Abstract
Four Newcastle disease virus (NDV) strains were isolated from domestic, commercial geese that showed clinical signs that were believed to be the result of NDV infections. The genetic, antigenic, and pathogenic characteristics of the 4 NDVs were compared with those of NDV strains that were isolated from chickens. The complete genomes of 2 of the NDV strains contained 15,186 nucleotides (nt); the other 2 contained 15,192 nt, and exhibited the typical genomic organization of genotype II NDV and molecular characteristics of VIId NDVs. Phylogenetic analysis confirmed that the genotype II and VIId NDVs that were isolated from geese belonged to the same clusters as the corresponding genotypes of the chicken isolates. A serologic assay demonstrated that the antigenic relatedness among the NDVs was associated with their genotypes, rather than their hosts, and that amino acid substitutions in the F and/or HN proteins may contribute to the antigenic differences among these NDV genotypes. Geese infected with genotype VIId NDVs that were isolated from geese and chickens showed similar pathologic characteristics. NDVs that were isolated from geese did not differ in genetic, serologic, and pathogenic characteristics from those isolated from chickens, indicating that these NDVs were derived from chicken NDVs. Given the significance of geese in NDV epidemiology, effective biosecurity measures should be adopted to prevent the interspecies transmission of NDVs.
Introduction
Newcastle disease is an acute and highly contagious viral disease that causes severe economic losses within the poultry industry. The etiologic agent of the disease is species Newcastle disease virus (NDV; family Paramyxoviridae, genus Avulavirus). Genus Avulavirus includes 9 serogroups of avian paramyxoviruses (APMV-1–9). Newcastle disease caused by NDV (also known as APMV-1) remains the most economically important viral disease within the poultry industry.1–3,14 It has been widely reported that a variety of NDV strains cause highly contagious respiratory, enteric, and neurologic diseases in various bird species.1,3 NDV has a single-stranded, negative-sense, nonsegmented RNA genome of 15,186, 15,192, or 15,198 nucleotides (nt) that contains 6 genes that encode the nucleoprotein (NP), phosphoprotein (P), matrix protein (M), fusion protein (F), hemagglutinin-neuraminidase (HN), and large RNA-directed RNA polymerase (L).10,29 Based on genomic size and a phylogenetic analysis of the F gene, NDV strains can be divided into 2 distinct clusters, class I and class II, within a single serotype. Class I and class II viruses have been recently classified into 1 and 18 genotypes, respectively.9,11 Distinct NDV genotypes were involved in each of the 4 major NDV panzootics that have occurred since the first report of Newcastle disease in 1926. 3
In addition to domestic avian species, at least 241 bird species can be infected naturally or experimentally with NDV. All bird species are probably susceptible to NDV infection; however, the outcome of infection may vary considerably in different species. 2 In domestic poultry, chickens are considered to be the most susceptible and natural host of NDV, and virulent NDV strains can cause serious disease in chickens, with almost 100% mortality.1,3 Waterfowl are considered to be a natural reservoir for NDV in addition to caged pet birds, pigeons, doves, and cormorants.1–3 Most NDV isolates obtained from waterfowl are categorized into class I and genotype I of class II, and they exhibit low virulence in chickens.20,22,26,35 Moreover, waterfowl, such as geese and ducks, which are infected with NDV strains that are virulent in chickens, do not typically show any clinical signs.1,3 However, in 1997, an outbreak of Newcastle disease in goose flocks, which resulted in high morbidity and mortality, was first reported in southern and eastern China. 37 Further analysis indicated that the goose NDV isolate Ch/97-1 might have originated or evolved from a chicken VIId NDV isolate that was introduced into China from Taiwan, China. 39 In addition, a similar disease was reproduced by experimentally infecting geese with NDV isolates from geese, as well as other genotype VI, VII, VIII, and IX NDVs. 26 Since the first reported Newcastle disease outbreak in a goose flock caused by genotype VIId NDV in 1997 in China, this kind of virulent NDV has been isolated frequently from chicken and goose flocks.6,23,24 The natural outbreaks and experimental reproduction of Newcastle disease in geese suggest that geese may play a critical role in the epidemiology of NDV. 34
Epidemiologic investigations showed that genotype VIId virulent NDVs have been circulating continuously in commercial chicken and goose flocks in China since the late 1990s.17,22,24,31,35 More significantly, adaptive evolution of genotype VIId NDVs via mutations that alter specific amino acids has been reported.8,31,35 Whether genotype VIId NDV isolates in chickens and geese evolved into separate phylogenetic clades, such as pigeon-specific NDVs (pigeon paramyxovirus-1), needs to be further elucidated. In addition, correlation analyses between avirulent NDV strains that were isolated from geese and chickens are scarce, despite the fact that avirulent NDVs are constantly isolated from goose and chicken flocks.22–24 In our study, whole genomic sequences of representative strains from domestic commercial goose and chicken flocks were analyzed genetically to elucidate the relationships and differences among genotype II and VIId NDV strains. Furthermore, the pathogenesis of genotype II and VIId NDV strains that were isolated from chickens and geese were compared in geese. In addition, the antigenic relatedness of genotype II and VIId NDVs that were isolated from different hosts were evaluated using cross–hemagglutination inhibition (HI) and cross–virus neutralization (VN) assays.
Materials and methods
Viruses, eggs, and animals
The NDV strains go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, go/CH/HLJ/LL01/08, and go/CH/LFJ/1/06 were isolated from domestic commercial geese that were not immunized with NDV vaccines. Some geese showed clinical signs in the flocks from which the NDVs were isolated. Some geese died in the flocks from which strains go/CH/LHLJ/1/06 and go/CH/HLJ/LL01/08 were isolated. Strain ck/CH/LHLJ/1/06 was isolated from commercial chickens that were immunized with the live La Sota vaccine using 10-d-old specific pathogen–free (SPF) embryonated chicken eggs (Harbin Veterinary Research Institute, Harbin, China) via the allantoic cavity. The NDV vaccine strain La Sota was also used in our study. The NDV strains go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, and ck/CH/LHLJ/1/06 were plaque-purified in primary chicken embryo fibroblasts, and NDV strains go/CH/LHLJ/2/06 and go/CH/LFJ/1/06 were purified by the end-point dilution method using embryonated SPF chicken eggs. A stock of each purified virus was produced by inoculating each virus into embryonated SPF chicken eggs via the allantoic cavity and collecting the infectious allantoic fluid at 48 or 72 h post-inoculation. The titers of the viruses were determined by inoculating 10-fold dilutions into groups of five 9-d-old SPF embryonated chicken eggs. The 50% embryo infectious dose (EID50) was calculated by the method of Reed and Muench (Table 1). 32
Characteristics of the 6 Newcastle disease virus strains used in this study.*
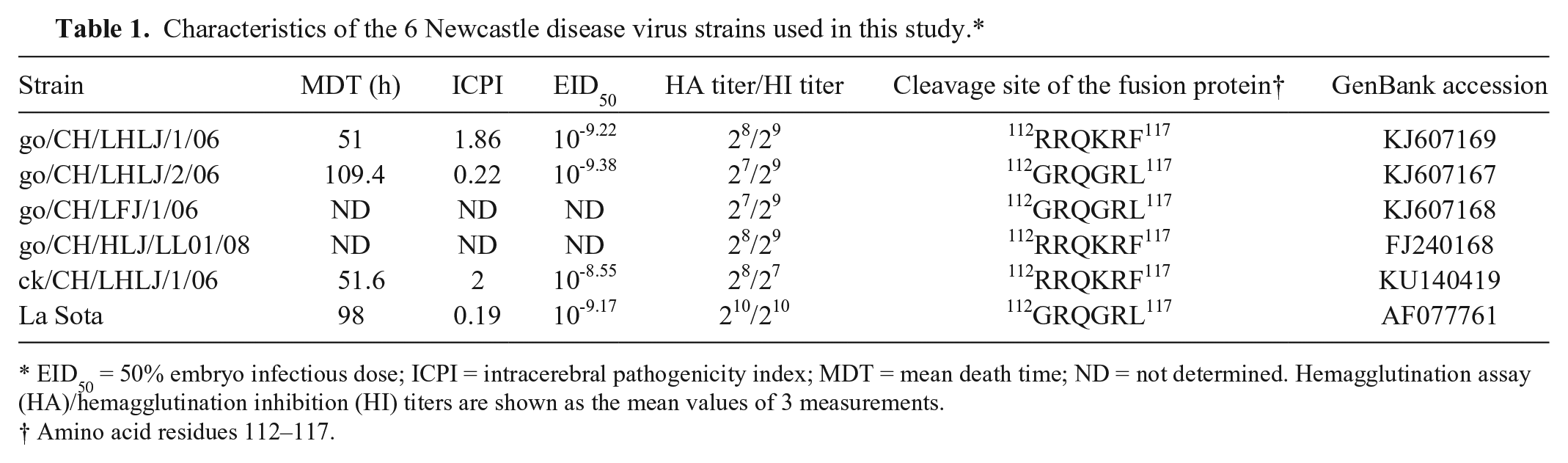

EID50 = 50% embryo infectious dose; ICPI = intracerebral pathogenicity index; MDT = mean death time; ND = not determined. Hemagglutination assay (HA)/hemagglutination inhibition (HI) titers are shown as the mean values of 3 measurements.
Amino acid residues 112–117.
Healthy 1-d-old female commercial Chinese geese were obtained from the Laboratory Animal Center, HVRI, Chinese Academy of Agricultural Sciences, China. The birds were housed in isolators with negative pressure, and food and water were provided ad libitum until they were 40-d-old. One week before experimental NDV infections, the sera of the birds were confirmed to be negative for goose parvovirus, avian influenza virus subtypes H5, H9, H7, and anti-NDV maternal antibodies using an enzyme-linked immunosorbent assay and a HI test, respectively.
RNA extraction, reverse transcription polymerase chain reaction, and genome sequencing
Viral RNA was extracted (RNAiso Plus reagent, TaKaRa, Shiga, Japan) from 200 μL of the viral stocks according to the manufacturer’s instructions. Fourteen pairs of primers, including those used for amplifying the 3′- and 5′-ends of the genomic RNA, were designed based on the consensus sequences of the published complete genome sequences of NDVs, as described previously. 36 The complete viral genome was amplified (PrimeScript one step RT-PCR kit, TaKaRa) according to the manufacturer’s instructions. Extraction and purification of PCR products were performed (E.Z.N.A. gel extraction kit, Omega Bio-Tek, Jinan, China) according to the manufacturer’s instructions. The purified PCR products were cloned into the T vector (pMD18-T vector, TaKaRa), and each fragment of the viral genome was sequenced at least 5 times to determine a consensus sequence.
GenBank accessions
The full genomic sequences of strains go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, go/CH/HLJ/LL01/08, go/CH/LFJ/1/06, and ck/CH/LHLJ/1/06 have been deposited in GenBank under accessions KJ607169, KJ607167, GU143550, KJ607168, and KU140419, respectively.
Sequence comparisons and phylogenetic analysis
The nucleotide sequences of our NDV strains were assembled, aligned, and compared with those of other NDV reference strains (Lasergene, DNASTAR, Madison, WI). BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) was used to search GenBank for homologous gene sequences. Sixty NDV reference strains with complete genomic sequences were selected for phylogenetic analysis based on the results of the BLAST search, as we wanted to investigate the epidemiologic link among the NDVs that were isolated from geese and other avian species. Of these viruses, 26 were isolated from geese and 43 were isolated from chickens. The characteristics of these NDVs are shown in Supplemental Table 1. A phylogenetic analysis was conducted on the basis of the F gene and the complete genomic sequences of 65 NDV strains by the neighbor-joining method with 1,000 bootstraps (MEGA v.6.0, http://www.megasoftware.net).
Pathogenicity tests
The pathogenicity of NDV strains go/CH/LHLJ/2/06 and La Sota, and go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06, which represent genotype II and VIId strains, respectively, was assessed by determining the intracerebral pathogenicity index (ICPI) in 1-d-old chickens and the mean death time (MDT) in 9-d-old SPF embryonated chicken eggs.
Animal experiments
Forty-six 40-d-old geese were randomly divided into 5 groups and housed in different isolators with negative pressure. Groups 1–4 contained 10 geese each, and group 5 had 6 geese. Geese in groups 1–4 were given inoculations of the go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, ck/CH/LHLJ/1/06, and La Sota strains, respectively, through the intranasal and intraocular routes with a dose of 106 EID50 in 0.1 mL of diluent per goose. Geese in group 5 were mock-inoculated with sterile phosphate-buffered saline (PBS) as a negative control. The animal experimental procedures were approved by the Ethical and Animal Welfare Committee of the HVRI, China. All experiments were conducted at appropriate biosafety levels. Geese were monitored daily for clinical signs. Morbidity and mortality were observed for 4 wk post-challenge. Serum samples and oropharyngeal and cloacal swabs were collected at 0, 4, 8, 12, 16, 20, 24, and 28 d post-inoculation (dpi). Tissue samples, including brain, Harderian gland, lung, trachea, liver, spleen, proventriculus, the bursa of Fabricius, kidney, and cecal tonsil, were collected from dead geese for the detection of viral loads and a histologic examination, as described previously. 12 After 4 wk, a booster dose of 106 EID50 of each virus was administered by intravenous inoculation to each bird in groups 1–4 prior to serum preparation. Geese were exsanguinated 1 wk after the last inoculation, and serum samples were collected. Sera of uninfected geese were collected and used as negative controls. All serum samples were inactivated at 56°C for 30 min and stored at −80°C until used.
Real-time RT-PCR
Equal amounts (100 mg) of tissue samples were homogenized in 400 μL of PBS, and total RNA was extracted (RNAiso Plus reagent, TaKaRa) according to the manufacturer’s instructions. RNA was quantified and qualified using a spectrophotometer (Nano Vue Plus, GE Healthcare, Piscataway, NJ). The obtained RNAs (2 μg) were used to evaluate the viral load of NDV in tissue samples using real-time reverse transcription (RT)-PCR, as described previously. 15 The primers and probe were designed based on the M gene sequence of the NDV strains used in this study. The primers and probe were as follows: forward (5′-CTCAGTGATGTGCTCGGACC-3′), reverse (5′-CCTGGGGAGAGGCATTTGCTA-3′), and probe (5′-[FAM]TTCTCTAGCAGTGGGACAGCCTGC[BHQ1]-3′). Real-time RT-PCR was performed on a LightCycler (Roche Diagnostics, Mannheim, Germany) using the PrimeScript one step RT-PCR kit (TaKaRa). The real-time PCR was performed as follows: RT at 42°C for 5 min, denaturation at 95°C for 10 s, and 40 cycles at 95°C for 5 s and 60°C for 20 s. All of the samples were tested in triplicate in each reaction. Standard templates with known concentrations, as well as a no-template negative control, were used for the quantitative analysis.
Virus recovery
Oropharyngeal and cloacal swabs were collected from each goose and used for virus recovery as previously described. 25 Each of the individual samples was inoculated into at least three 10-d-old SPF embryonated chicken eggs via the allantoic cavity. NDV viral particles in the allantoic fluids of inoculated eggs were identified using a HI analysis with a specific antibody.
Serologic assay
The HI and cross-HI assays were performed as described previously using 4 hemagglutinating units of antigen (go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, ck/CH/LHLJ/1/06, and La Sota) and their respective anti-serum. R values were calculated using the formula of Archetti, 4 and the antigenic relationship between 2 viruses was evaluated based on the R values.4,21,36
The cross-VN assay was performed to further determine the antigenic relationships among these 4 NDV strains. Each equal volume of 2-fold serially diluted serum was mixed with a constant amount of virus (100 EID50) and incubated for 1 h at 37°C. Then, 0.2 mL of each virus–serum mixture was inoculated into five 10-d-old SPF chicken embryos via the allantoic cavity. Four days after incubation, the presence of a virus in the allantoic fluid was determined by a hemagglutination assay (HA), and neutralizing antibody titers were calculated using the Reed and Muench method. 32 The R value was calculated, and Pearson correlation analysis (SPSS software, IBM, Armonk, NY) was used to evaluate the correlation between the R values of the cross-HI and -VN assays.
Results
Virus isolation and identification
Five NDV strains, go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, go/CH/LHLJ/2/06, go/CH/LFJ/1/06, and ck/CH/LHLJ/1/06, were identified in the allantoic fluid from inoculated eggs by HA and HI assays. The range of HA titers of each isolate was 27–28, and their HA activity was inhibited by anti–La Sota chicken serum at HI titers of 27–29 (Table 1). As a reference, the HI titer of the La Sota strain using the La Sota strain as antigen was 210. All viruses were purified and stored at −80°C prior to further use. Avian influenza subtype H5, H7, and H9 viruses were not detected in the allantoic fluids using HI assays and their specific serum or by a real-time PCR assay, which simultaneously detects the H5, H7, and H9 subtypes of avian influenza viruses (data not shown). 27
Genomic characterization and phylogenetic analysis
The complete genomic sequences of the go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, and ck/CH/LHLJ/1/06 strains were 15,192 nt in length; the genomes of the go/CH/LFJ/1/06 and go/CH/LHLJ/2/06 strains were 15,186 nt. The genomic organization of all 5 strains followed the “rule of six,” and the gene order was 3′-NP-P-M-F-HN-L-5′. All strains had the same length of leader and trailer sequences, which consisted of 55 and 114 nt, respectively. Detailed information regarding the genomic characteristics of these isolates and the representative strains ZJ1 (genotype VIId) and La Sota (genotype II) is summarized in Supplemental Table 2.
The go/CH/LHLJ/1/06 strain, which was isolated from a goose, and the go/CH/HLJ/LL01/08 and ck/CH/LHLJ/1/06 strains, which were isolated from chickens in this study, were all classified as genotype VIId of class II, along with 7 other strains from geese and 16 strains from chickens. The go/CH/LHLJ/2/06 and go/CH/LFJ/1/06 strains were classified as genotype II of class II, along with other NDV strains that were isolated from both geese and chickens (Fig.1 and Fig.2).
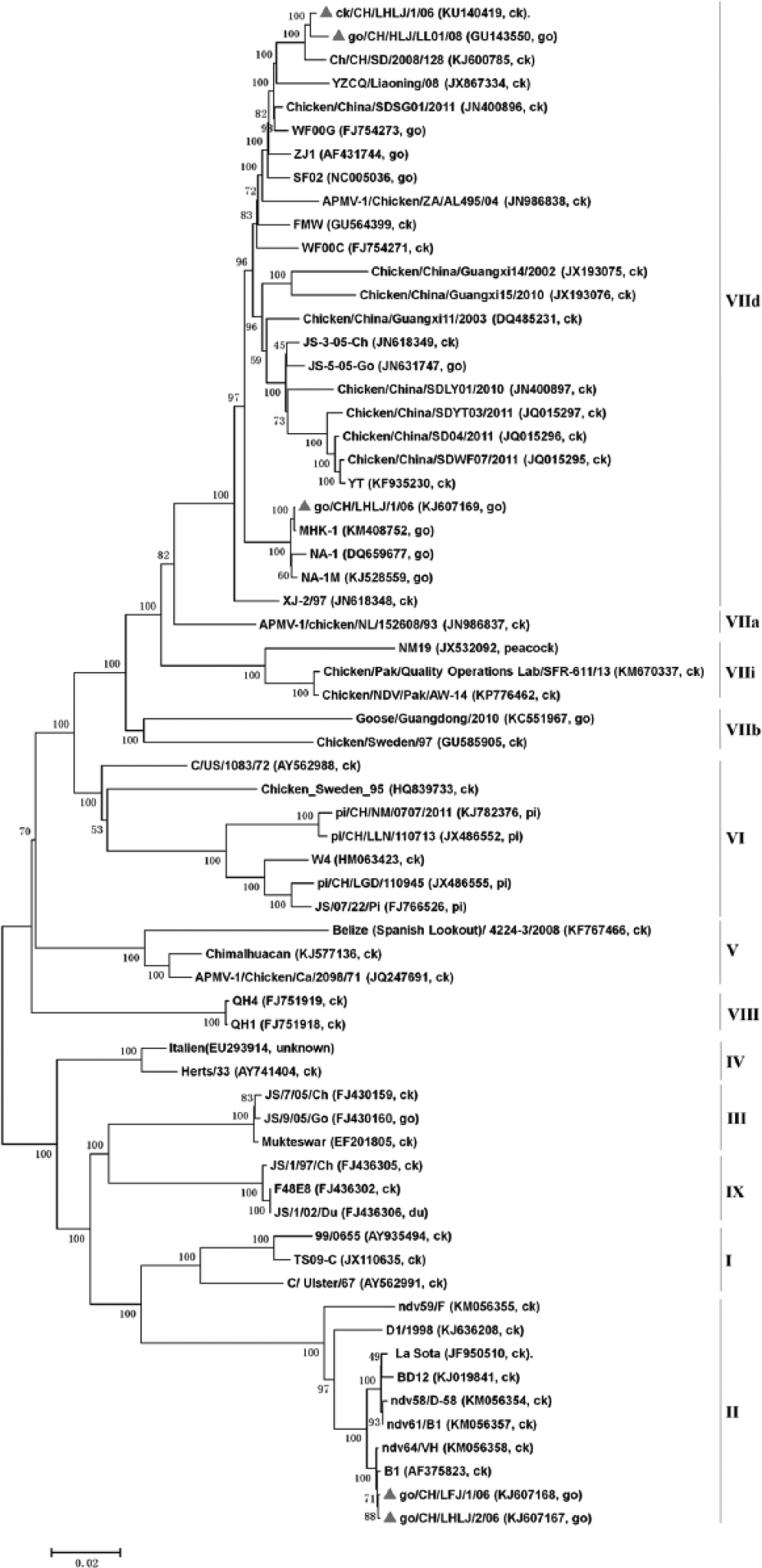

Phylogenetic tree constructed based on the complete genomic sequences of 65 Newcastle disease virus (NDV) strains. The 5 strains in our study are marked with red triangles. GenBank accessions and hosts of the NDV isolates are shown in brackets. The genotype of each strain is indicated at the right. The tree was constructed using the neighbor-joining algorithm of MEGA6.0, and 1,000 bootstrap replicates were performed to assign confidences to the groupings. ck = chicken; go = goose; du = duck; pi = pigeon.
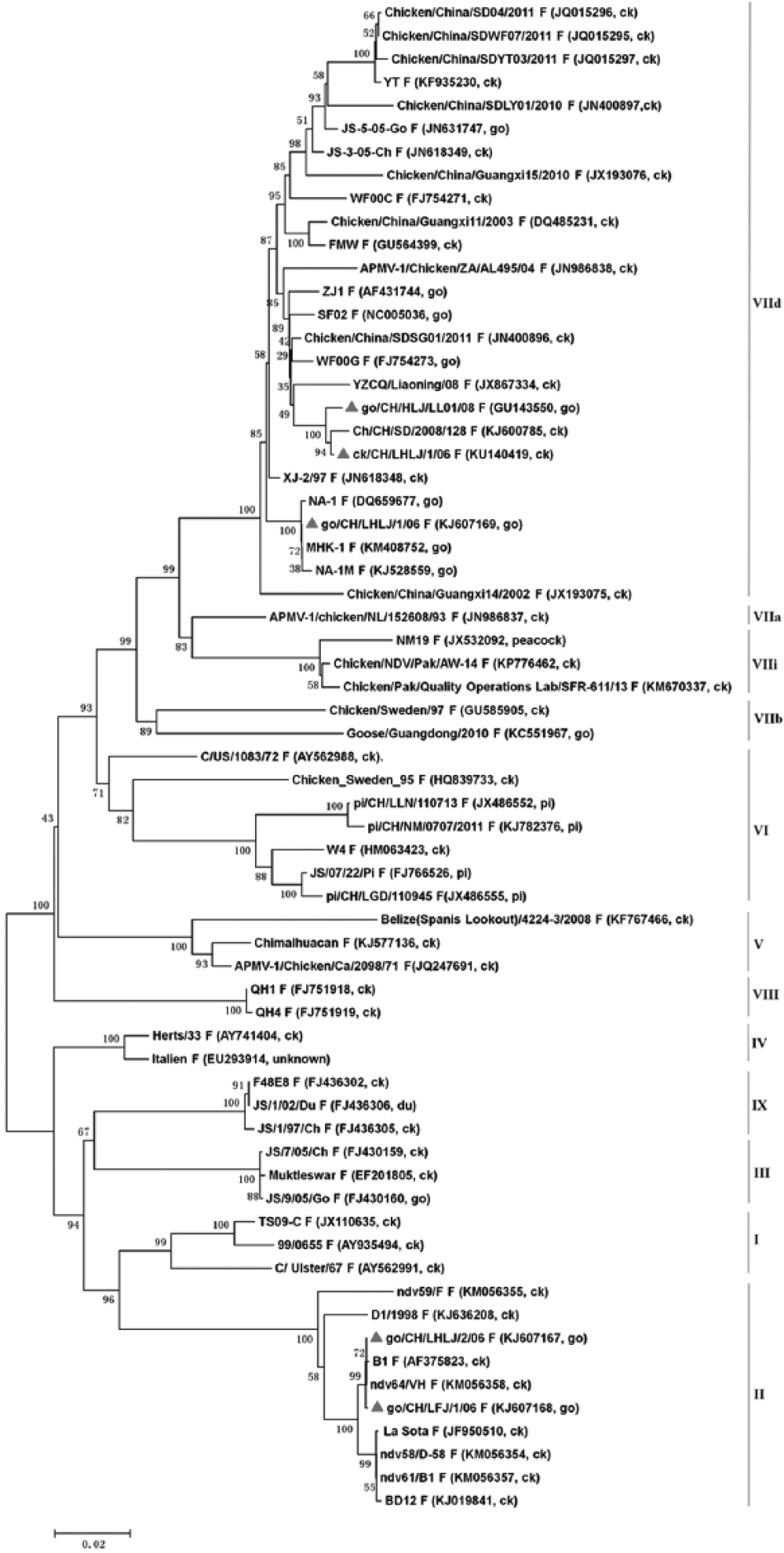
Phylogenetic tree constructed based on the F gene of 65 Newcastle disease virus (NDV) strains. The 5 strains in our study are marked with red triangles. GenBank accessions and hosts of the NDV isolates are shown in brackets. The genotype of each strain is indicated at the right. The tree was constructed using the neighbor-joining algorithm of MEGA6.0, and 1,000 bootstrap replicates were performed to assign confidences to the groupings. ck = chicken; go = goose; du = duck; pi = pigeon.
A genomic sequence comparison showed that the nucleotide identity between chicken and goose genotype II NDVs was 95.5–99.9%; the identities within the chicken and goose genotype II NDVs were 95.5–99.9% and 99.9%, respectively (Table 2). Similarly, the nucleotide identities among chicken and goose genotype VIId NDVs were 94.2–99.9%; the identities within the chicken and goose genotype VIId NDVs were 94.2–99.8% and 96.0–99.9%, respectively (Table 2). In addition, the NP, P, M, F, HN, and L genes also shared high homologies among the genotype II and VIId NDVs that were isolated from geese and chickens, respectively (Table 2).
Homology comparisons of chicken- and goose-origin genotype II and VIId Newcastle disease virus (NDV) strains.
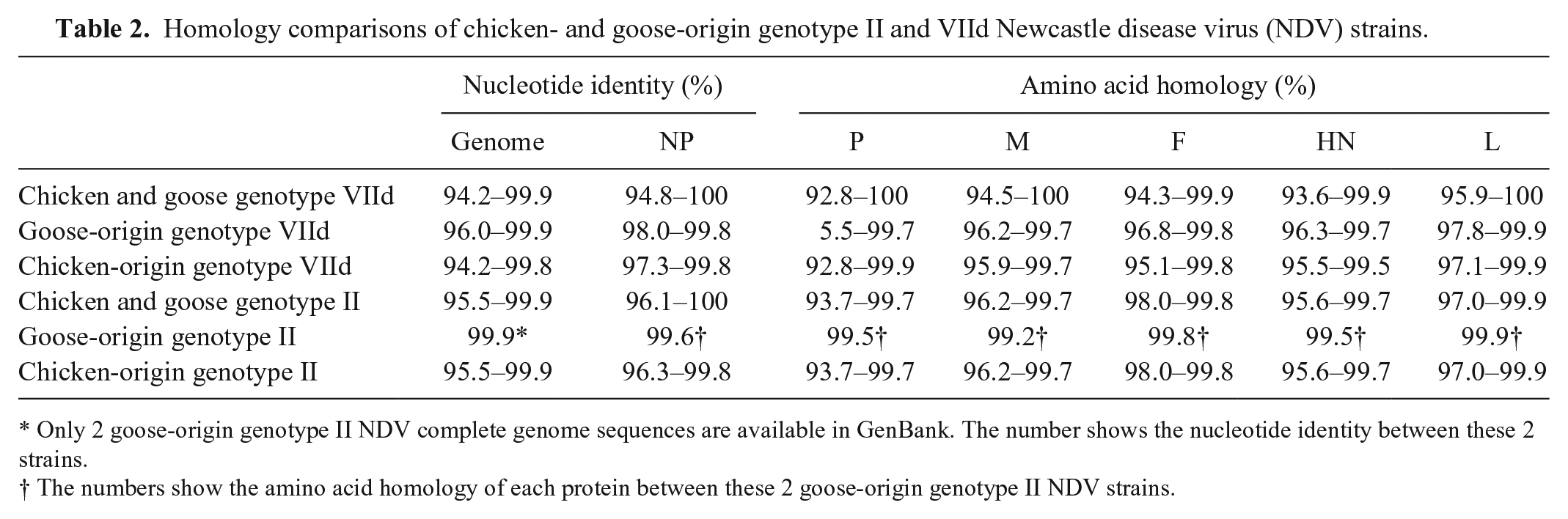
Only 2 goose-origin genotype II NDV complete genome sequences are available in GenBank. The number shows the nucleotide identity between these 2 strains.
The numbers show the amino acid homology of each protein between these 2 goose-origin genotype II NDV strains.
The F protein cleavage site motifs of the go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, and ck/CH/LHLJ/1/06 strains are 112RRQKRF117; this motif in the go/CH/LHLJ/2/06, go/CH/LFJ/1/06, and La Sota strains is 112GRQGRL117. In our study, the F protein of the go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, go/CH/LFJ/1/06, and La Sota strains contained all 6 conserved potential N-glycosylation sites (residues 85, 191, 366, 447, 471, and 541); however, the go/CH/HLJ/LL01/08 and ck/CH/LHLJ/1/06 strains contained an N-to-A mutation at position 543 that resulted in the loss of the N-glycosylation site at position 541 to 543. Cysteine residues that were located at positions 25, 27, 76, 199, 338, 347, 362, 370, 394, 399, 401, 424, and 523 of the F protein were found in the avirulent go/CH/LHLJ/2/06, go/CH/LFJ/1/06, and La Sota strains; the virulent go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, and ck/CH/LHLJ/1/06 strains showed a C-to-R substitution at residue 27. Of the neutralizing epitopes found at residues 72, 74, 75, 78, 79, 157 to 171, and 343 of the F protein in most NDV strains,28,33,40 1 amino acid substitution at residue 78 (K-to-R) was found in the go/CH/LHLJ/1/06 strain and another was found at residue 72 (D-to-G) in the go/CH/HLJ/LL01/08 strain (Supplemental Table 3).
Of the neutralizing epitopes identified in the HN protein in most NDV strains,5,16,18,19 identical amino acid substitutions were found at positions 263, 494, and 514 in the virulent strains (ck/CH/LHLJ/1/06, go/CH/LHLJ/1/06, and go/CH/HLJ/LL01/08) and avirulent strains (go/CH/LHLJ/2/06, go/CH/LFJ/1/06, and La Sota; Supplemental Table 4). Residue 347 of the HN protein is a major determinant of an immunodominant epitope, and the HN 347K mutation leads to antigenic variation in genotype VII NDV strains.8,16 Significantly, the residue at position 347 of the HN protein of the go/CH/LHLJ/2/06 and go/CH/LFJ/1/06 strains, which were isolated from geese in different provinces of China, is a K (Supplemental Table 4), which indicates that these strains have very similar molecular characteristics and might be antigenic variants.
Virulence studies
The MDT and ICPI of the go/CH/LHLJ/1/06 strain were 51 h and 1.86, respectively, and 51.6 h and 2, respectively, for the ck/CH/LHLJ/1/06 strain, demonstrating that these strains are velogenic in chickens (Table 1). The MDT and ICPI of the go/CH/LHLJ/2/06 strain were 109.4 h and 0.22, respectively, and 98 h and 0.19, respectively, for the La Sota strain, suggesting that they are lentogenic strains.
Pathogenicity in geese
Although apparent clinical signs were not observed in all of the NDV-infected geese, 2 birds died in the groups that were challenged with the go/CH/LHLJ/1/06 or ck/CH/LHLJ/1/06 strains. The geese in the control group did not show any clinical signs, and none of these geese died. Similar gross anatomic lesions were observed in dead geese that were infected with the genotype VIId go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strains. In general, hemorrhage and edema were found in the proventriculus. Hemorrhagic changes in the trachea and congestion in the lung and brain were also observed. Livers of dead geese showed slight enlargement and congestion. In addition, these 2 VIId NDV strains, which originated from different hosts, induced similar histologic changes in the tissues of dead geese. Hyperplasia of colloid cells was found in the brain (Fig. 3A, 3D). The pathogenic changes of the proventriculus (Fig. 3B, 3E) included desquamation of mucosal epithelial cells, atrophy of simple tubular glands in the lamina propria, lamina propria edema with inflammatory cell infiltration, and edema of the muscularis mucosae. In addition, deciliation of the tracheal epithelium was observed (Fig. 3C, 3F). No pathologic change was observed in the corresponding tissues of birds in the control group (Fig. 3G–I).
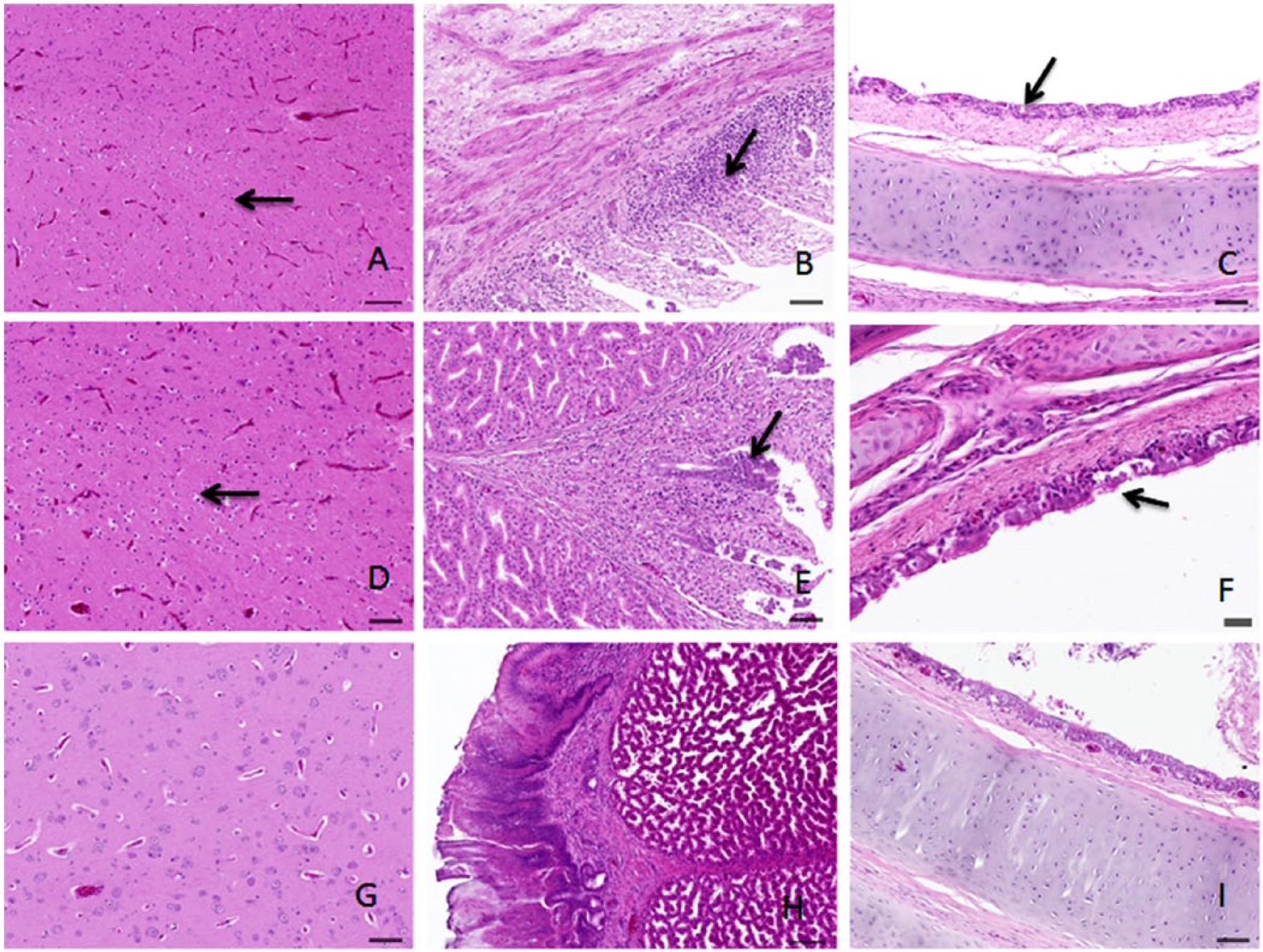
Histologic changes in tissues from dead geese infected with Newcastle disease virus (NDV) strains are shown. The brain tissue samples from geese infected with the go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strains show colloid cell hyperplasia (A and D, respectively). The proventriculus tissue samples from geese infected with the go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strain show desquamation of mucosal epithelial cells, atrophy of simple tubular glands in the lamina propria, lamina propria edema with inflammatory cell infiltration, and edema of the muscularis mucosae (B and E, respectively). The trachea tissue samples from geese infected with the go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strains show deciliation of the epithelium (C and F, respectively). Histologic changes are indicated by arrows. Normal brain, proventriculus, and trachea tissue samples are from a goose in the control group (G–I, respectively).
The go/CH/LHLJ/1/06 viral RNA could be detected in 10 tissue samples in both dead geese at 3 dpi (Table 3). However, viral RNA could only be detected in 6 out of the 10 tissue samples, including the trachea, lung, kidney, liver, proventriculus, and spleen, in the dead geese that were infected with the ck/CH/LHLJ/1/06 strain. In addition, the viral RNA copy numbers in the tissues of dead geese infected with the go/CH/LHLJ/1/06 strain were significantly higher than those in the dead geese that were infected with the ck/CH/LHLJ/1/06 strain (p < 0.05), with the exception of the kidney (Fig. 4).
Detection of viral RNA in tissue samples of dead geese.*
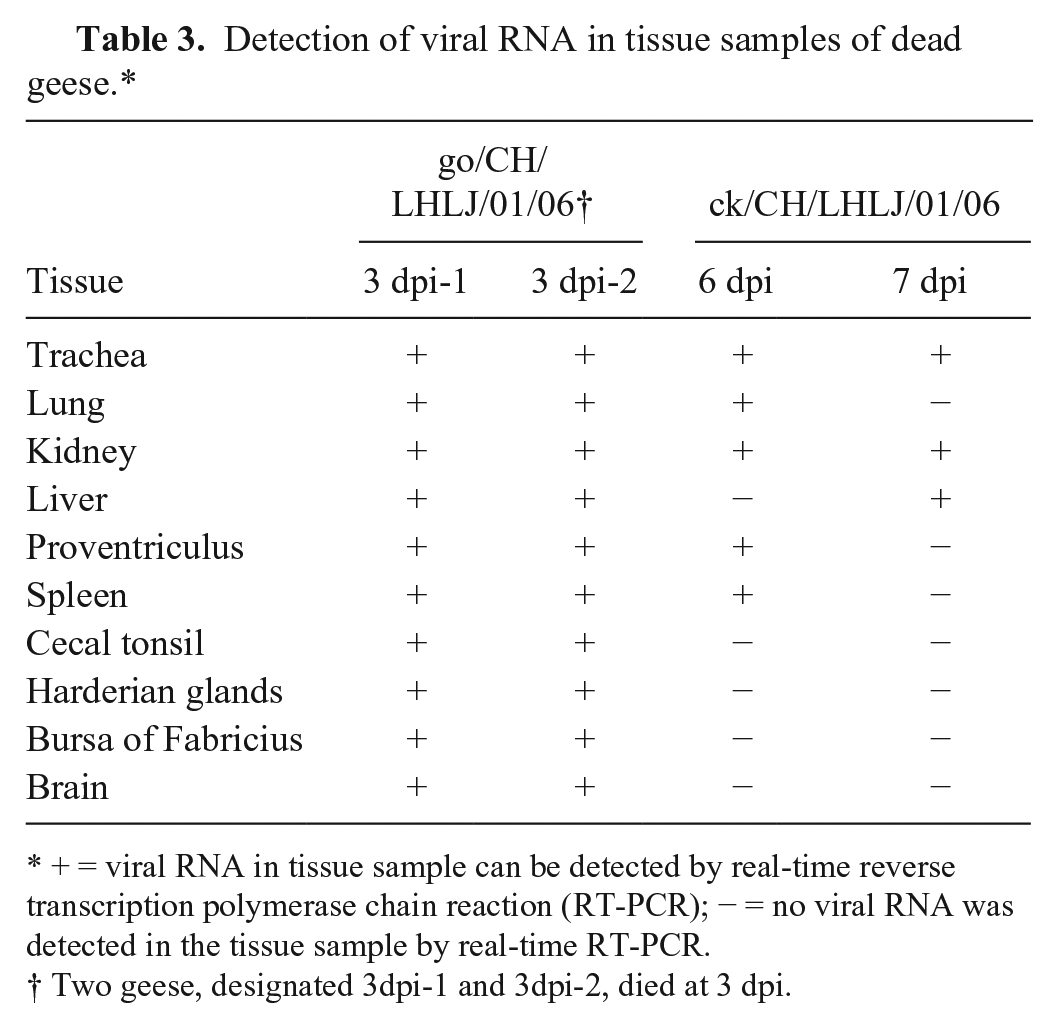
+ = viral RNA in tissue sample can be detected by real-time reverse transcription polymerase chain reaction (RT-PCR); − = no viral RNA was detected in the tissue sample by real-time RT-PCR.
Two geese, designated 3dpi-1 and 3dpi-2, died at 3 dpi.
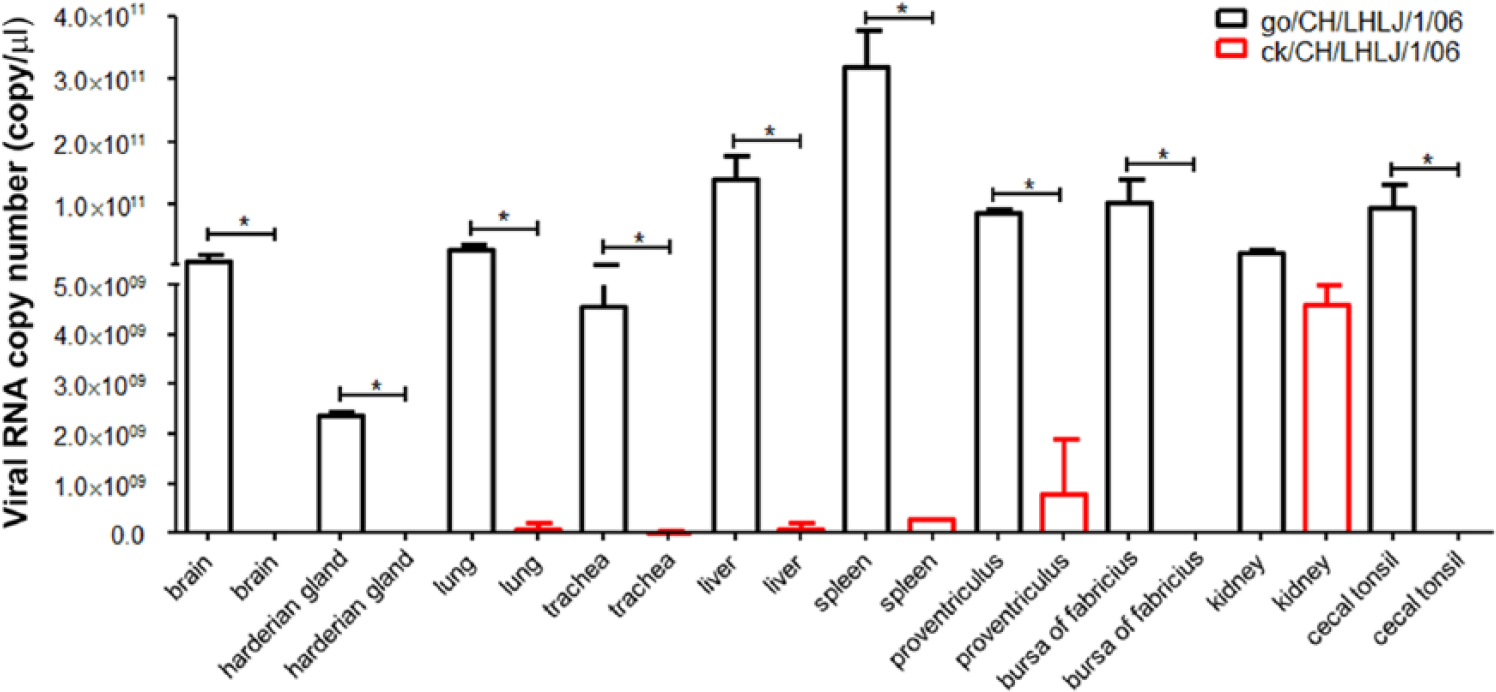
Average viral RNA copy number in tissue samples from dead geese infected with Newcastle disease virus (NDV) strains. The viral RNA in the tissue samples of geese that died at 3 dpi in the go/CH/LHLJ/1/06-infected group, and geese that died at 6 and 7 dpi in the ck/CH/LHLJ/1/06-infected group were detected by real-time RT-PCR. The average viral copy number of each tissue from dead geese infected with the go/CH/LHLJ/1/06 or ck/CH/LHLJ/1/06 strain was calculated using a standard curve based on a 10-fold dilution series of a standard template of known concentration. The results are represented as per-group means. Error bars indicate standard deviations (SD) of the mean. Data were compared using the Student t-test (*p < 0.05).
Virus shedding
Four days after challenge with the go/CH/LHLJ/01/06, go/CH/LHLJ/02/06, and ck/CH/LHLJ/01/06 strains, 25%, 10%, and 30% of the geese were positive for virus recovery in the oropharyngeal swabs, respectively (Table 4); data from the cloacal swabs showed that 50%, 0%, and 10%, respectively, of the geese tested positive for these strains. No virus was isolated from any geese that were inoculated with the La Sota strain or from the unchallenged group. Furthermore, no virus could be detected in any of the swab samples at 8, 12, 16, 20, 24, and 28 dpi.
Virus recovery from geese infected with the go/CH/LHLJ/01/06, go/CH/LHLJ/02/06, ck/CH/LHLJ/01/06, and La Sota strains.*
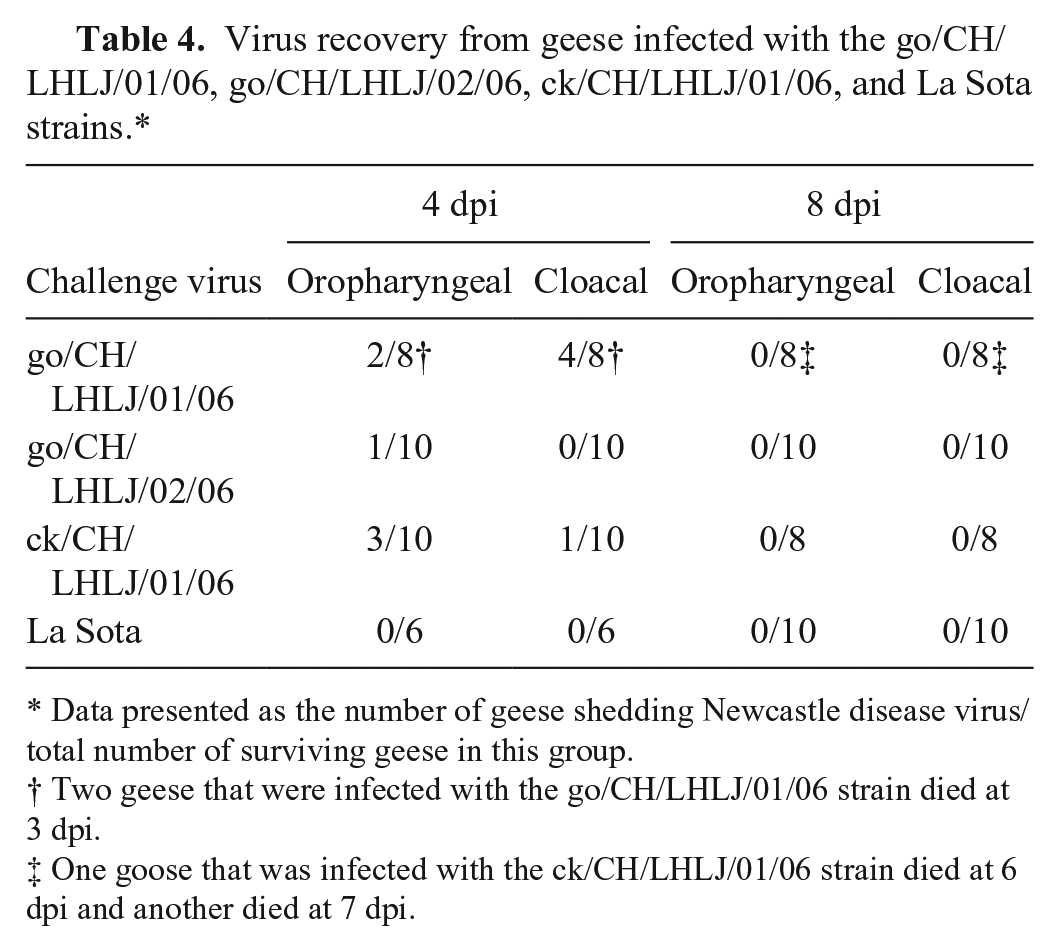
Data presented as the number of geese shedding Newcastle disease virus/total number of surviving geese in this group.
Two geese that were infected with the go/CH/LHLJ/01/06 strain died at 3 dpi.
One goose that was infected with the ck/CH/LHLJ/01/06 strain died at 6 dpi and another died at 7 dpi.
Serologic assay
The cross-HI assay showed that there were no significant antigenic differences between the go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strains (R = 0.707); however, significant antigenic differences were found between the go/CH/LHLJ/1/06 and go/CH/LHLJ/2/06 strains (R = 0.203), and the go/CH/LHLJ/1/06 and La Sota strains (R = 0.5; Table 5). When strain ck/CH/LHLJ/1/06 was compared with the go/CH/LHLJ/2/06 and La Sota strains, R values of 0.225 and 0.25, respectively, were obtained, indicating that there were significant antigenic differences between them. Interestingly, the go/CH/LHLJ/2/06 strain showed a significant (R = 0.319) antigenic difference from the La Sota strain, even though they belonged to the same genotype.
R values for the cross–hemagglutinin inhibition (HI) and cross–virus neutralization (VN) assays among different Newcastle disease virus strains.*
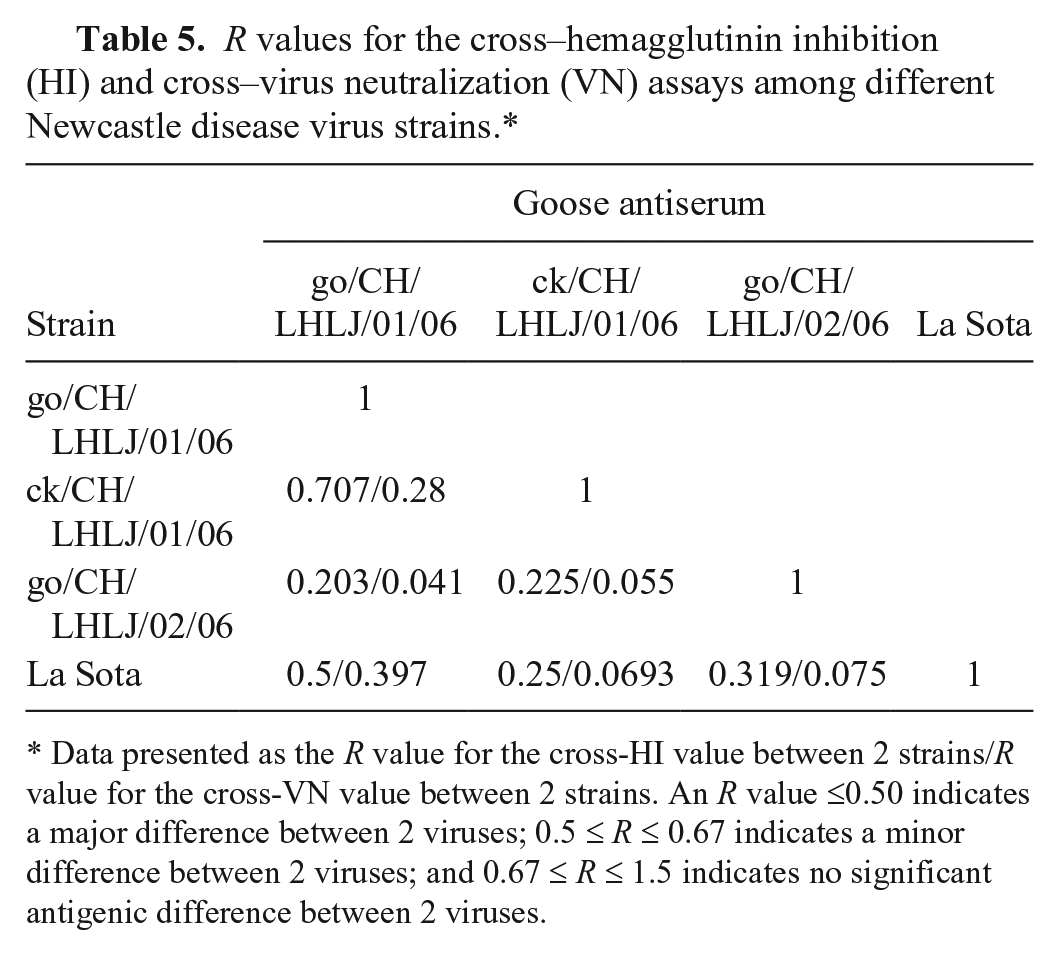
Data presented as the R value for the cross-HI value between 2 strains/R value for the cross-VN value between 2 strains. An R value ≤0.50 indicates a major difference between 2 viruses; 0.5 ≤ R ≤ 0.67 indicates a minor difference between 2 viruses; and 0.67 ≤ R ≤ 1.5 indicates no significant antigenic difference between 2 viruses.
Much more striking antigenic differences among these 4 strains were revealed by the cross-VN assay, compared with those revealed by the cross-HI assay (Table 5). Pairwise comparisons of the go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, ck/CH/LHLJ/1/06, and La Sota strains yielded R values that were all < 0.5, indicating that there are significant antigenic differences among these 4 strains. A relevant analysis of these 2 serologic assays was performed (SPSS software, IBM) because the R values of the cross-VN assay were lower than those of the cross-HI assay. The results showed that the R values were positively correlated (r = 0.823, p = 0.044) between the cross-HI and cross-VN assays.
Discussion
Genomic sequence comparison showed that genotype II and VIId NDVs isolated from geese have molecular characteristics similar to genotype II and VIId NDVs isolated from chickens. Genotype II and VIId NDVs isolated from geese and chickens clustered together, respectively, in phylogenetic trees based on the F gene and the whole genome. Consequently, we conclude that there are no obvious differences in the general genomic characteristics among NDV strains that were isolated from chickens and geese.
Epidemiologic analysis showed that genotype VIId was the predominant genotype of the NDVs that were responsible for Newcastle disease outbreaks in chicken and goose flocks in the past decade in mainland China and Taiwan.17,22–24,31,38,39 Therefore, it can be concluded that genotype VIId NDVs from geese might have been derived from an earlier chicken VIId NDV. Our results also indicate that genotype VIId NDVs are still co-circulating among chicken and goose flocks in China. The remaining 2 strains (go/CH/LHLJ/2/06 and go/CH/LFJ/1/06) in our study were genotype II, sharing nucleotide homologies of 99% and 99.8% with the vaccine strains La Sota and B1, respectively, indicating that they might be re-isolations of these vaccine strains, or that they evolved from these vaccine strains through the chicken-to-goose transmission pathway because these vaccine strains are commonly used in chicken flocks in China.
We further evaluated whether the F protein cleavage site of NDVs that were isolated from geese is consistent with the MDT and ICPI of NDV pathogenicity in chickens.13,30 As shown in Table 1, the genotype VIId go/CH/LHLJ/1/06, go/CH/HLJ/LL01/08, and ck/CH/LHLJ/1/06 strains had the sequence motif 112RRQKRF117, which is a representative velogenic site that is commonly identified in genotype VIId NDVs. The F cleavage motif of the go/CH/LHLJ/2/06 and go/CH/LFJ/1/06 strains is 112GRQGRL117, which is identical to that of the La Sota strain and characteristic of lentogenic NDVs. These results are in agreement with the MDT and ICPI that were measured in embryonated chicken eggs and 1-d-old chickens (Table 1), suggesting that unlike pigeon parmyxovirus-1, the F cleavage site is a significant factor in determining the virulence of NDVs isolated from geese.
We identified several amino acid substitutions in the neutralizing epitopes of the F and HN proteins. Compared with the genotype II La Sota strain, the go/CH/LHLJ/1/06 strain has a K-to-R substitution at residue 78 of the F protein; strain go/CH/HLJ/LL01/08 has a D-to-G substitution at residue 72 of the F protein. Amino acid substitutions at residues 263, 347, 494, and 514 of the HN protein were found in NDV strains used in this study in comparison with La Sota, and the 347K substitution, which can result in antigenic differences, was present in 2 genotype II strains, go/CH/LHLJ/2/06 and go/CH/LFJ/1/06, which were isolated from different provinces in China. To investigate the antigenic relatedness of NDVs that were isolated from geese and chickens, cross-HI and -VN assays were performed in our study. The cross-HI results showed that there were no significant antigenic differences among the genotype VIId strains that were isolated from chickens and geese, whereas significant antigenic differences among genotype VIId and II NDVs were found. Interestingly, the R value for the go/CH/LHLJ/02/06 and La Sota strains was 0.319, indicating there are antigenic differences between these 2 strains that have the same genotype. The antigenic difference between the go/CH/LHLJ/02/06 and La Sota strains may be attributed to the 347K amino acid substitution in the HN protein of the go/CH/LHLJ/02/06 strain.7,8,16 Nevertheless, the R values calculated via the cross-VN assay were all < 0.5, suggesting that there are major antigenic differences among go/CH/LHLJ/1/06, go/CH/LHLJ/2/06, ck/CH/LHLJ/1/06, and La Sota. The antigenic differences demonstrated by the cross-HI and -VN assays may be attributed to the fact that the cross-HI assay is based on the ability of antibodies to recognize antigenic sites on the HN protein, whereas the cross-VN assay is based on the ability of antibodies to recognize neutralizing epitopes on both the F and HN proteins, although the influences of unidentified antigenic sites on the F and/or HN proteins cannot yet be ruled out.
Unsurprisingly, the genotype II NDVs (go/CH/LHLJ/2/06 and La Sota) did not cause clinical signs or death in geese, whereas genotype VIId NDVs (go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06) caused 20% mortality in geese. Similar gross and histologic lesions (proventriculus, trachea, lung, brain, and liver) were observed in dead geese that were infected with the go/CH/LHLJ/1/06 and ck/CH/LHLJ/1/06 strains. The time to death of geese infected with the go/CH/LHLJ/1/06 strain (3 dpi) was earlier than that of geese infected with the ck/CH/LHLJ/1/06 strain (6 and 7 dpi), suggesting that the go/CH/LHLJ/1/06 strain was more pathogenic in geese than the ck/CH/LHLJ/1/06 strain, which might be attributed to the host adaptation of the go/CH/LHLJ/1/06 strain in geese. Alternatively, pathogenicity might be associated with the virus strains because different NDVs showed various degrees of pathogenicity in their hosts, although further investigation is required to address this issue. In addition, virus shedding results showed that geese infected with the go/CH/LHLJ/1/06, ck/CH/LHLJ/1/06, or go/CH/LHLJ/2/06 strain shed virus through the oropharyngeal and cloacal routes, indicating that geese may play important roles in NDV transmission and epidemics.
Footnotes
Authors’ contributions
Q Xu and J Sun contributed equally to this study. Q Xu and J Sun contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. M Gao and S Zhao contributed to conception of the study and contributed to analysis and interpretation of data. H Liu and T Zhang contributed to conception and design of the study. Z Han and X Kong contributed to conception of the study and contributed to interpretation of data. S Liu contributed to conception of the study, contributed to acquisition and interpretation of data, and drafted the manuscript. All authors critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from National “Twelfth Five-Year” Plan for Science & Technology Support (2015BAD12B03), Special Fund for Agro-scientific Research in the Public Interest (201303033), and the China Agriculture Research System (CARS-41-K12).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
