Abstract
Fifty-three cattle of unknown serologic status that were not persistently infected (PI) with bovine viral diarrhea virus (BVDV) were commingled with 10 cattle that were PI with different strains of BVDV, and were monitored for an extended commingle period using a reverse-transcription real-time PCR (RT-rtPCR) BVDV assay on various sample types. Transient infections with BVDV were also assessed by virus isolation, virus neutralization (VN) assays, and direct buffy coat 5′-UTR sequencing. Infections were demonstrated in all cattle by RT-rtPCR; however, the detection rate was dependent on the type of sample. Buffy coat samples demonstrated a significantly greater number of positive results (p ≤ 0.05) than either serum or nasal swab samples. Presence of elevated BVDV VN titers at the onset inversely correlated with the number of test days positive that an individual would be identified by RT-rtPCR from buffy coat samples, and directly correlated with the average Ct values accumulated over all RT-rtPCR test days from buffy coat samples. Both single and mixed genotype/subgenotype/strain infections were detected in individual cattle by direct sample 5′-UTR sequencing. A BVDV-2a strain from a PI animal was found to be the predominant strain infecting 64% of all non-PI cattle; BVDV-1b strains originating from 3 PI cattle were never detected in non-PI cattle. Although direct sample 5′-UTR sequencing was capable of demonstrating mixed BVDV infections, identifying all strains suspected was not always efficient or possible.
Introduction
Significant economic losses associated with bovine viral diarrhea virus (BVDV) infections in cattle are reported worldwide.10,15,27,39 These viruses are a diverse group of single-stranded, positive-sense RNA viruses (species Bovine viral diarrhea virus, genus Pestivirus, family Flaviviridae), and are classified into 2 species or genotypes, BVDV-1 and BVDV-2. Each of these genotypes is further subdivided into several subgenotypes (BVDV-1a–1q and BVDV-2a–2c) that are distributed worldwide.1,31,41,44 Presently, only BVDV-1a, BVDV-1b, BVDV-2a, and BVDV-2b subgenotypes have been identified in North America, and BVDV-2b only rarely19,38,45; however, the prevalence of the 1a, 1b, and 2a subgenotypes has changed over time.21,42
Two distinct BVDV infections defined in cattle are “acute, primary, or transient infections” and “persistent infections.”2,47 The primary economic losses attributed to “acute” BVDV infections are associated with respiratory or reproductive diseases.9,13,20,40,43 The second type of BVDV infection is the result of fetal infections with a noncytopathic BVDV biotype, at 2–4 mo of gestation. The resulting neonate will be immunotolerant to the homologous BVDV strain, remain persistently infected (PI) with that strain of BVDV for its life, and serve as an important reservoir for the transmission of virus in a susceptible population.8,33,34,49
To date, BVDV control efforts have been focused primarily on detecting PI cattle. This process can ignore detection and management of transient BVDV infections, which are frequently associated with morbidity and mortality in cattle because of the immunosuppressive effects of BVDV and its potentiating effect on secondary infections. Of particular importance is the documented association of transient BVDV infections and bovine respiratory disease (BRD) in feedlot cattle.9,16,18,46 Exposing susceptible cattle to BVDV PI cattle by commingling them is a means of introducing BVDV infections into a herd, and has been demonstrated experimentally to be a very successful challenge model to demonstrate morbidity rates.17,25,36 In prior commingle studies, there have been variations observed in the clinical outcomes,17,20,35 the number of PI cattle involved,17,25,30,35 and the genetic characterization of the BVDV from PI cattle into genotypes, subgenotypes, or strain within a subgenotype, or these factors were not defined.17,20,25,30,36
We commingled a large group of non-PI cattle (n = 53) of unknown serologic status, typical of most feedlot practices in the United States, with 10 confirmed BVDV PI cattle that were each infected with a phylogenetically different strain of BVDV-1a, -1b, or -2a. Our objectives were 3-fold: 1) to determine the effect of PI contact with non-PI cattle, 2) determine the extent of transmission of BVDV strains to the non-PI cattle, monitored with a commercial RT-rtPCR assay on various sample types (buffy coat, serum, nasal swab, and ear notch samples), and 3) to further characterize BVDV infections by virus isolation in tissue culture (VI), virus neutralization (VN) assays, and by 5′-UTR sequencing of BVDV RNA from whole blood buffy coat samples.
Materials and methods
Animal selection and commingling
Ten PI cattle (P1–P10; obtained from multiple feedlots in southwest Kansas), 2 with BVDV-1a genotype infections (PI-2, PI-5), 6 with BVDV-1b genotype infections (PI-1, PI-3, PI-4, PI-6, PI-8, and PI-10), and 2 with BVDV-2a genotype infections (PI-7, PI-9) were chosen for the commingle study (Fig. 1).
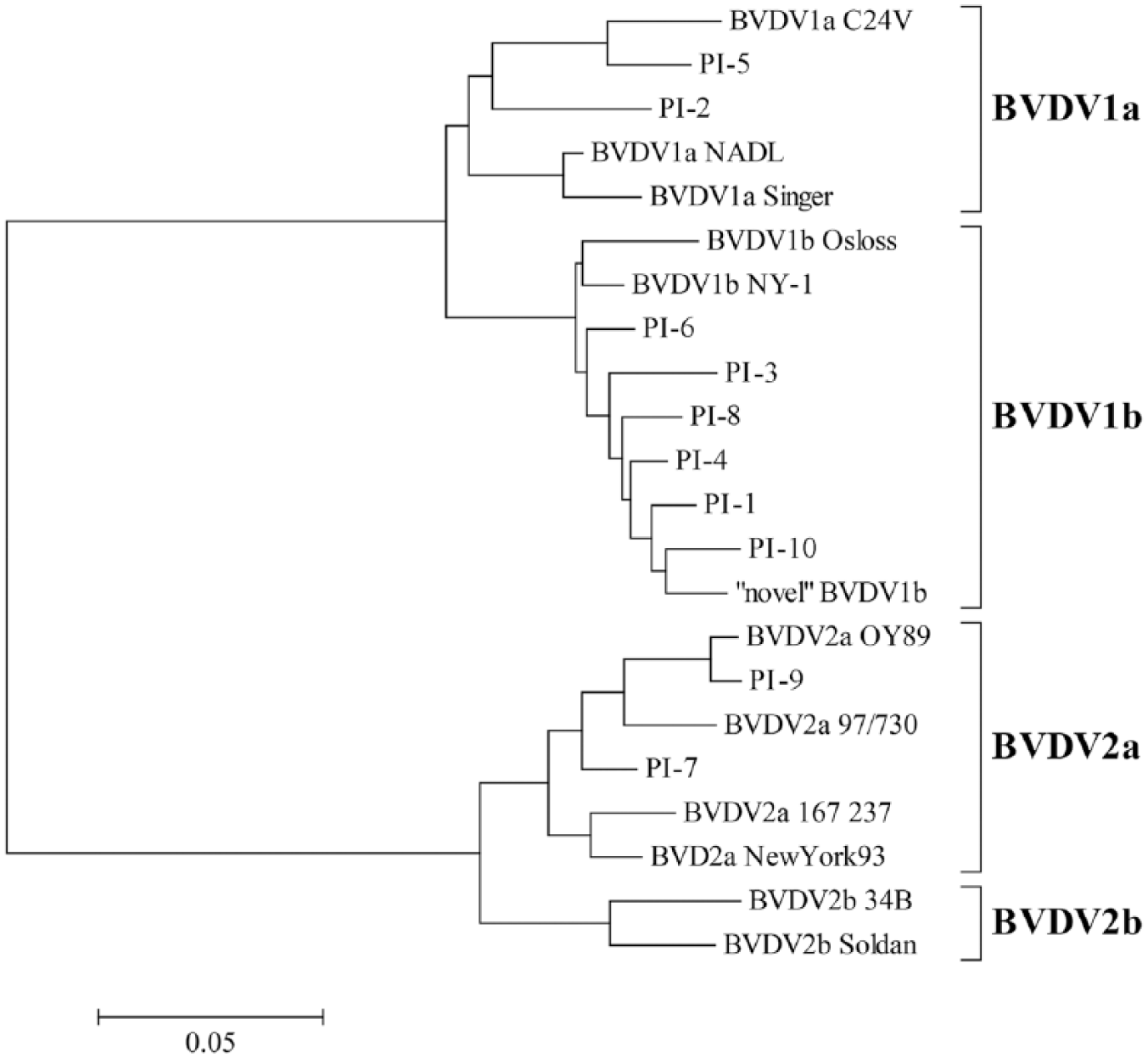
Phylogenetic tree of the 5′–untranslated region of bovine viral diarrhea virus (BVDV) that includes prototype strains of BVDV subgenotypes [and GenBank accessions] (BVDV-1a: C24V [AF091605], NADL [M31182], and Singer [DQ088995]; BVDV-1b: Osloss [M96687] and NY-1 [FJ387232]; BVDV-2a: OY89 [AB003621], 97/730 [AF026770], 167 237 [U65055], and NewYork93 [AF502399]; BVDV-2b: 34B [AF244952], and Soldan [U94914]), and subgenotype or strains of BVDV in PI cattle used in the commingle (BVDV-1a: PI-2, PI-5; BVDV-1b: PI-1, PI-3, PI-4, PI-6, PI-8, and PI-10; BVDV-2a: PI-7, PI-9), and the “novel” BVDV-1b strain.
The BVDV-positive status of all 10 PI cattle was confirmed several weeks prior to commingling and again 2 wk prior to commingling by RT-rtPCR on 3-mm ear notch samples using BVDV kit A (BoVir-SL bovine viral diarrhea virus real-time RT-PCR kit, Enfer Diagnostics, Manhattan, KS) and kit B (VetMAX-Gold BVDV detection kit, Applied Biosystems, Thermo Fisher Scientific, Austin, TX). The BVDV-positive status was also re-affirmed on the day of their arrival (d-1) by VI on buffy coats and nasal swab, and by RT-rtPCR with kit B on buffy coat, serum, and nasal swabs samples collected on d-1 and again on the last day of the study (d+27). The average threshold cycle (Ct; the first cycle at which fluorescence is detected above the baseline) values from 2 ear notch sampling dates (2 wk before the start of the study and on d+27) from all PI cattle used in the commingle was 25.9, indicating a high, stable viral load.
Fifty-three non-PI cattle, averaging 243 kg, with unknown vaccination history or BVDV-antibody status, were obtained from various stocker operations in Kansas that procure animals from multiple geographic locations across the United States. All of the non-PI cattle, on d-1, received a single prophylactic dose (1.5 mL/45 kg) of ceftiofur crystalline free acid (Excede, Pfizer Animal Health, New York, NY) and were weighed, assigned a study identification number, ear tagged, and the following samples collected: whole blood in EDTA tubes (Vacuette K3EDTA tube, Greiner Bio-One, Monroe, NC), serum in clot tubes (Vacuette serum clot activator tube, Greiner Bio-One), and nasal swabs in viral transport medium (Universal viral transport medium, BD Diagnostics, Sparks, MD). All samples were transported to the Kansas State Veterinary Diagnostic Laboratory (KSVDL) on ice within 1 h after collection.
Prior to commingling, the PI status of all 53 non-PI cattle was evaluated by RT-rtPCR on 3-mm ear notch samples using kit A. Subsequently, the BVDV-negative status of the non-PI cattle was affirmed by VI on buffy coat and nasal swab samples, and by RT-rtPCR on buffy coat, serum, and nasal swab samples collected on d-1 using the BVDV RT-rtPCR kit B. Kit B was used in all subsequent BVDV RT-rtPCR evaluations on samples collected during the commingle.
The commingling study was performed in a single dirt pen at the Kansas State University Cow Calf Unit–Junietta. Appropriate amounts of feed, water, and bunk space were provided for the cattle in accordance with standard feedlot practices. The animals were handled in accordance with the protocols approved by the Kansas State University, Institutional Animal Care and Use Committee (KSU-IACUC permit 2919).
After sample collection on d-1, all non-PI cattle were placed in the designated study pen. PI BVDV cattle were placed in a quarantine pen with no fence-line contact with non-PI cattle. The day after the non-PI cattle arrived, they were commingled with all PI cattle in a common study pen. All of the cattle remained in this pen (except during sampling times and morbidity treatments) for the remainder of the study and were observed twice daily for clinical signs of disease or injury using prescribed protocols. 18 Cattle that exhibited signs of illness (including lethargy, coughing, inappetence, difficulty breathing, excessive nasal discharge, or depressed appearance) were removed from the group and examined clinically, which included rectal temperature. Animals with rectal temperatures ≥40°C were treated with a subcutaneous dose of enrofloxacin (12.5 mg/kg; Baytril, Bayer Animal Health, Shawnee Mission, KS) and returned to the study pen.
Sample collection after commingling
Samples were collected from all non-PI cattle prior to collecting the samples from PI cattle to avoid any possibility of sample contamination. On d+1 through d+8, d+10, d+13, d+17, d+20, d+23, and d+27 post-commingling, whole blood, serum, and nasal swabs were collected from each non-PI animal. At the conclusion of the study (on d+27), serum, nasal swab, whole blood buffy coat, and ear notch samples were collected from all PI cattle. On d+8, d+13, and d+20, ear notch biopsies were obtained from each of the non-PI cattle. To minimize the potential for BVDV nucleic acid contamination between animal samplings, aseptic techniques were followed throughout the ear notching process. Within 1 h of collection, all samples were transported to the KSVDL on ice. Whole blood samples were processed immediately by centrifuging (2,350 × g, 10 min), and buffy coats were collected. Buffy coat cells were washed and aliquoted to a final volume of 0.5 mL in phosphate-buffered (0.01 M, pH 7.3) physiologic saline solution (PBS).
RNA recovery
RNA was recovered from serum (50 µL), buffy coats (100 µL), nasal swabs (100 µL), and ear notch samples (3 mm; Sklar Tru-Punch sterile disposable 3 mm biopsy punch, VWR International, Radnor, PA) for use in RT-rtPCR analysis. A magnetic bead–based nucleic acid extraction method was used to recover viral RNA following manufacturer’s instructions (MagMAX-96 viral RNA isolation kit, Applied Biosystems) using a 96 deep-well magnetic particle processor (MagMAX Express-96 deep well plates; MagMAX Express-96 magnetic particle processor, Applied Biosystems). The RNA control from kit B was added to each sample according to the manufacturer’s instructions, and the resulting elution was stored at −80°C.
BVDV RT-rtPCR
A one-step RT-rtPCR assay was performed using kit B to identify BVDV RNA in the serum, buffy coat, nasal swab, and ear notch samples. The RT-rtPCR assays were carried out in a 96-well reaction plate (MicroAmp optical 96-well reaction plate, Applied Biosystems), with each sample assayed in a 25-µL final reaction by following the kit manufacturer’s instructions (7500 Fast real-time PCR system, Applied Biosystems). Each plate also included 2 positive controls (BVDV control RNA; RNA control [10,000 copies] from kit B) and 2 negative controls (nuclease-free water). The RNA control was an internal positive control added to the lysis solution prior to RNA isolation to distinguish PCR inhibition, reaction failure, or human error from true negatives. RT-rtPCR data analysis was performed with the real-time PCR instrument’s software (7500 Fast system SDS v1.4 module, Applied Biosystems) using a manual Ct analysis in which the threshold is set at 10% of the final signal for the positive controls.
Prior to data analysis, the RT-rtPCR assay was validated by verifying the Ct values of positive and negative control reactions (BVDV control RNA target, Ct ≤ 28; RNA control target, Ct ≤ 31; and no-template control reaction, no signal or Ct = 40). The results were then evaluated, and interpretations were made as follows: BVDV-positive samples had BVDV test Ct values ≤38 with accompanying RNA control Ct values of 30–40; the sample was considered suspect BVDV-positive when the BVDV Ct value was 38–40 and RNA control Ct value was 30–40; the sample was considered BVDV-negative when the BVDV Ct was ≥40 and the RNA control Ct was 30–34. All samples that fell under the suspect BVDV-positive category (BVDV Ct values 38–40) and RNA control Ct out of acceptable range were subjected to further analysis as recommended in kit B.
Virus isolation
VI was performed on frozen (–80°C) whole blood buffy coat and nasal swab samples collected from each animal on d-1, d+8, d+13, and d+20. The samples were brought to room temperature, vortexed, and centrifuged (2,000 × g, 5 min). Supernatants were diluted 1:5 with cell culture growth medium, and 200 µL of each were inoculated in duplicate into 96-well plates that were freshly seeded with embryonic bovine lung cells. The BVDV NY-1 strain from KSVDL was used as the positive control, and a cell culture medium blank served as the negative control. After incubation (4 d, 37°C, 5% CO2) the plates were washed with 1× PBS (pH 7.4), fixed with 80% aqueous acetone, and stained with α-BVDV antibody (Bovine viral diarrhea virus monoclonal antibody [D89], VMRD, Pullman, WA) in an indirect fluorescent antibody (IFA) assay. After the plates were incubated (30 min, 37
Virus neutralization
The BVDV VN assays were used to detect BVDV antibody titers of non-PI cattle on d-1and on d+27. Six selected BVDV strains were used for VN (n = 5, from PI animals PI-1, PI-5, PI-7, PI-8, and PI-9; and the 6th strain, a “novel” BVDV-1b from non-PI animal 12) isolated from buffy coats. A working VN stock for all of the viruses was prepared in Madin–Darby bovine kidney (MDBK) cells; the infectivity level for the VN indicator viruses were determined, and frozen stocks were prepared for strain-specific antibody testing.
The BVDV VN antibody levels were determined using a varied serum–constant virus method and used a specific strain of BVDV as the indicator virus. Serial 2-fold dilutions of heat-inactivated serum were mixed with a constant quantity of the indicator BVDV (50–300 TCID50), incubated (1 h, 37°C), and 0.1 mL inoculated onto a monolayer of 1-d-old MDBK cells in 96-well plates. Cultures were incubated (3 d, 37°C, 5% CO2), then fixed (room temperature, 15 min) in 80% aqueous acetone, and the presence of BVDV was determined by IFA. The IFA procedure was conducted by incubation (30 min, 37°C) with 50 µL/well of undiluted α-BVDV antibody (D89), then washed in PBS, and 50 µL of diluted (1:75) fluorescein isothiocyanate conjugated goat anti-mouse IgG (Jackson Immuno Research Laboratories, West Grove, PA) was added to each well. The plates were incubated (30 min, 37°C), washed with PBS and 50 µL/well of 50% glycerol/1× PBS added. Immunofluorescence reactivity was observed with an inverted epifluorescence microscope equipped with a metal halide light source at 100× magnification. VN titers were determined on 4 replicates of serial 2-fold dilutions of the sera, based on 100% inhibition of the indicator virus and 50% endpoints, as determined by the Spearman–Kärber method. Cattle with titers ≤1:2 were considered seronegative to the indicator BVDV strain. Specific titer information was not assessed for cattle that had BVDV titers >1:4,096. Animals with at least a 4-fold rise in titer from the initial sampling date were considered to have seroconverted to the indicator BVDV strain. The geometric mean of d-1 BVDV VN titers was calculated using the specific titer of all non-PI cattle on d-1 (prior to commingling) to the 6 BVDV indicator viruses described previously.
BVDV genotyping
BVD viruses from all PI cattle were subjected to 5′-UTR sequence analysis to ascertain the BVDV genotype and strain distinction using RNA recovered from buffy coat lysates collected on d-1 and d+27. Non-PI cattle whole blood buffy coat samples collected on d+8, d+13, or d+20 that tested positive for BVDV by RT-rtPCR (Ct ≤ 38) or suspect-BVDV positive (Ct = >38 but <40) were processed for 5′-UTR sequence analysis. The buffy coat total RNA was recovered (QIAamp viral RNA mini kit, Qiagen, Valencia, CA) and was used in a RT-PCR to amplify a 293-bp fragment of the BVDV 5′-UTR region using primer sequences (BVD-SF: AGGCTAGCCATGCCCTTAGT; BVD-SR: CTCCATGTGCCATGTACAGC) modified from those described previously. 48 The 30-µL PCR reaction comprised 1× buffer, 0.33 µM of each primer, 0.33 mM of each dNTP, 4 units of RNase inhibitor (Qiagen), 5 µL of buffy coat RNA, and 1.2 µL of commercial RT-PCR mix (One-Step RT-PCR kit, Qiagen). The RT-PCR was completed as follows: 50°C, 30 min; 95°C, 15 min; followed by 40 cycles at 95°C, 20 s; 58°C, 30 s; 72°C, 45 s. The amplicons were analyzed (QIAxcel multi-capillary electrophoresis system, Qiagen) to confirm that only a single amplicon (293 bp) was generated. Samples positive for the predicted product size were purified (QIAquick PCR purification kit, Qiagen), and the concentrations were measured (Nanodrop 8000 UV-Vis spectrophotometer, ThermoFisher Scientific, Wilmington, DE). The resulting PCR product was diluted to the desired concentration and sequenced from both ends using an outsourced sequencing facility. Forward and reverse sequences were assembled, the assembled sequences were aligned, and pair-wise comparisons showing the percent identities were generated (CLC Main Workbench, CLC bio, Cambridge, MA). Assembled sequences were used to generate a neighbor-joining phylogenetic tree (MEGA 6, http://megasoftware.net/).
Statistics
Data were entered into Excel (Microsoft, Redmond, WA) and analyzed using a commercial statistical program (Stata 12 Reference A-F, StataCorp, College Station, TX). To investigate the association of non-PI cattle VN titers on d-1 with other parameters, linear regression accounting for repeated measures was used (xtmixed) with an autoregressive covariance structure. For comparison of the mean RT-rtPCR Ct value of positive non-PI cattle between test parameters, an independent sample t-test was used, assuming unequal variance. To assess the equality of the proportions of non-PI cattle BVDV RT-rtPCR–positive by sample type, a binomial probability test was completed. For all analyses, values of p ≤ 0.05 were considered to be statistically significant.
Results
Animal health during commingle
During the study, 43% (23 of 53) of the non-PI cattle were treated after exhibiting signs of BRD and rectal temperatures of ≥40°C (Supplementary Table 1). Seven of the treated cattle (17, 23, 27, 29, 32, 38, 40) continued to qualify as individual cases and were treated with enrofloxacin. On d+20, 1 animal (17) showed BRD signs, and was treated and returned to the study pen. Despite treatment, this animal continued to be ill for the remainder of the study and went into lateral recumbency and was unable to rise. The animal was euthanized, and autopsy and histologic examination at KSVDL revealed chronic-active necrosuppurative bronchopneumonia with fibrinous pleuritis affecting ~60% of the total lung fields with a cranioventral distribution. The tracheobronchial and mediastinal lymph nodes were markedly enlarged. The carpal, stifle, and hock joints had acute fibrinous arthritis. Pooled lung tissues and nasopharyngeal swabs from this animal were subjected to bovine respiratory PCR panel (real-time PCRs assaying for Mycoplasma bovis, bovine respiratory syncytial virus, bovine coronavirus, bovine herpesvirus 1 [BoHV-1], parainfluenza virus 3, and BVDV) at the KSVDL Molecular Diagnostic Laboratory. The samples were PCR-positive for M. bovis, BoHV-1, and BVDV. Attempts at VI from these samples were negative. Direct 5′-UTR sequencing from RNA recovered from pooled lung tissues revealed that animal 17 was infected with 2 subgenotypes of BVDV, the BVDV-1b strain from PI-8 and the BVDV-2a strain from PI-9.
Virus isolation
All PI cattle were VI-positive for BVDV on d-1, whereas all non-PI cattle were VI-negative on buffy coat and nasal swab samples collected on d-1. On d+8, 26% (14 of 53) of the non-PI cattle were VI positive for BVDV from buffy coats (Supplementary Table 1). Attempts to isolate BVDV from non-PI cattle by VI on d+13 as well as d+20 buffy coat samples were unsuccessful. Attempts to isolate BVDV from nasal swabs were successful in all of the PI cattle (d+27 swabs), but were unsuccessful for all of the non-PI cattle (d+8, d+13, and d+20 swabs).
BVDV RT-rtPCR from whole blood buffy coat, serum, nasal swabs, and ear notch samples
Approximately 52% (384 of 742) of the buffy coat samples, 26% (191 of 742) of the serum samples, and 24% (174 of 742) of the nasal swab samples collected from non-PI cattle during the commingle were BVDV RT-rtPCR positive. Of the 53 exposed non-PI cattle, 52 (98%) were BVDV-positive from buffy coat samples, 46 (87%) were BVDV-positive on serum samples, and 49 (92%) were BVDV-positive on nasal swab samples, by RT-rtPCR on at least one sampling date. Animal 42 was the only animal that was RT-rtPCR negative on all buffy coat sampling dates after commingling (but was low/suspect PCR-positive at least one time on other sample types). Approximately 82% (130 of 159) of the total ear notch samples collected on d+8, d+13, and d+20 of the commingle were BVDV RT-rtPCR negative/undetected. Ear notch samples were RT-rtPCR positive in 8 of the non-PI cattle (15%); however, only animal 28 was RT-rtPCR positive on more than one ear notch collection day (on d+8 and d+13).
The RT-rtPCR assay for detection of viral RNA was conducted at 14 time points. The range of time points for which the cattle were positive for viral RNA ranged from 0 to 13 (Supplementary Table 1). The number of non-PI cattle (93%, 49 of 53) positive for BVDV RNA by RT-rtPCR results peaked on day 8 post-commingle. Approximately 8% (61 of 742) of the buffy coat samples, 7% (53 of 742) of serum samples (Supplementary Table 3), and 17% (125 of 742) of nasal swab samples were identified as BVDV suspect-positive (Ct values ≥38.0 but <40). The RT-rtPCR suspect-positive results from these samples were generally recognized at the beginning and/or the end of an individual animal’s BVDV infection period, with the exception of nasal swab samples, for which no pattern could be associated for suspect samples.
Plotting the average BVDV RT-rtPCR Ct values of all non-PI buffy coat, serum, and nasal swab samples collected post-commingling indicated that the average Ct value of all non-PI cattle became suspect-positive by d+1 (Ct ~39), positive by d+4 to d+7 (Ct < 38), peaked between d+8 and d+13, became suspect-positive nearly 2 wk after commingling, and then negative by d+23 (except for buffy coats, the majority of which remained suspect-positive for the remainder of the study). The average Ct for all positive ear notch samples was 37.1, and the lowest BVDV-positive Ct value detected from an ear notch sample was from animal 40 on d+8 (Ct 34.4). Seventeen non-PI cattle were identified as RT-rtPCR suspect-positive on ear notch samples, and 10 of those cattle were from samples collected on d+8. Three cattle (3, 18, 32) were RT-rtPCR suspect-positive on consecutive ear notch collections; however, none of the cattle were found to be RT-rtPCR suspect-positive on all 3 ear notch sampling dates.
During the course of the commingle period, the percentage of buffy coat samples that were BVDV RT-rtPCR positive (52%, 384 of 742) was significantly higher (p ≤ 0.05) than either nasal swab samples (24%, 174 of 742) or serum samples (26%, 191 of 742).
Genotyping of BVDV strains
The 5′-UTR BVDV sequencing results on buffy coat lysates collected from all PI cattle on d-1 and d+27 indicated that the strain sequences were consistent with those identified on initial characterization of the viruses.
Sequencing buffy coat lysates from d+8 indicated that 38 cattle were infected with a single strain of virus, 4 cattle with multiple strains that could be differentiated, and 8 cattle with multiple BVDV strains that could not be differentiated (Supplementary Table 2). One BVDV suspect (animal 6, Ct 38.5) was infected with a single strain of BVDV. Two cattle were BVDV RT-rtPCR positive (11, Ct 30.8; 25, Ct 37.3), but the sequence could not be obtained. Mixed BVDV infections were defined by different genotypes obtained by repeated sequencing attempts or the chromatogram of a strain displaying multiple peaks in multiple nucleotide sites in multiple sequencing attempts.
The genotype or strain breakdown of the d+13 RT-rtPCR positive and suspect-positive cattle indicated that 73% (33 of 45) of those cattle were infected with a single BVDV strain. Three cattle were confirmed to have infections with >1 BVDV strain, 8 cattle had 1 strain of BVDV defined but sequencing trace data could not define the additional strain(s), and 1 animal was RT-rtPCR positive (Ct = 35.6) but sequencing was not attempted (Supplementary Table 2).
The genotype or strain breakdown of the d+20 RT-rtPCR positive and suspect-positive cattle showed that 20 animals were infected with a single BVDV strain, 8 cattle were infected with >1 BVDV strain, 6 cattle had 1 strain defined but had undefined evidence of mixed infections, 2 animals were infected with 3 different BVDV strains, and 1 animal was RT-rtPCR positive (Ct 36.3) but no sequence was obtained (Supplementary Table 2).
Sequencing results demonstrated that BVDV-2a was the most frequently detected genotype in exposed non-PI cattle (41 of 53). BVDV-2a strain from PI-9 accounted for a large majority of infections (34 of 53); the other 2a strain from PI-7 was detected in 10 non-PI cattle. BVDV-1a sequences were detected in 14 non-PI cattle, of which 13 were infected with BVDV-1a strain from PI-5; BVDV-1a strain from PI-2 was detected in 1 animal (2) and only once (d+20; Table 1). BVDV-1b strains from PI cattle were detected in 12 non-PI cattle (Table 1), of which 1b strains from PI-8, PI-1, and PI-4 were detected in 5, 4, and 3 cattle, respectively. Sequences for BVDV-1b strains from PI-3, PI-6, and PI-10 were undetected in any buffy coat samples (d+8, d+13, or d+20) from any of the non-PI cattle.
Frequency of bovine viral diarrhea virus (BVDV) genotypes, subgenotypes, and strains detected by direct 5′–untranslated region sequencing of nucleic acid from whole blood buffy coat samples of non–persistently infected (PI) cattle collected on 3 dates (days 8, 13, and 20) post-commingling with PI cattle.
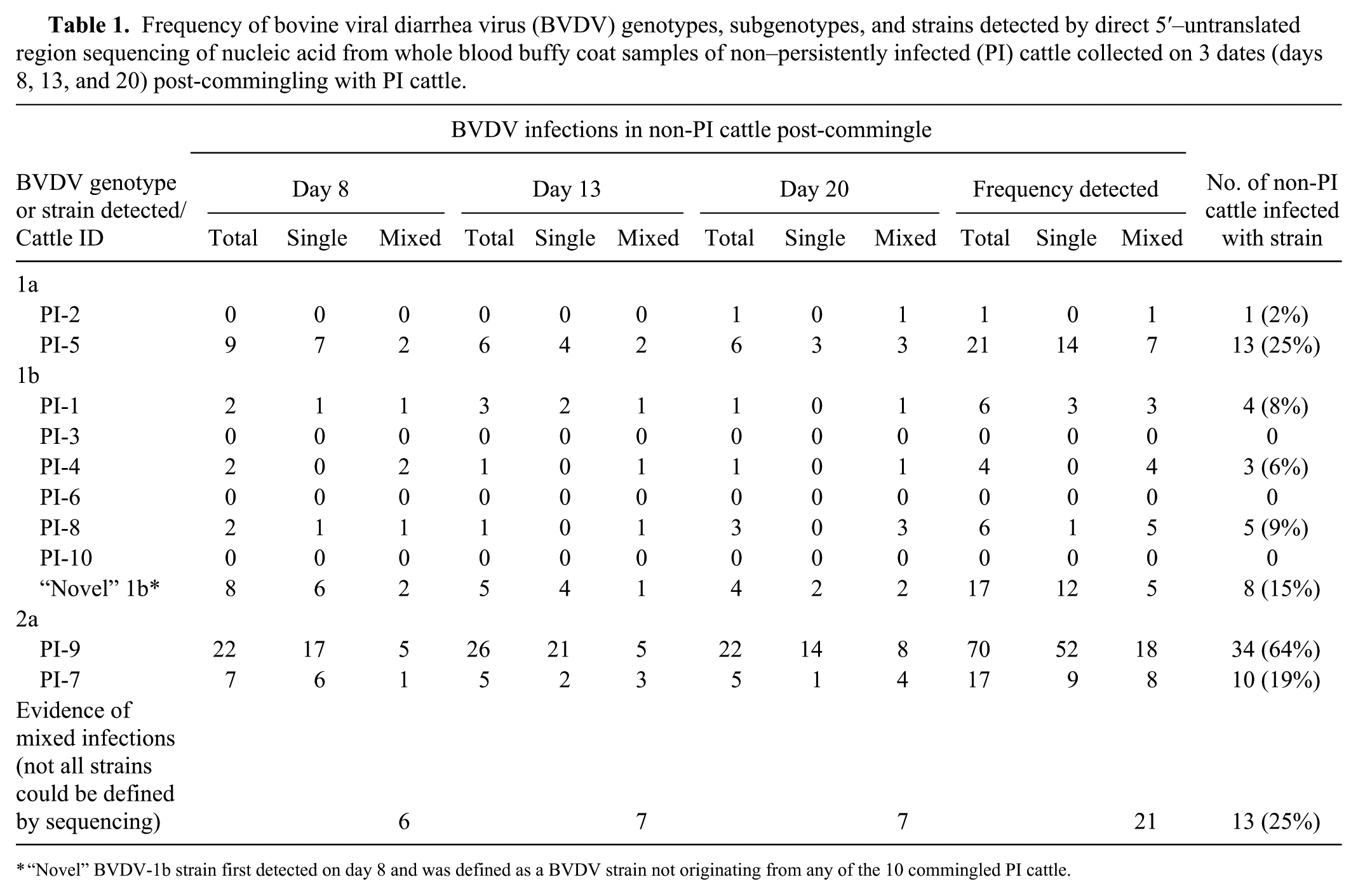
“Novel” BVDV-1b strain first detected on day 8 and was defined as a BVDV strain not originating from any of the 10 commingled PI cattle.
“Novel” BVDV-1b strain
On d+8, a BVDV-1b strain was identified in buffy coat cells that did not correspond to BVDV sequences from any of the 10 PI cattle (Fig. 1). It was subsequently identified in 8 different cattle (12, 14, 19, 26, 40, 46, 47, 48) over the 3 sampling dates (d+8, d+13, and d+20; Supplementary Table 2). This “novel” BVDV-1b strain was detected in 8 non-PI cattle as single strain infections or as mixed infection in multiple sampling dates (Supplementary Table 2).
To evaluate the BVDV that resulted in d+1 BVDV infections, buffy coat samples from 5 cattle (12, 19, 26, 41, 47) collected on d+1 through d+7 were selected for further 5′-UTR sequence analysis. Sequencing results identified animal 26 as infected with the “novel” BVDV-1b strain on d+1 through d+7. Animal 26 appeared to be infected with the “novel” BVDV-1b as a single strain infection until d+20 when a mixed infection with BVDV-2a strain from PI-9 was also demonstrated. The remaining 4 cattle (12, 19, 41, 47) had evidence of BVDV sequences on d+1, but the genotype and strain designation could not be clearly defined by sequencing after repeated attempts. Sequencing of d+2 samples demonstrated the presence of the “novel” BVDV-1b strain in 3 of these cattle (19, 41, 47). Animal 12 buffy coat samples continued to show evidence of undefined BVDV sequences until d+3 when it was possible to define the “novel” BVDV-1b strain sequence. The presence of the “novel” BVDV-1b strain continued to be detected in all 5 cattle through d+7, except for animal 41 who demonstrated the “novel” BVDV-1b through d+6 and then became BVDV RT-rtPCR negative or undetected on d+7.
All remaining animals that were found to be infected with the “novel” BVDV-1b on the 3 original sequencing dates were BVDV RT-rtPCR negative until d+3 (14, 40), d+4 (48), or d+5 (46; Table 1). Although not evaluated by 5′-UTR sequencing on samples from d+1 through d+7, animal 49 was suspected of being infected with the “novel” BVDV-1b strain early in the study, similar to animal 41.
Virus neutralization
VN results indicated that 66% (35 of 53) of the non-PI cattle were seronegative (≤1:2) on d-1 to all BVDV indicator viruses (Supplementary Table 3). Five of the non-PI cattle (6, 25, 38, 42, 43) began the study with VN titers ≥1:16 to all of the indicator viruses and 1 animal (8) had d-1 VN titers ≥1:16 to all 3 of the BVDV-1b indicator viruses (Supplementary Table 3).
Approximately 80% (35 of 44) of the d-1 VN seronegative (≤1:2) or low titered (<1:16) cattle seroconverted (>4-fold rise in titer) to all of the BVDV indicator viruses (Supplementary Table 3). Overall, seroconversion titers were variable to the VN indicator viruses; some animals responded with high titers to all indicator viruses, while some responded with high titer to only 1 or 2 of the indicator viruses.
Non-PI cattle that began the study with higher BVDV VN titers to the BVDV indicator viruses demonstrated either a lack of detectable BVDV infection during the commingle period as interpreted by BVDV RT-rtPCR from buffy coat lysates, or a reduced duration of infection when compared to cattle whose BVDV VN titers were negative or had low titers (Supplementary Table 3). VN titers on d-1 were associated with several parameters. For each increase in VN value by 1 on d-1, the VN value on d+27 was increased by 7.46 values, and there was a negative association with VN values on d-1 and number of test days RT-rtPCR positive for all samples types including buffy coat, nasal swab, and serum samples.
When the cumulative buffy coat BVDV RT-rtPCR number of positive test days from individual cattle was compared to the d-1 VN geometric mean titers (GMT) to the 6 indicator BVDV strains, it was possible to inversely correlate a reduction in positive BVDV RT-rtPCR test days with an elevated d-1 VN GMT. Similarly, a lower level of detectable BVDV, as indicated by higher average Ct values generated from buffy coat samples over all the PCR test days, could also be correlated to an elevated d-1 VN GMT. Animal 42 had a d-1 VN GMT of 574.7 as measured against the 6 BVDV indicator viruses, and BVDV was undetected from buffy coat samples on all PCR test dates (Supplementary Table 3). Similarly, 4 cattle (6, 18, 25, 43) had d-1 VN GMTs of 128–287, and each was PCR positive from a buffy coat sample on only a single PCR test date (Supplementary Table 3). This contrasted with the 48 other non-PI cattle whose d-1 VN GMTs averaged 1.8 (range: <2–20.2), and the average number of test days that cattle were PCR positive for BVDV was ~7.7 (range: 3–13).
Discussion
Commingling of non-PI cattle with PI cattle resulted in rapid transmission of BVDV and transient infections, indicated by at least one RT-rtPCR–positive result, in all non-PI cattle. The transmission of BVDV from PI cattle to non-PI cattle was further confirmed by 5′-UTR sequencing and VN assays. After commingling of the PI cattle with non-PI cattle, 45% (23 of 53) of the non-PI cattle showed clinical signs of the disease on at least one occasion and required treatment. In our study, 6 cattle required treatments for 4–10 d, and 1 animal developed severe BRD and was euthanized. Although identifying other pathogens associated with BRD was not a major objective of our study, it is very probable that cattle that demonstrated clinical signs requiring treatment were clinically ill because of the synergistic effect of BVDV with other BRD pathogens. 40 This was clearly demonstrated with animal 17 that was euthanized and subsequently identified to be infected with M. bovis, BoHV-1, and 2 subgenotypes of BVDV (BVDV-1b from PI-8 and BVDV-2a from PI-9).
In previous studies, the severity and duration of a BVDV infection have been shown to be inversely proportional to the levels of VN antibodies against BVDV. Calves with passively acquired BVDV VN antibody titers of <8 developed severe clinical disease; whereas, calves that had BVDV VN titers of ≥16 but <256, although not ill, did replicate BVDV that could be detected as viremias. 6 Such studies assessed BVDV viremias by VI and did not utilize molecular tests such as RT-rtPCR assays. In our study, duration of BVDV RT-rtPCR–positive status on buffy coat samples in an animal was inversely correlated with the presence of higher BVDV VN titers at the onset of commingling with PI cattle. This suggests that, although not sufficient to elicit sterile immunity to BVDV or to totally prevent viremias or infection, cattle with the higher VN titers would have short-spanned viremia and reduced amounts of circulating BVDV in their blood.
Significantly more (p ≤ 0.05) BVDV RT-rtPCR positives were detected from buffy coat samples than serum or nasal swabs. A majority of the individual non-PI cattle demonstrated a pattern of increased BVDV RNA detection, as indicated by lower Ct values, which then peaked and subsequently decreased over the course of the infection. Nasal swab samples differed from buffy coat and serum samples in that a brief spike in BVDV RT-rtPCR–positive samples was observed on d+2 post-commingle, only to subside, then resurge on d+6, and peak between d+8 and d+10. The frequency of suspect-positive BVDV RT-rtPCR results was higher with nasal swabs (18%) than with either buffy coat (8%) or serum samples (7%). Suspect nasal swab samples did not indicate any pattern of either early or late stages of BVDV infection, as was identified in buffy coat or serum samples. Suspect results on nasal swab samples might not necessarily represent systemic infection in that individual but rather the presence of BVDV in that sampling or the environment and could potentially aid in BVDV surveillance in a herd.
Detection of BVDV in ear notches by RT-rtPCR from some of the non-PI cattle correlated with previous reports.11,20 Interestingly, of all of the sample types tested, only ear notches showed the ability to distinguish PI cattle (average Ct 25.9) from transiently infected cattle (average Ct 37.1; lowest Ct 34.4) into 2 distinct non-overlapping Ct populations. Although this particular observation warrants further investigation, it would appear prudent to confirm PI status of an individual through follow-up confirmatory BVDV RT-rtPCR testing on samples collected 3 to 4 wk later.11-13
VI from buffy coats, considered to be the “gold standard” for detecting acute BVDV infections,14,24,43 was not as effective as the RT-rtPCR for detecting an acute BVDV infection. The success rate for VI in the current commingle was similar to another commingle study. 17 VI was successful only on 29% of RT-rtPCR buffy coat samples (14 of 49) collected on d+8. At d+8 after commingle, the number of viremic non-PI cattle (as identified by RT-rtPCR) peaked as observed with previous studies.35,37,50
Acute mixed BVDV infections have been demonstrated in natural22,26,42 and experimental infections7,25,32 and are implicated for change in BVDV biotypes by recombination events,28,38 and as the basis for fatal mucosal disease in PI cattle.3,5 In our study, by 5′-UTR sequencing, single and various combinations of mixed infections were observed between different genotypes (BVDV-1 and BVDV-2), subgenotypes (BVDV-1a and BVDV-1b), or different strains within a subgenotype. BVDV-2a appeared to be the predominant subgenotype infecting the non-PI cattle. Predominant BVDV strains could also be identified within each subgenotype; however, the exclusion of BVDV-1b strains from 3 PI cattle was unexpected. These findings suggest that BVDV originating from some PI cattle would not compete as effectively as other PI strains in causing transient BVDV infections. This finding was similar to another commingle study in which multiple PI cattle were used concurrently. 30 In that study, variations in the infectivity of BVDV from PI cattle were attributed to differences in the virulence of the PI BVDV strains or to differences in socialization patterns of the PI cattle and how that might affect transmission to non-PI cattle.
A dual mechanism of BVDV superinfection exclusion for acutely infected cells in tissue culture has been reported; one at the level of viral entry involving a blocking mechanism of structural protein E2, and the other at the level of viral replication. 29 Whether similar exclusion mechanisms might have interfered with the replication of certain strains of BVDV in non-PI cattle in our study can only be speculated. It may be possible that multiple PI cattle could serve as a source of viral genetic variants that may be “selected” by non-PI cattle when they become infected. 4 Combining RT-rtPCR and direct sample sequencing approaches in future commingling studies would afford the opportunity to evaluate natural transmission, selection, and virulence of BVDV strains in acute BVDV infections.
Unexpectedly, a BVDV-1b strain, referred to as the “novel” BVDV-1b strain, which did not correspond to any of the strains from PI cattle, was detected in 8 non-PI cattle on the 3 sampling dates selected for 5′-UTR sequencing. Based on the BVDV RT-rtPCR interpretations and sequences generated from buffy coat lysates, 6 non-PI cattle appeared to have been exposed to and became infected with this “novel” BVDV-1b strain just prior to the commingling. Transmission of the “novel” strain from these 6 cattle could have resulted in a transmission chain of acute BVDV infections that were identified on d+8, d+13, and d+20 in other non-PI cattle. Given that all non-PI cattle were BVDV RT-rtPCR negative from all sample types evaluated prior to the commingle on d-1, it seems plausible that animals identified as being infected with the “novel” strain on d+1 became infected in the movement period preceding their commingling with the PI cattle. The inadvertent introduction of unintended BVDV strains into large PI commingle studies has been reported previously,17,20 and stresses the importance of enforcing rigorous biosecurity requirements to prevent BVDV transmission during cattle movement. Quarantining or isolating all new or replacement animals before commingling with resident cattle as well as retesting has been suggested where BVDV eradication programs have been implemented. 23
Footnotes
Acknowledgements
Heather Sloan and Taghreed Mahmood are acknowledged for their excellent technical assistance. We also thank Drs. Ben Hause and Ivan Leyva-Baca for their critical review of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Financial support for this study was provided by Applied Biosystems (Thermo Fisher Scientific Corporation) (KSU Grant BG1344) by supplying reagents, supplies, salary support for Dr. L Peddireddi; by the Kansas State Veterinary Diagnostic Laboratory; and by the Beef Cattle Institute at Kansas State University for salary support for Dr. K Foster and for cattle care.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
