Abstract
Although inactivating mutations of tumor suppressor genes are well described in cell lines of canine osteosarcoma (OS), expression of tumor suppressor proteins in spontaneous disease is poorly characterized. We determined the immunohistochemical expression of p53, PTEN, Rb, and p16 in a large cohort of dogs with OS. Formalin-fixed, paraffin-embedded samples of canine OS were analyzed retrospectively. Primary tumor samples from 145 dogs, collected between 2003 and 2008, were evaluated by tissue microarray. Streptavidin–biotin complex immunohistochemistry was performed using monoclonal antibodies for Rb and PTEN and polyclonal antibodies for p16 and p53. The average age of dogs was 7.6 y, and 118 of 145 (81%) were purebred. Most commonly represented purebreds were Greyhound (23%), Rottweiler (11%), and Labrador Retriever (10%). Immunohistochemical detection of p53, PTEN, Rb, and p16 was 81%, 61%, 66%, and 66%, respectively. The staining pattern for p16 was primarily cytoplasmic; the predominant pattern for PTEN, Rb, and p53 was cytoplasmic and nuclear. Exclusively cytoplasmic staining was noted in 19% of samples positive for p53 and 8% of samples positive for Rb. Kaplan–Meier curves showed that protein expression was not associated with significant differences in overall survival (
Introduction
Osteosarcoma (OS) is the most common primary bone malignancy in dogs. 29 Tumors display aggressive biologic behavior with extensive local destruction and micrometastases that occur early during the disease. Although OS is comparatively rare in humans, clinical and biologic similarities and the relative frequency of canine OS make it an excellent model for spontaneously occurring human disease. 10 Canine and human OS share similar molecular features, demonstrated by gene expression profiling and comparative genomic hybridization. 1 Genomic instability is a hallmark of both the human and canine disease, involving a broad variety of oncogenes and tumor suppressor genes. Many genes altered in OS have effects on the cell cycle17,21,36 or DNA repair.18,37
Progression through the G1/S checkpoint is determined by cyclin D–cyclin-dependent kinase (CDK)4 complexes, which ultimately regulate phosphorylation of retinoblastoma (Rb). The
In cell lines of OS, inactivation of tumor suppressor genes
Abnormal activity of the nuclear phosphoprotein p53 has been noted in spontaneously occurring canine OS.15,30 P53 is considered the “guardian of the genome” and links DNA repair, cell cycle arrest, and apoptosis. Wild-type p53 has a short half-life and cannot be detected by immunohistochemistry (IHC). Mutant p53 has greater stability, allowing IHC as a tool for indirectly evaluating p53 function. Mutant p53 can acquire oncogenic functions that are independent of wild-type p53. 26 Loss of p53 in mesenchymal stem cells and osteoblast precursors promotes commitment toward osteoblasts but blocks terminal differentiation to mature osteocytes. 35 Double-mutant Rb;p53 mice develop early-onset OS with complete penetrance. 5
Reduction in phosphatase and tensin homologue deleted on chromosome 10 (PTEN) protein has been noted in canine OS cell lines and spontaneous tumors.
22
Reduced gene expression has also been documented by reverse-transcription quantitative (RT-q)PCR in spontaneous canine OS,
1
and
Although tumor suppressor proteins (TSPs) have been studied in vitro and in vivo, studies evaluating protein expression in a larger cohort of spontaneous canine OS are limited. Our objectives were: 1) to construct a tissue microarray (TMA) of canine OS suitable for immunohistochemical characterization, and 2) to determine the frequency of TSP expression in a large cohort of dogs with OS. We hypothesized that proteins p16, p53, PTEN, and Rb can be detected in spontaneous cases of canine OS.
Materials and methods
Tumor material
Cases were identified by a computer-based search of The Ohio State University (OSU) College of Veterinary Medicine Pathology archives for diagnoses of OS from the years 2003–2008. All samples had been routinely fixed in 10% neutral-buffered formalin for 24–72 h, followed by demineralization as necessary. Demineralization solutions had been chosen according to pathologist preference. The 3 demineralization solutions routinely used in the OSU Department of Veterinary Biosciences are hydrochloric acid and EDTA (Decalcifier-S, US Biotex, Webbville, KY), formic acid, formaldehyde, and methyl alcohol (Surgipath decalcifier I, Leica, Richmond, IL), and hydrochloric acid (Surgipath decalcifier II, Leica).
Material was available for use from 150 of the cases reviewed. Prior to constructing the tissue array, hematoxylin and eosin (H&E)-stained slides were reviewed, and the diagnosis of osteosarcoma was confirmed by a board-certified pathologist (DS Russell). Neoplasms were subtyped according to the World Health Organization (WHO) International Histologic Classification of Tumors of Domestic Animals. 32
Tissue microarray and IHC
TMAs were constructed (Manual tissue arrayer MTA-1, Beecher Instruments, Sun Prairie, WI). Briefly, on a H&E-stained slide, representative areas of tumor were defined as foci that represented the predominant morphologic phenotype (i.e., osteoblastic, fibroblastic, chondroblastic, etc.); foci of necrosis and reactive bone were excluded. The TMA was then constructed from corresponding areas in the formalin-fixed, paraffin-embedded block. The recipient block was constructed to hold 54 cores (9 × 6) spaced 2.5 mm between sample centers. Representative punches (1.5 mm wide) of tumor were removed and inserted into a recipient block containing paraffin treated identically to the donor block. Tumors were evaluated in triplicate and placed adjacent to each other in the array, compensating for tumor heterogeneity. Prior to sectioning, the array block was smoothed, leveled, and warmed to 37°C for 15 min to encourage binding between the cores and recipient block. The block was faced to ensure that all cores had representative sectioning.
For IHC, 4-µm sections were mounted on poly-L-lysine slides, deparaffinized in xylene, and rehydrated in ethanol. Antigen retrieval was completed by incubating slides in target retrieval solution (Dako, Carpinteria, CA) at a 1:10 dilution in a decloaking chamber. To block endogenous peroxidase activity, the slides were placed in 3% H2O2 for 10 min. To prevent nonspecific antibody binding, the slides were incubated with 3% nonfat dry milk (NFD) in phosphate-buffered saline (PBS) for 30 min. Primary antibodies, dilutions, and incubation times were: vimentin (V9, mouse monoclonal, Dako), dilution 1:100, 30 min incubation; PTEN (A2B1, mouse monoclonal, Santa Cruz Biotechnology, Santa Cruz, CA), dilution 1:100, 120 min incubation; p16 (H-156, rabbit polyclonal, Santa Cruz Biotechnology), dilution 1:100, 120 min incubation; p53 (CM1, rabbit polyclonal, Covance, Emeryville, CA), dilution 1:100, 120 min incubation; Rb (G3-245, mouse monoclonal, BD Pharmingen, San Jose, CA), dilution 1:100, 120 min incubation. Western blot validation of antibody clone specificity is described elsewhere. 19 After rinsing in PBS, slides were incubated with biotinylated universal (anti-mouse IgG/rabbit IgG) antibody at a dilution of 1:200 for 30 min (Vectastain universal elite ABC kit, Vector Laboratories, Burlingame, CA). Rinsed slides were incubated for 30 min with Vectastain elite ABC reagent. Finally, sections were incubated with 3,3′-diaminobenzidine (DAB, MilliporeSigma, St. Louis, MO) tablets prepared in tap water for 5 min. Slides were counterstained with Harris hematoxylin, dehydrated, and coverslipped.
Control slides were incubated with a mouse or rabbit universal negative control antibody, with staining steps and incubation times otherwise identical to the primary antibody (FLEX universal negative control antibodies for mouse and rabbit, Dako). To confirm preservation of intermediate filaments in the formalin-fixed and demineralized tissues, vimentin IHC was used as a positive control.
Immunohistochemical scoring
All IHC was evaluated by a single board-certified pathologist (DS Russell). The sample was considered positive if >10% of interpretable neoplastic cells among all 3 cores stained positive. The intensity of immunohistochemical staining was interpreted semi-quantitatively and scored as negative (score 0), weak (score 1), moderate (score 2), and intense (score 3). Staining was determined to be cytoplasmic, nuclear, or cytoplasmic and nuclear.
Survival analysis
Survival times were calculated from the time of initial diagnosis to the time of death. Patients were excluded from analysis if death was unrelated to osteosarcoma, if survival <30 d (i.e., excluding for complications directly associated with surgery), and if standard of care (i.e., surgery and chemotherapy) was not performed. Statistical analyses were performed (Prism 6, GraphPad Software, La Jolla, CA). The Kaplan–Meier approach was used to compare survival with expression of each individual TSP. Animals lost to follow-up were censored at the time of last known contact with a veterinarian. The log-rank test was used for comparisons; all values of
Results
Sample distribution
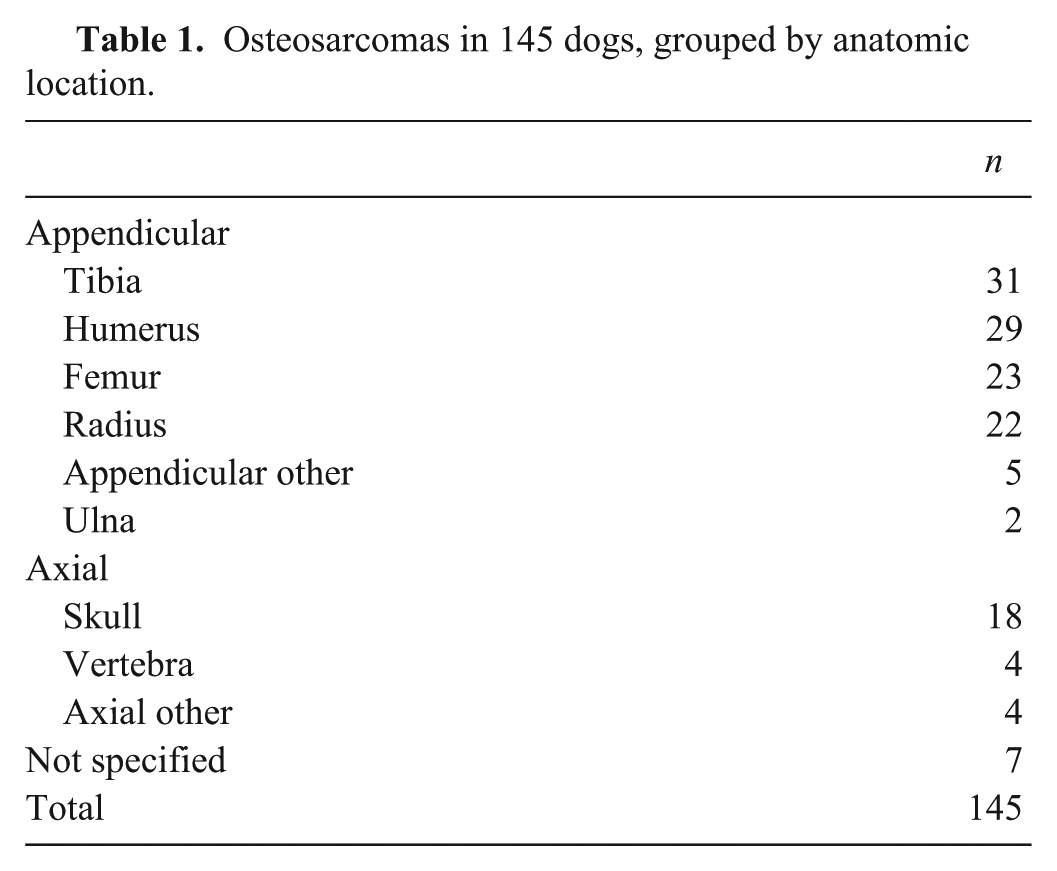
Of the 150 samples identified, 5 samples stained negatively for vimentin and were excluded from subsequent analysis. Of 145 tumors included in our study, 112 were appendicular, 26 were axial, and in 7 the location was not specified (Table 1). The average age at the time of biopsy was 7.6 y (median: 8 y). One hundred eighteen of 145 (81%) dogs were purebred, 24 of 145 (17%) dogs were mixed breed, and 3 (2%) dogs were not specified. The most frequently represented breed was Greyhound (
Osteosarcomas in 145 dogs, grouped by anatomic location.

IHC
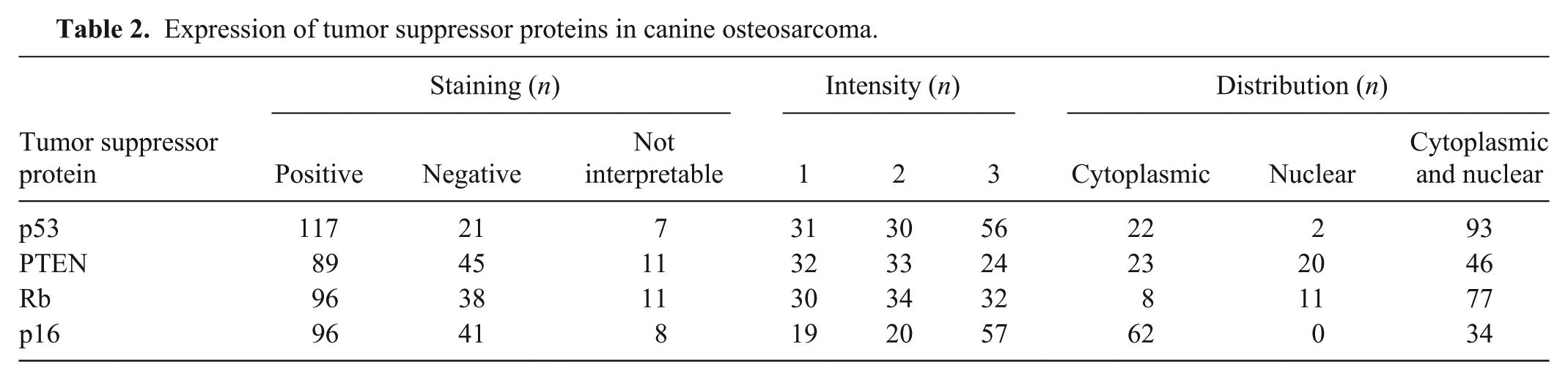
Arrays were generally of high quality, although among the 4 TSPs, 5–8% of samples could not be interpreted because of inadequate cellularity of the punches. Immunohistochemical findings for the 4 TSPs are shown in Table 2. Findings are subgrouped into the 3 most frequent breeds, location (axial and appendicular), and histologic subtype (Supplementary Table 1).
Expression of tumor suppressor proteins in canine osteosarcoma.
P16 IHC was positive in 66% of cases; the most common distribution pattern was cytoplasmic (65%; Fig. 1). Rb IHC was positive in 66% of cases; cytoplasmic and nuclear staining was the most frequent pattern (Fig. 2). For PTEN IHC, 61% of tumors stained positively. Nuclear and cytoplasmic staining was most common (52%); in many cases, nuclear localization was most intense within the nucleolus (Fig. 3). For p53 IHC, 81% of all samples stained positively. Staining was most commonly intense. The most common staining pattern was cytoplasmic and nuclear (80%); exclusively cytoplasmic staining was noted in 19% of tumors (Fig. 4).
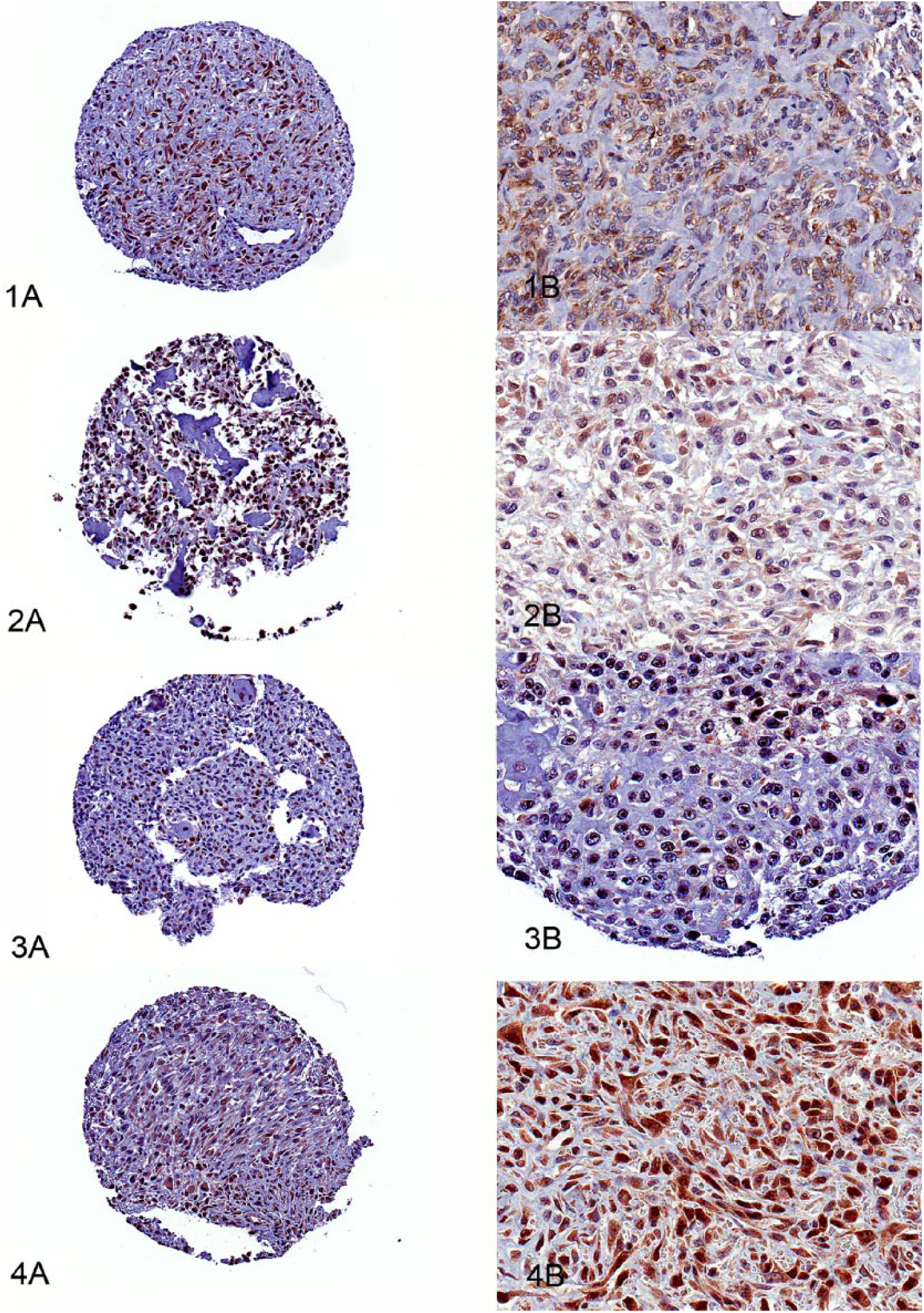
Immunohistochemical expression of tumor suppressor proteins in canine osteosarcoma at low (
Survival analysis
Seventy-two patients were included for survival analysis; all animals were either dead or censored at the end of the study period (Supplementary Fig. 1). The median survival time for all dogs was 279 d (30–986 d). Expression of each TSP was not significantly associated with survival time (
Discussion
We demonstrated that TMA can be used effectively for high-throughput immunohistochemical detection of TSPs in OS. Products of tumor suppressor genes
The most frequently detected protein was p53, expressed in 81% of samples. Normally, p53 is in low concentration in the cell, and the wild-type protein has a short half-life. 20 Its presence in the majority of canine OS samples suggests abnormal p53 activity. The frequency of p53 expression in this dataset is similar to previous IHC-based studies against archived canine OS tissues.6,30 The extranuclear localization of p53 immunostaining could be further evidence of abnormal p53 activity. Cytoplasmic and nuclear expression was found in 80% of samples, and exclusively cytoplasmic staining was noted in 19% of samples. Cytoplasmic accumulation of p53 could be the result of an abnormal nuclear localization signal or cytoplasmic translocation as the result of binding by a 70-kDa heat-shock protein (HSP70). 11 Our survival data for p53 expression found no significant association with overall survival, and this contrasts with findings of one previous report. 6 This contrast might reflect differences in study cohorts, antibody performance, histologic evaluation technique (i.e., TMA cores vs. conventional histologic sectioning), and slide interpretation.
PTEN expression was absent in 31% of samples. Western blot of canine OS cell lines has previously demonstrated that PTEN protein was absent or severely reduced in 3 of 5 cell lines, with large deletions in the
Rb and p16 were not detected in 26% and 28% of neoplasms, respectively. Absence of these proteins, which regulate G1 to S transition, may suggest disturbed cell cycle regulation in canine OS. Some variation in Rb expression might be related to cell cycle stage: even in the presence of intact Rb, cells in G0 or mid-G1 might lack nuclear expression. 38 Some degree of nuclear Rb staining was present in 92% of cases, and 80% had concurrent cytoplasmic staining. A small percentage (8%) of cases had exclusively cytoplasmic staining. A variety of poorly differentiated human tumors have been associated with cytoplasmic mislocalization of Rb. 14 The spatial distribution of Rb is determined by a nuclear localization signal in the C-terminus 39 and association of the N-terminus with nuclear matrix proteins. 9 Phosphorylation during G1 to S transition results in decreased affinity for the nuclear compartment.23,24,34 CDK-dependent Rb phosphorylation may play an important role in nucleocytoplasmic shuttling. 13 Mislocalization has been linked to reduced CDK activity, mutation of CDK-dependent Rb phosphorylation sites, or inhibition of the exportin-1 pathway. 13
Limitations of these data are inherent to the retrospective nature of our study. Given that IHC detects the presence or absence of protein, these data do not necessarily take into account epigenetic modifications or protein functionality. For example, even in the presence of p16 and Rb,
Supplemental Material
DS1_JVDI_10.1177_1040638718770239 – Supplemental material for Immunohistochemical detection of p53, PTEN, Rb, and p16 in canine osteosarcoma using tissue microarray
Supplemental material, DS1_JVDI_10.1177_1040638718770239 for Immunohistochemical detection of p53, PTEN, Rb, and p16 in canine osteosarcoma using tissue microarray by Duncan S. Russell, Lauren Jaworski, William C. Kisseberth in Journal of Veterinary Diagnostic Investigation
Supplemental Material
DS2_JVDI_10.1177_1040638718770239 – Supplemental material for Immunohistochemical detection of p53, PTEN, Rb, and p16 in canine osteosarcoma using tissue microarray
Supplemental material, DS2_JVDI_10.1177_1040638718770239 for Immunohistochemical detection of p53, PTEN, Rb, and p16 in canine osteosarcoma using tissue microarray by Duncan S. Russell, Lauren Jaworski, William C. Kisseberth in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Florinda Jaynes for assistance with IHC and Dr. Jaime Modiano for critical review of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
We thank the Department of Veterinary Clinical Sciences, The Ohio State University, for funding support.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
