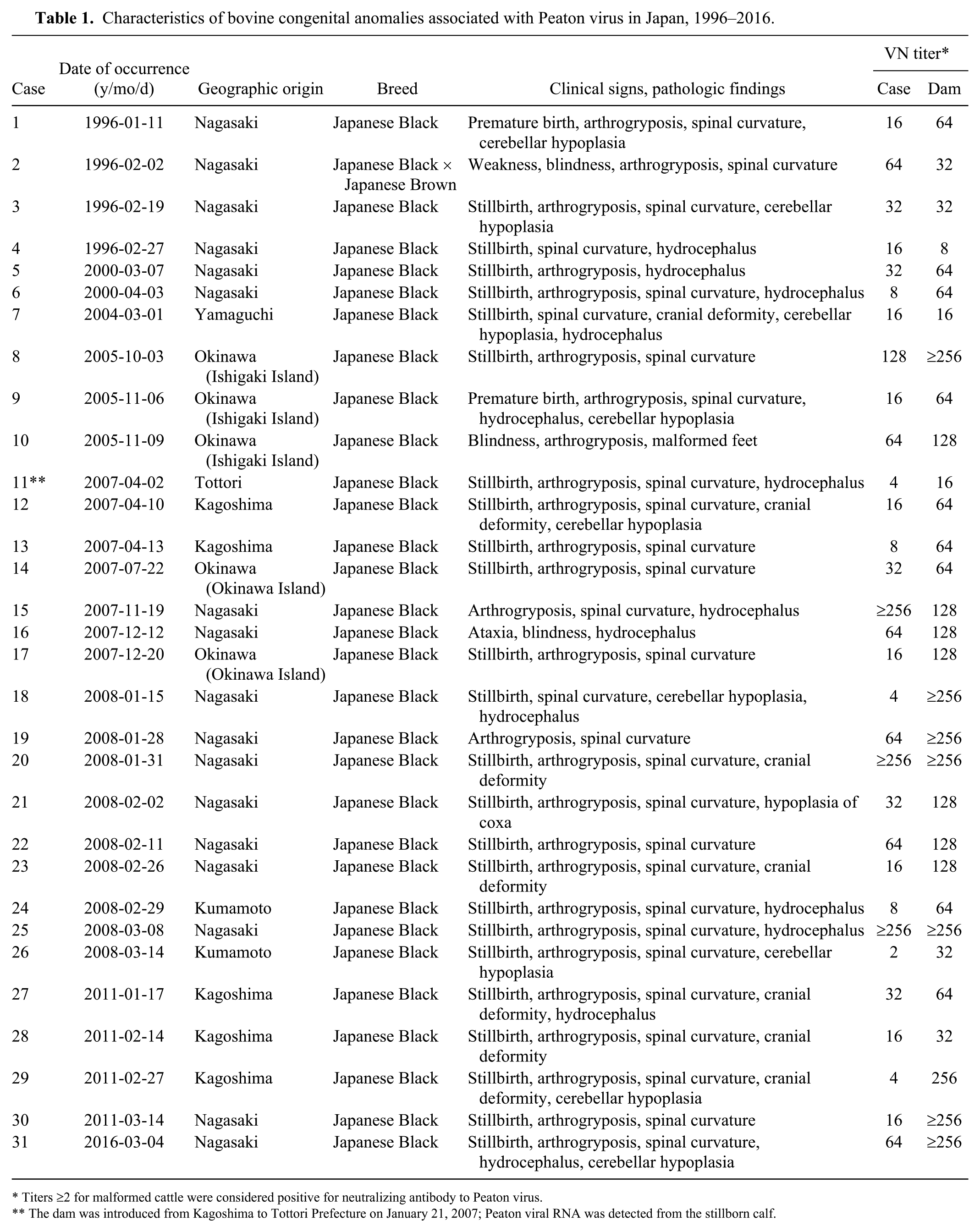
Titers ⩾2 for malformed cattle were considered positive for neutralizing antibody to Peaton virus.
The dam was introduced from Kagoshima to Tottori Prefecture on January 21, 2007; Peaton viral RNA was detected from the stillborn calf.
Virus neutralization test
Precolostral sera and/or body fluids from abnormal calves and their dams were examined by a VN test using bovine arboviruses (AKAV, AINOV, PEAV, Chuzan, and Ibaraki) and bovine viral diarrhea virus (BVDV). Ascites, pleural effusion, and/or cerebrospinal fluid were used in some cases when it was difficult to collect blood samples from stillborn calves. Sera of naive calves with no previous exposure to PEAV were used as the sentinel control group and were collected 4 or 5 times between June and November. Specimens inactivated at 56°C for 30 min were serially diluted 2-fold with serum-free Eagle minimum essential medium (MEM; Nissui Pharmaceutical, Tokyo, Japan) in 96-well cell culture microplates. Each serum dilution was mixed with an equal volume of 100 TCID50 of virus and incubated at 37°C for 1 h. Then, hamster lung (HmLu-1) cells suspended in GIT medium (Wako Pure Chemical Industries, Osaka, Japan) were added to each well. After incubation at 37°C for 7 d, the antibody titer was determined as the reciprocal of the highest serum dilution showing 50% inhibition of the cytopathic effect (CPE).
Processing of tissues for histopathology and immunohistochemistry
Tissue samples of the CNS, skeletal muscle, and other organs were collected from the abnormal calves. The samples were fixed with 10% buffered formalin, processed routinely, and stained with hematoxylin and eosin for histologic examination. One pair of serial sections was submitted for immunohistochemical detection of PEAV antigens. After deparaffinization of the paired sections, endogenous peroxidase activity was inhibited by treatment with 3% H2O2 in absolute methanol for 20 min at room temperature. In each section pair, one section was incubated with a diluted rabbit antiserum (1:2,000) against the PEAV NS-3/P/99 strain, 21 whereas the other was not, serving as the negative control. The sections were then labeled (HISTOFINE SAB-PO kit, Nichirei, Tokyo, Japan) with a biotinylated secondary antibody for anti-rabbit IgG and streptavidin-conjugated peroxidase. Localized peroxidase conjugates were visualized with 3,3’-diaminobenzidine. Sections were counterstained with hematoxylin.
RNA extraction and detection of viral genome
Tissue specimens obtained from the brain and spinal cord of abnormal calves were rinsed, minced, and homogenized in serum-free MEM. The supernatant of a 10% homogenate of each tissue was used for viral RNA preparation. Total RNA was extracted (QIAamp viral RNA mini kit, Qiagen, Hilden, Germany). Primer pair AKAI172F/AKAI560R (5’-CAGAAGAAGGCCAAGATGGT-3’/5’-AAGTTGACATCCATTCCATC-3’) and a RT-PCR kit (OneStep, Qiagen) were used to detect the viral S RNA segment. Reverse transcription was conducted at 50°C for 30 min. This mixture was then heated at 95°C for 15 min to stop the reaction and to activate HotStart Taq DNA polymerase (Qiagen). The resulting complementary DNA was amplified by 35 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s and extension at 72°C for 45 s, followed by one step of final extension at 72°C for 10 min. PCR products were electrophoresed on 1.5% agarose gel and visualized by staining with ethidium bromide.
Sequence analysis of the PCR products
Cerebrum and spinal cord tissues of a stillborn calf (case 11) were positive by the RT-PCR assay. The obtained PCR fragments were purified (QIAquick PCR purification kit, Qiagen) and directly sequenced in both orientations with primers AKAI206F (5’-CACAACCAAGTGTCGATCTTAC-3’) and AKAI560R (BigDye terminator v.3.1 sequencing kit, ABI3100-Avanti genetic analyzer, Thermo Fisher Scientific, Waltham, MA). Nucleotide sequences were analyzed (DNASIS Pro v.2.0, Hitachi Software Engineering, Tokyo, Japan; University of Wisconsin Genetic Computer Group (UWGCG) software package, Pharmacopeia, Princeton, NJ). Nucleotide sequence data were deposited in the DNA Data Bank of Japan under accession AB568273.
Virus isolation
The supernatant of homogenized tissues described above was also used for virus isolation. In addition, blood samples were collected from sentinel calves distributed throughout Nagasaki, Kagoshima, and Okinawa Prefectures. Heparinized blood samples were separated into erythrocytes and plasma by centrifugation, and erythrocytes were washed 3 times with ice-cold phosphate-buffered saline. The supernatants of 10% homogenates of tissues, plasma, and washed erythrocytes were stored at −80°C until use. These specimens were inoculated into monolayer cultures of HmLu-1 and/or baby hamster kidney (BHK-21) cells. After incubation at 37°C for 7 d, cell culture fluids were pooled and sub-inoculated into freshly prepared cell cultures at least twice until the infected cells exhibited CPE. The isolated viruses were identified by immunobinding assay as described previously using monoclonal antibodies recognizing the Gc protein of PEAV.17,37
Results
Clinical, pathologic, and epidemiologic features
Characteristic arthrogryposis and/or spinal curvature was consistently observed in 30 of 31 affected calves (Table 1, Fig. 1). Hydrocephalus (13 of 31) and cerebellar hypoplasia (9 of 31) were seen occasionally. However, hydranencephaly, as found in clinical cases of AKAV and AINOV infections, was not detected in the affected calves. Histologic examination indicated atrophy, dysplasia, and loss of skeletal muscle fibers accompanied by adipose replacement without polymyositis (18 of 30; Fig. 2), and loss of neurons in the ventral horn of the spinal cord (15 of 30) in many cases (Fig. 3). Nonsuppurative encephalomyelitis was recognized in only one case (11). No obvious changes to other main organs were observed in any of the cases.
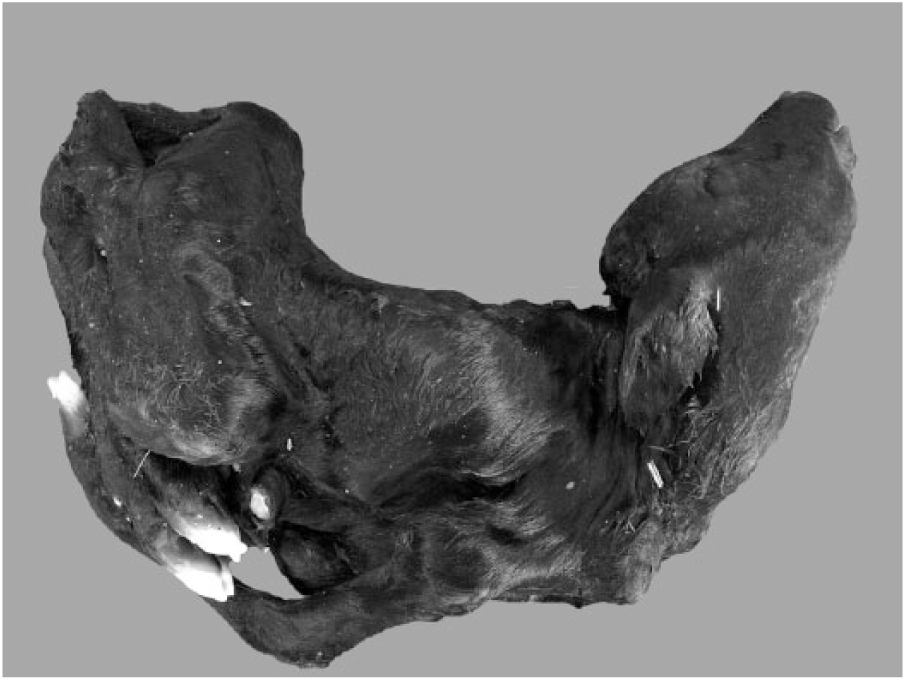
A Peaton virus–infected calf with arthrogryposis and spinal curvature (case 25).
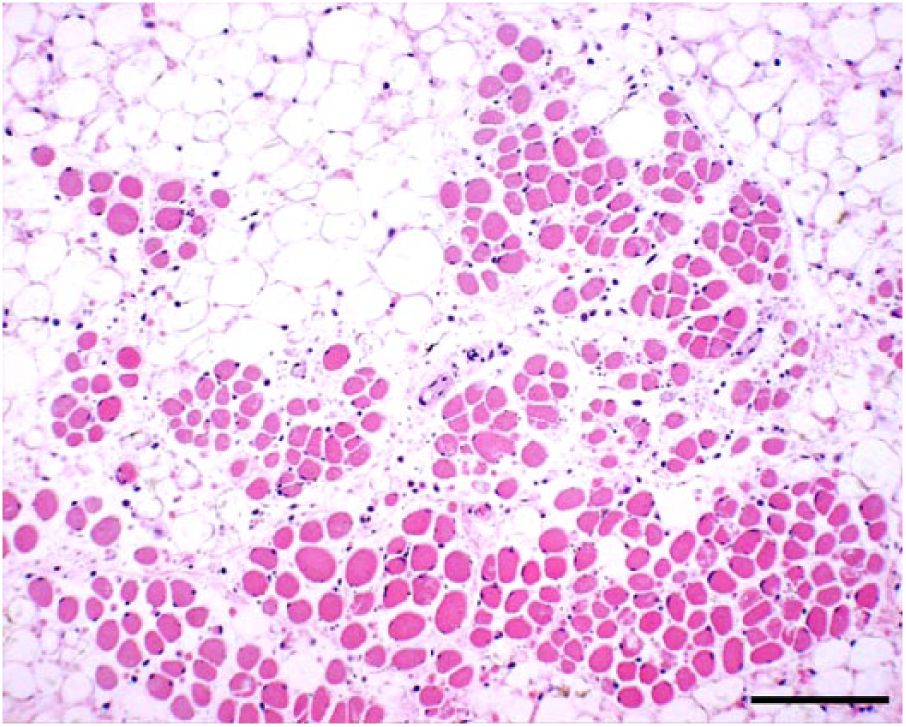
Atrophy, dysplasia, and loss of skeletal muscle fibers accompanied by adipose replacement attributed to Peaton virus infection in a stillborn calf. H&E. Bar = 0.25 mm.
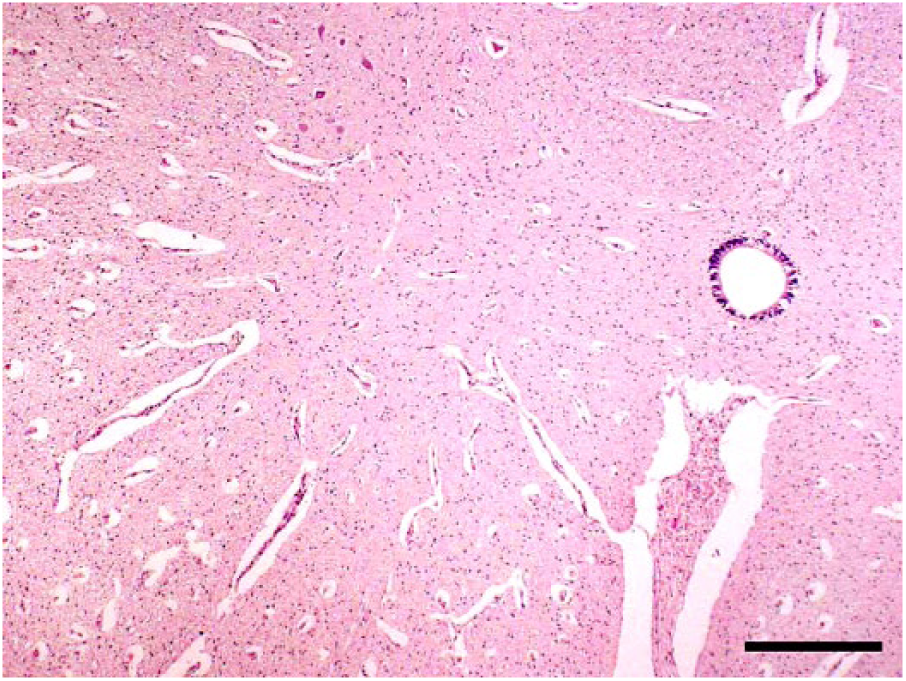
Loss of neurons in the ventral horn of the spinal cord of a Peaton virus–infected stillborn calf. H&E. Bar = 1 mm.
Thirty-one abnormal calves, of which 24 were stillbirths at the end of gestation, possessed antibodies specific to PEAV (Table 1). No antibodies against other arboviruses were detected in any affected calves. Antibodies to BVDV were found in an affected calf, but the VN titer was very low (⩽4). All dams also possessed neutralizing antibodies against PEAV, but had not shown any clinical signs. These cases of congenital abnormalities occurred between July and the following April in 1996–2016 in 6 prefectures (Nagasaki, Okinawa, Kumamoto, Kagoshima, Yamaguchi, and Tottori) in southwestern Japan. Of 31 calves, 24 were found in Kyushu district (the westernmost area of the mainland) and 5 in Okinawa Prefecture (the southernmost area consisting of many small islands). Twelve abnormal calves were delivered between November 2007 and March 2008.
Detection of viral antigens and genes, and virus isolation from clinical samples
Despite several attempts, no PEAV antigen was detected in the tissue samples from lesions in the CNS or skeletal muscles of the affected calves. No viruses were isolated. However, the viral gene (expected size of 389 bp) was amplified from the brain and spinal cord tissue of a stillborn calf (case 11) delivered in April 2007 in Tottori Prefecture (Table 1).
Sequence analysis of the PCR products
Except for the primer-binding sites, the nucleotide sequence of the amplified gene from case 11 was determined and compared with the corresponding region of the genomic sequences of PEAV strains reported previously.21,34 The PCR fragment showed identities of 90.4%, 99.0%, and 99.7% with positions at nucleotides 211–523 in the S RNA segment of the CSIRO110, NS-3/P/99, and MZ-4/C/99 strains, respectively, and was identical with that of the KSB-1/P/06 strain isolated in September 2006 in Kagoshima Prefecture (Supplementary Table 1).
Virologic and serologic analyses of sentinel calves
Ten strains of PEAV were isolated from bovine blood samples collected in Okinawa (1987, 2001, 2005, and 2009), Nagasaki (1999), and Kagoshima (2006 and 2010) Prefectures (Supplementary Table 1). Two strains of the virus were also obtained from Culicoides biting midges captured at cowsheds in Miyazaki in 1999 and Kagoshima in 2010.13,21
Retrospective serologic investigation using the sera of sentinel calves collected in 1985–2009 in Nagasaki Prefecture (71–75 per year) revealed that seroconversion to PEAV occurred 6 times, or once each in 1985 (positive rate: 2%), 1987 (8%), 1995 (11%), 1999 (38%), 2007 (27%), and 2009 (7%). In the spring of the years after high seroconversion rates were observed, the numbers of abnormal calves suspected of PEAV infection were 4 (1996), 2 (2000), and 7 (2008; Table 1). In Okinawa Prefecture, seroconversion was frequently observed in sentinel calves in 1996–2009. PEAV seemed to have circulated in this area during each of 6 different years: 1996 (positive rate: 4%), 1998 (3%), 1999 (3%), 2005 (5%), 2007 (11%), and 2009 (4%). During this period, however, only 5 suspected cases were reported in Okinawa Prefecture (Table 1).
Discussion
Detection of virus-specific antibodies in the precolostral serum of abnormal offspring can be useful to identify etiologic agents. In general, only transient viremia of the related orthobunyaviruses has been observed in infected animals.19,33 Therefore, virus isolation and viral gene detection are generally unsuccessful in malformed calves, because the virus disappears from the infected fetuses during gestation. 6 Surveillance in sentinel animals is useful to detect new or unexpected arboviral disease.11,23,27,30 The detection of specific antibodies in 31 malformed calves and viral circulation in the affected regions strongly suggested an association between PEAV and the congenital abnormalities. Although signs and pathologic findings were very similar to other cases, antibody titers in body fluids from several affected calves (cases 18, 26, and 29) were low. Because VN tests for Simbu serogroup viruses often produce nonspecific reactions, 20 more evidence of in utero infection is needed to support the involvement of PEAV in these cases. The amplification of a PEAV-specific gene from affected calf 11 supported our suspicion. Of interest is that the sequence of the amplified product was identical with the corresponding region of the PEAV KSB-1/P/06 strain isolated in Kagoshima in the autumn of 2006. The dam of case 11 was transported from Kagoshima to Tottori in the winter of 2007, suggesting that the infection occurred in Kagoshima. Serosurveillance and virus isolation from sentinel cattle indicated that almost all of the other clinical cases were epidemiologically consistent with the circulation of PEAV.
The clinical and pathologic features of malformed calves associated with PEAV resembled those caused by AKAV and AINOV. Arthrogryposis and spinal cord lesions were frequently found in the malformed calves confirmed to have PEAV infection. However, the brain lesions of the PEAV-infected calves appeared to be milder than those in calves infected with AKAV or AINOV. Also, fatty replacement in skeletal muscle was an observation specific to PEAV. These findings suggest that the tissue tropism and pathogenic capacities of PEAV are different than those of AKAV and AINOV. On the other hand, fatty replacement in skeletal muscle has been observed in malformed calves affected by SBV and SHAV.2,11,26 Comparison of the lesions caused by these orthobunyaviruses will clarify tissue tropism and lesion pathogenesis in malformed calves. There were no reports of bovine abortions associated with PEAV during the study period.
Until vaccines were used more widely, large outbreaks of congenital abnormalities caused by AKAV and AINOV occurred repeatedly. 9 During the largest epidemic caused by AKAV (1972–1975), 42,000 affected calves were reported throughout Japan. AINOV was also involved in at least 700 cases of fetal death and congenital abnormalities in cattle in 1995–1996. However, reported cases of bovine malformation associated with PEAV were highly sporadic (1–12 cases in each epidemic season). Insufficient awareness and motivation to report might have affected these data. However, no obvious increase of bovine congenital abnormalities was reported after PEAV spread in the endemic regions (data not shown). These findings imply that the pathogenicity of PEAV is relatively low in comparison with that of AKAV and AINOV. As of 2018, it seems that PEAV has a low or limited impact on cattle reproduction in Japan. Although vaccination is an effective measure to prevent fetal death and congenital abnormalities caused by orthobunyaviruses, its application to PEAV must be carefully considered from a cost-effectiveness perspective.
Serosurveys have indicated that PEAV is widely distributed in Africa, Asia, and Australia.20,22,28,32 Although PEAV potentially affects reproductive performance in domestic ruminants, no association between reproductive disorders and PEAV has been reported from its regions of prevalence outside Japan. This could be explained by a lack of prevalence data for this agent and/or reporting systems. In some regions, animals may be infected and develop immunity to PEAV before reaching reproductive age. If the virus is introduced into a nonendemic region, outbreaks of malformation will be more obvious. High seroprevalence (58–71%) against PEAV was observed in lower-latitude areas, such as in Indonesia and Tanzania,20,22 suggesting that the virus exists in tropical and/or subtropical zones. It has been postulated that long-distance movement of Culicoides biting midges is associated with the spread of arboviruses. 7 Antibodies against PEAV in sentinel cattle have been detected frequently in the most-southwestern islands of Japan during long-term monitoring, suggesting that PEAV has been endemic in neighboring subtropical regions. 14 The abnormal calves confirmed to be infected with PEAV were localized to western Japan, which suggests that the infected midges may have been brought by air streams from overseas.
If PEAV is introduced into a large naive ruminant population, the impact will become more significant. The large outbreak of abnormal deliveries in previously SBV-free regions suggests this possibility. 1 Several possible reassortants between closely related orthobunyaviruses were identified by genetic analyses of field-isolated viruses, and such reassortants had features distinct from those of their parent viruses. 3 Genetic reassortment between PEAV and other orthobunyaviruses could possibly change viral antigenicity and pathogenicity. A previous phylogenetic study indicated that a reassortment occurred between AINOV and PEAV in nature. 34
Our findings did not show high animal losses caused by PEAV infection. However, a differential diagnosis of congenital abnormalities caused by orthobunyaviruses including AKAV, AINOV, SHAV, and PEAV would be necessary in regions where these viruses are co-distributed. We recommend that PEAV be added to the list of possible etiologic agents that can be found in malformed calves. Further work to clarify differences in pathogenicity among Simbu serogroup viruses will contribute to the development of strategies to prevent and reduce animal losses caused by these potentially teratogenic viruses.
Supplemental Material
DS1_JVDI_10.1177_1040638718796269 – Supplemental material for Congenital abnormalities in calves associated with Peaton virus infection in Japan
Supplemental material, DS1_JVDI_10.1177_1040638718796269 for Congenital abnormalities in calves associated with Peaton virus infection in Japan by Yoichi Matsumori, Maki Aizawa, Yoshiko Sakai, Daisuke Inoue, Michiko Kodani, Osamu Tsuha, Akira Beppu, Yoshimasa Hirashima, Ryota Kono, Akifumi Ohtani, Tohru Yanase, Hiroaki Shirafuji, Tomoko Kato, Shogo Tanaka and Makoto Yamakawa in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank veterinary officers at Nagasaki, Tottori, Okinawa, Kagoshima, Kumamoto, and Yamaguchi Prefectural livestock hygiene service centers and technical staff at Kyushu Research Station, National Institute of Animal Health, for excellent technical support and sampling.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the “Research and Development Projects for Application in Promoting New Policy of Agriculture, Forestry and Fisheries” from the Ministry of Agriculture, Forestry and Fisheries of Japan.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
