Abstract
Swine and poultry viruses, such as porcine reproductive and respiratory syndrome virus (PRRSV), porcine epidemic diarrhea virus (PEDV), and highly pathogenic avian influenza virus (HPAIV), are economically important pathogens that can spread via aerosols. The reliability of methods for quantifying particle-associated viruses as well as the size distribution of aerosolized particles bearing these viruses under field conditions are not well documented. We compared the performance of 2 size-differentiating air samplers in disease outbreaks that occurred in swine and poultry facilities. Both air samplers allowed quantification of particles by size, and measured concentrations of PRRSV, PEDV, and HPAIV stratified by particle size both within and outside swine and poultry facilities. All 3 viruses were detectable in association with aerosolized particles. Proportions of positive sampling events were 69% for PEDV, 61% for HPAIV, and 8% for PRRSV. The highest virus concentrations were found with PEDV, followed by HPAIV and PRRSV. Both air collectors performed equally for the detection of total virus concentration. For all 3 viruses, higher numbers of RNA copies were associated with larger particles; however, a bimodal distribution of particles was observed in the case of PEDV and HPAIV.
Introduction
Infectious agents that can be transmitted via air include some of the most economically important pathogens affecting intensively reared animals. These include foot-and-mouth disease virus (FMDV), porcine reproductive and respiratory syndrome virus (PRRSV), porcine epidemic diarrhea virus (PEDV), and influenza A virus (IAV) in pigs, and Newcastle disease virus, Marek’s disease virus, and avian influenza virus (AIV) in poultry. These pathogens cause significant economic losses as a result of their pathogenicity and ease of transmission, including the ability to travel long distances2,9,18,21,23 and, in some instances, also present a zoonotic risk. 9
Risk of airborne transmission is most problematic in areas of high farm and animal density where regional spread can be difficult to contain. In such scenarios, biocontainment and biosecurity to limit herd-to-herd (or flock-to-flock) transmission is a major challenge. As a result, livestock producers are investing significant resources to prevent airborne transmission of infectious agents such as PRRSV and AIV.1,5
One of the outstanding questions about airborne transmission concerns the composition and size of particles with which airborne viruses are associated. The size of particles in air influences the distances to which the particles (with their associated viruses) are transported, the survivability and infectivity of the viruses in aerosols, and the locations at which they are deposited in the respiratory tract after inhalation. There is limited information on particle size association of pathogens with aerosols, and much existing literature has been generated in humans or with animals under experimental conditions that simulate natural infection processes rather than from natural infections in field settings.3,12,13,16 Furthermore, such information has not provided insights into the particle size association of viruses transported between herds, in part because of the limitations of air sampling methods. Sampling of bioaerosols can be performed by impaction (onto plates, membranes, or growth media), impingement into liquid, or capture by filtration. In addition, most air samplers have been tested under experimental conditions using flow rates of 2–22 L/min (Lpm), which are more suitable for conditions of low air flow and high pathogen load. More recently, a cyclonic air collector (Midwest Microtek, Brookings, SD) has been used in field studies to assess airborne dissemination of swine viruses (i.e., PRRSV and IAV).10,21 However, this apparatus does not separate particles by size.
Mathematical modeling of airborne viruses provides a complementary tool to interpret and predict the risk of airborne transmission events. However, current models do not incorporate information on particle size. For example, atmospheric dispersion models for FMDV are based on data of disease onset, virus strain, number and type of animals involved, and stage of the disease.15,17,20,23,27 Because infectivity is linked to particle size,3,28 characterization of pathogen-associated particle size in aerosols is important to understand and to predict the likelihood of transmission of infective viruses among farms. Furthermore, the efficiency of air sanitation methods such as air filtration and electrostatic ionization technology are particle size dependent.1,4 Therefore, knowledge about sizes of particles harboring viruses should help identify the most cost-effective biocontainment and biosecurity measures to prevent the dissemination of infectious viruses within and among farms.
We assessed and compared the efficiency of 2 air samplers to detect 3 different viruses (PRRSV, PEDV, and highly pathogenic avian influenza virus [HPAIV]) during active outbreaks that occurred on swine and poultry farms. We describe the size distribution and concentrations of these viruses in indoor and outdoor air. Despite the heterogeneous conditions of these measurements and possible confounding factors, we believe that the results of our study show the strengths and weaknesses of using both methods in field conditions and provide important insights to determine the ability of airborne transmission of these viruses.
Materials and methods
All procedures were conducted in accordance with the University of Minnesota Institutional Biosafety Committee (IBC) and Institutional Animal Care and Use Committee (IACUC). Permission to sample the air at the HPAI-affected flocks was obtained via U.S. Department of Agriculture–Animal and Plant Health Inspection Service (USDA-APHIS) veterinarians as part of the regulatory response to the HPAI outbreak in the U.S. Midwest in 2015 (https://goo.gl/xb5wmX).
Farm selection/description
The study was conducted between March and May of 2015 in 9 swine or poultry farms. Veterinarians serving swine herds in southern Minnesota were contacted and asked to alert the investigators when acute outbreaks of PRRSV or PEDV were suspected on client farms. Four nursery farms were included in the study (2 for PRRSV, and 2 for PEDV) and diagnoses were confirmed as described below. Recruitment of poultry flocks was achieved via the USDA-APHIS veterinarians engaged in the emergency response to the H5N2 HPAIV outbreak in the U.S. Midwest 2015 outbreaks. Three turkey flocks in Minnesota and 1 layer flock each in Iowa and Nebraska were selected. The selected flocks had a confirmed diagnosis of HPAIV at the time of sampling, and air sampling was conducted within 2–11 days of the diagnostic confirmation.
Sampling procedures
To measure concentration and particle size distribution of viruses inside and outside the affected facilities, air samples were collected using 2 air samplers: the Andersen cascade impactor (ACI; Thermo Electron, Waltham, MA), 6 and the Tisch cascade impactor (TCI; Model 230, Tisch Environmental, Village of Cleves, OH). The TCI is a high-volume air sampler, and both the ACI and the TCI are size-differentiating air samplers that distribute captured particles into several size categories.
Air samples were first collected using the TCI air sampler immediately outside affected facilities (~5 m from the exhaust fan). The ACI sampler was also used to collect air samples outside poultry facilities at ~5 m from the exhaust fans. Subsequently, both air samplers were dissembled, cleaned, and disinfected as explained below, and brought inside. Air samplers inside the facilities were located in strategic locations chosen based on secure and stable areas close to the animals (within a 1-m range) but out of their reach to avoid any direct contact. Relative humidity (%) and temperature (°C) were recorded (Kestrel 3000, Nielsen-Kellerman, Broothwyn, PA) during each sampling event to document environmental conditions.
The ACI sampled air at 28.3 Lpm for 1 h, and separated particles into 8 size intervals based on their aerodynamic diameter: 0.4–0.7, >0.7–1.1, >1.1–2.1, >2.1–3.3, >3.3–4.7, >4.7–5.8, >5.8–9.0, and >9.0 µm. Particles smaller than 0.4 µm were captured by a backup filter. Samples from different stages (aluminum plates) of this device were eluted individually using a cell scraper and 1 mL of minimal essential medium. 6 All samples were transferred into 1.5 mL sterile plastic tubes, placed on ice, and stored at −80°C until tested. The ACI was then disassembled, and plates and stages were scrubbed with alkyl dimethyl benzyl ammonium chloride soap (Lysol, Reckitt Benckiser, Slough, England), rinsed, and finally disinfected with 70% ethanol prior to drying with paper towels. After disinfection, a randomly selected collection of plates and individual ACI stages were swabbed to serve as negative controls, and samples stored at −80°C.
The TCI sampler sampled air at 1,130 Lpm for 30 min, separating particles into 4 stages, with aerodynamic particle diameter cut points of 0.95, 1.5, 3, and 7.2 μm. Particles were captured using slotted glass fiber collection substrate (Model P/N TE-230-GF, New Star Environmental, Roswell, GA), and particles smaller than 0.95 µm were captured by a backup filter. After each sampling period, the substrates and filters were removed using sterile gloves, folded, placed into individual labeled Petri dishes, and refrigerated at 4°C prior to sample elution and processing, which took place within 24 h of collection. Between collection events, all stages were disinfected with 70% alcohol, air dried, and finally rinsed and dried with paper towels. After disinfection and drying, all stages were swabbed, and samples stored at −80°C. To extract the captured viruses from substrates and filters, samples were removed from the Petri dishes, cut individually in 1 cm2 squares using sterile scissors, placed in 50 mL conical centrifuge tubes, and eluted with 20 mL of an eluent solution (3% beef extract–0.05 M glycine solution at pH 9.1). As part of the elution process, each tube was inverted 10 times to mix the contents, followed by a 5-min set period; another mixing step of 10 inversions; and finally, each tube was vortexed for 5 min at 572 × g. Eluates from filters were measured, transferred to a 15 mL conical centrifuge tube, and the pH adjusted to ~7.0. All samples were transferred into 1.5 mL sterile plastic tubes and stored at −80°C until testing.
For each sampling event, there were 8 stages from the ACI, 4 stages from the TCI, and 2 backup filter samples. At each sampling location, 2 replicates were performed with the ACI sampler and 3 with the TCI sampler.
Total airborne particle counts by particle size were determined to describe the airborne particle distribution in the different farm environments (inside and outside) using an optical particle counter (AeroTrak 9306 handheld particle sizer, TSI, St. Paul, MN). Given its ability to separate particles from 0.3 to >10 µm into 6 size intervals, total airborne particles were counted during each sampling event, and 2 sampling repetitions or “replicates” of 30 min were performed at the respective locations.
Laboratory analysis
Air samples collected in swine facilities were tested for PRRSV and PEDV by quantitative reverse transcription polymerase chain reaction (qRT-PCR) in order to estimate the number of RNA viral copies per m3 of air using previously validated methods.2,8,24 In addition, samples were also assessed semiquantitatively where threshold cycle (Ct) values <35 were considered positive, 35–40 suspect, and >40 negative. Air samples from poultry facilities were screened using a RT-PCR based on the matrix gene for influenza viruses at the University of Minnesota Veterinary Diagnostic Laboratory (St. Paul, Minnesota). 25 Influenza-positive samples were re-tested using specific H5 and N2 RT-PCRs. Positive and suspect samples were also tested using a quantitative IAV RT-PCR (qRT-PCR) as published previously. 10
To confirm that pigs were indeed infected with PRRSV or PEDV, oral fluids were collected from pig pens at the same time as air sampling by hanging 0.4 m of cotton rope from pen railings following published procedures.11,22 Samples were then refrigerated, transported to the laboratory, and stored at −80°C.
Statistical analyses
Data from the qRT-PCR results, replicate, virus, location, type of air collector, and particle size were entered in a spreadsheet (Excel, Microsoft, Redmond, WA) and organized for analysis. First, we conducted a descriptive exploratory analysis of the results comparing the proportions of PCR-positive, -suspect, and -negative results of both air samples at the different farms. Results were analyzed by a 2-tailed Fisher exact test. Afterward, an analysis of variance with 2 fixed effects and 1 random effect was used to predict virus concentration in the air with different collectors at different particle size ranges.
For the quantitative analysis, negative stages of both air samplers were included in the analysis when 1 or more stages of the sampler had a Ct value within the positive or suspect range (Ct <40). Negative results included in the analysis were assigned a value of 100 RNA copies/mL according to the limit of detection of the qRT-PCR technique. In order to compare the efficiency of both air samplers collecting different particle size ranges, a new variable called “size” was created. If the geometric mean point of a specific size interval (square root of the product of the 2 ends of the interval) was <1 μm, ≥1 and ≤3 μm, or >3 μm, then “size” was equal to “small,” “medium,” or “large,” respectively.
Total quantity of virus (RNA copies/m3) was assessed for significance using a generalized linear mixed model (SAS Institute, Cary, NC). Three models were performed for each of the specific size intervals (small, medium, and large). The variables, replicate, and farm were considered as random. Air sampler (ACI vs. TCI), virus (PRRSV, HPAIV, and PEDV), and location of sampling (inside vs. outside) were treated as fixed effects. The model was fitted by the SAS Glimmix model using maximum likelihood estimation. The total quantity of virus (RNA copies/m3) was tested for interaction of air sampler by location and by virus. P values by the model of the multiple comparisons were calculated according to the Tukey–Kramer method.
Results
Sample description, total viral load, and air sampler comparison
The 9 farms included in our study housed an average of 5,500 pigs (range: 4,700–6,800) with a maximum of 4 rooms per barn, and an average of 500,000 birds (range: 4,205–1.8 M) with a maximum of 15 barns.
A total of 444 qRT-PCR samples from 68 air sampling events were tested. Table 1 summarizes the results from the sampling events by air sampler, type of virus, and location of sampling. The ACI sampler detected a higher percentage of positive sampling events compared with the TCI (23 of 26 [88%] vs. 24 of 42 [57%]). Sixty-nine percent (11 of 16), 61% (17 of 28), and 8% (2 of 24) were positive sampling events (Ct values <35) for PEDV, HPAIV, and PRRSV, respectively. Inside samples were more often positive (23 of 44, 52%) than air samples collected outside the facilities (7 of 24, 29%).
Number of positive and suspect air sampling events divided by the total samples tested by reverse transcription polymerase chain reaction (RT-PCR) for porcine reproductive respiratory syndrome virus (PRRSV), porcine epidemic diarrhea virus (PEDV), and highly pathogenic avian influenza virus (HPAIV) collected by the Andersen cascade impactor (ACI) and high-volume Tisch cascade impactor (TCI) from inside and outside (5 m) infected swine and poultry facilities.*
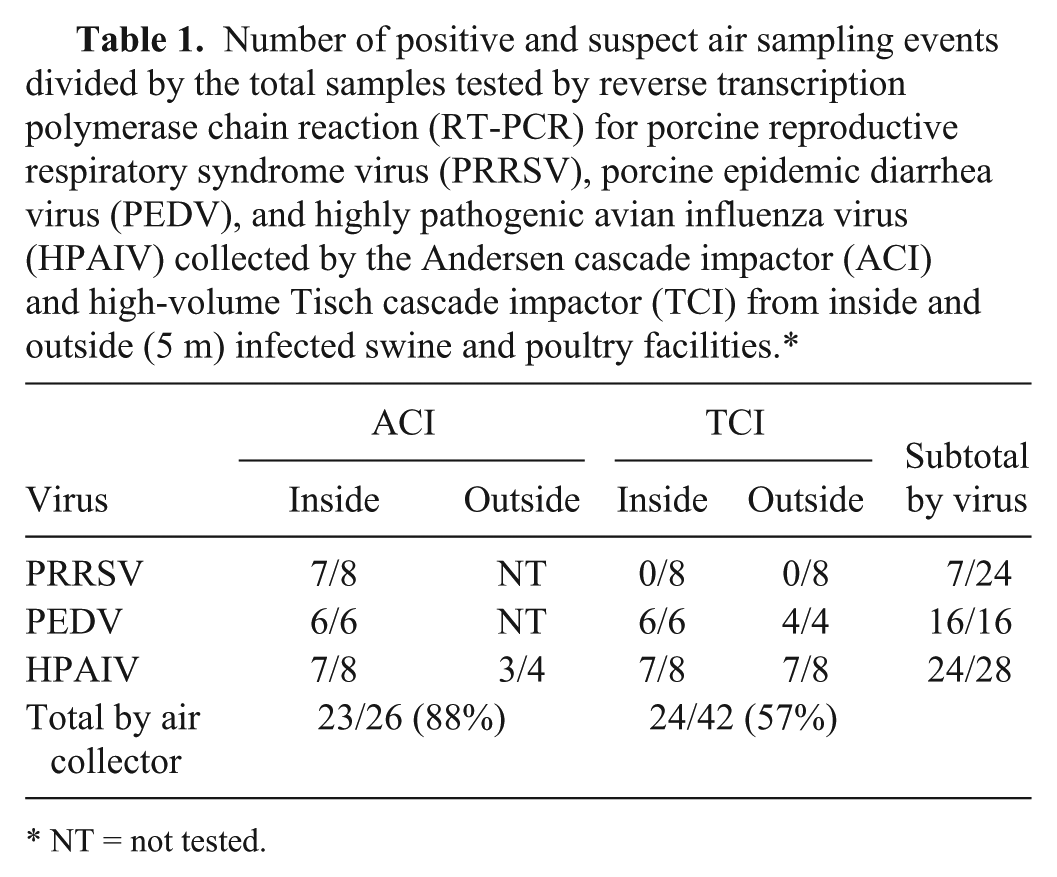
NT = not tested.
The results of the sampling events collected for PRRSV inside swine facilities with the ACI sampler differed significantly from those collected with the TCI sampler (p = 0.001). No other statistically significant differences were observed between collectors.
Overall, we found no difference in virus concentration between the ACI sampler (5.09 ± 1.22 log10 RNA copies/m3 of air) and the TCI air sampler (4.47 ± 1.35 log10 RNA copies/m3 of air; Fig. 1A). There was also no difference in virus concentration detected by the different collectors between animal species as shown in Table 2 for positive and suspect samples of HPAIV.
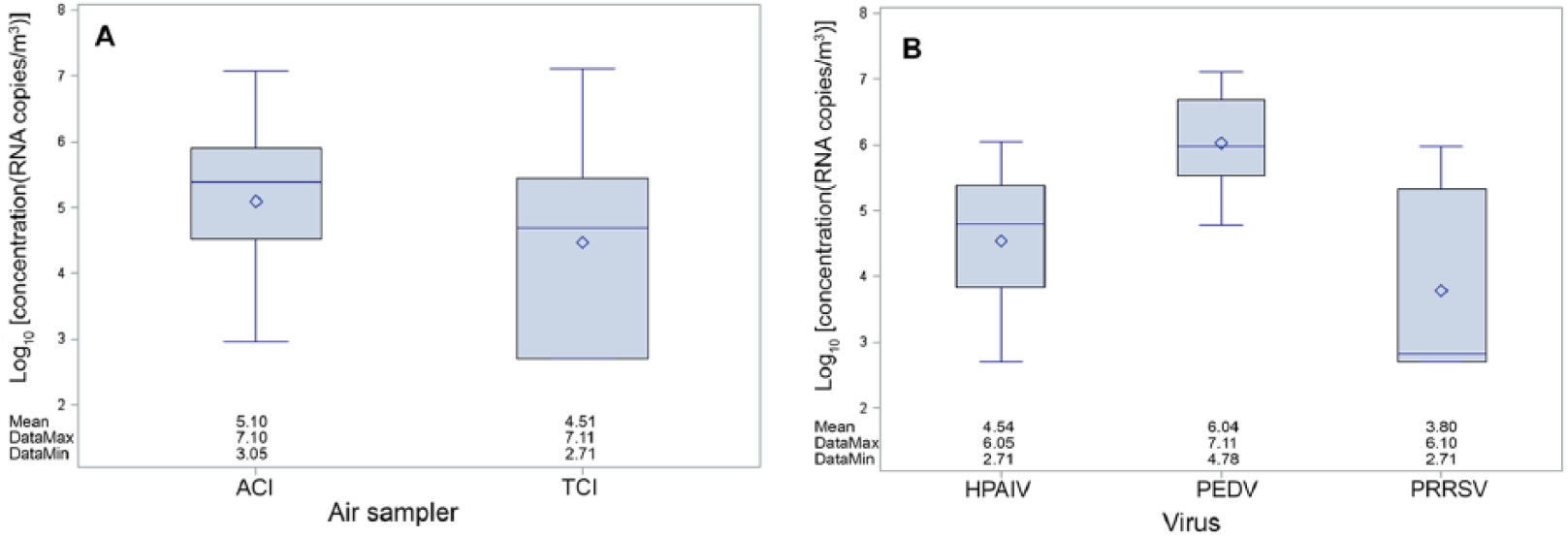
Box plots of virus concentration by virus type (
Quantity (geometric mean of RNA copies/m3 of air [geometric standard deviation] or GM [GSD]) of H5N2 highly pathogenic avian influenza virus in suspect and positive sampling events collected inside and outside using 2 size-selective air samplers: an Andersen cascade impactor (ACI) and a high-volume Tisch cascade impactor (TCI), in turkey and layer premises.*

N = total sampling events; n = positive or suspect sampling events (Ct < 40); NT = not tested.
The highest airborne virus concentrations were found for PEDV (6.04 ± 0.72 log10 RNA copies/m3 of air), followed by HPAIV (4.53 ± 0.97 log10 RNA copies/m3 of air), and then PRRSV (3.79 ± 1.35 log10 RNA copies/m3 of air; Fig. 1B).
Particle size distribution by virus type, sampling location, and air sampler type
All 3 viruses were detected inside barns with the ACI sampler, and 2 of 3 (PEDV and HPAIV) were detected inside and immediately outside by the TCI sampler. All viruses were associated with a wide range of particle sizes. Higher concentrations of viral RNA copies were detected in association with large particles, and these associations were different among the 3 viruses tested (Fig. 2). In the case of PRRSV, there were higher concentrations of viral RNA in particles >5.8 μm compared with those <5.8 μm (p ≤ 0.03). For PEDV, there were higher concentrations of viral RNA in particles >3.3 μm diameter compared with those of 0.4–1.1 μm (p ≤ 0.02). For HPAIV, a significantly higher viral RNA concentration was associated with the largest particles (>9 μm) compared with the smallest particles (<0.7 μm; p ≤ 0.017). Viral RNA was detected in the smallest size range (particles <0.4 μm) only for PEDV and HPAIV. No PRRSV RNA copies were detected in the smallest size ranges (<2.1 μm), and values were imputed as the limit of qPCR detection values (Fig. 2).
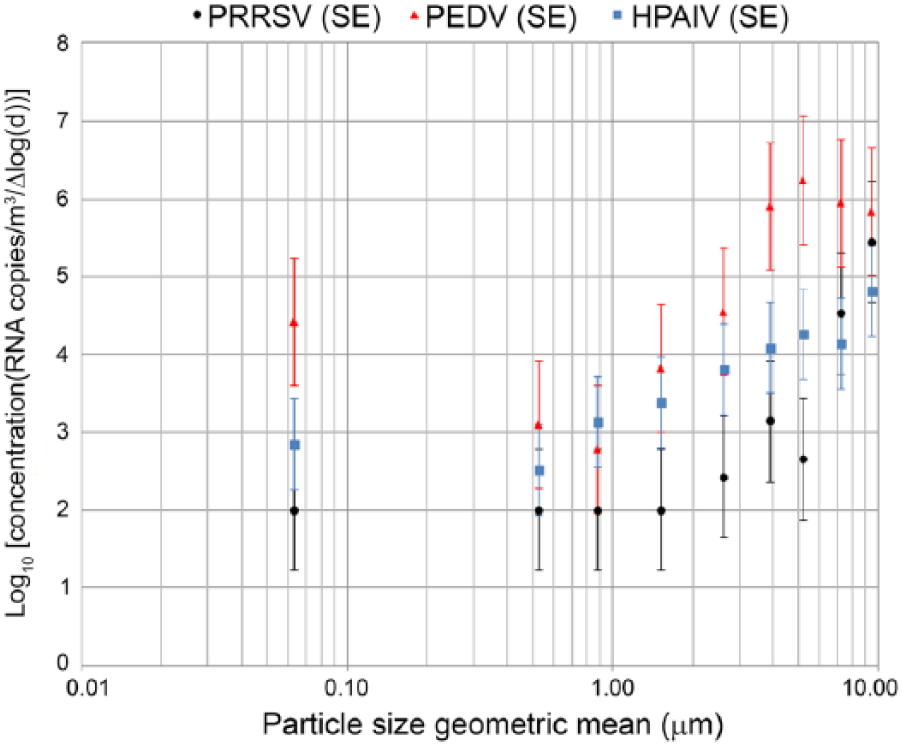
Distribution of viral RNA concentration (least squares means of log10 RNA copies/m3 of air and standard error [SE]) of porcine reproductive and respiratory syndrome virus (PRRSV), porcine epidemic diarrhea virus (PEDV), and highly pathogenic avian influenza virus (HPAIV) by particle size as detected by the Andersen cascade impactorb from aerosols generated by infected animals inside infected premises.
The particle size distribution analysis by air sampler, type of virus, and location is summarized in Figure 3. Variable “replicate” was removed from the final model because lack of significance and total concentration of viral particles was obtained as an average of all replicates. Three models were created, one for each particle size range (“small,” “medium,” and “large”). For PRRSV, in small and medium particle sizes, there were only low quantities of RNA (2.5–2.6 log10 RNA copies/m3) detected with the ACI, and no sample was positive with the TCI sampler; however, for the purposes of the model, the minimum detectable concentration value was imputed. For HPAIV, the ACI sampler yielded significantly higher concentrations of viral RNA associated with small and medium-sized particles inside (4.2 vs. 4.4 log10 RNA copies/m3) compared to the TCI sampler (2.2 and 3.5 log10 RNA copies/m3). For large particle sizes (>3 µm) collected inside, the ACI sampler yielded significantly higher concentrations of HPAIV and PRRSV (5 and 4.8 log10 RNA copies/m3, respectively) than the TCI sampler (3.8 and 2.3 log10 RNA copies of HPAIV and PRRSV, respectively). No difference between samplers was observed with regard to concentration of PEDV RNA in samples collected inside associated with any of the 3 particle sizes measured.
Distribution of viral RNA concentration (least squares means of log10 RNA copies/m3 of air and standard error) of porcine reproductive and respiratory syndrome virus (PRRSV), porcine epidemic diarrhea virus (PEDV), and highly pathogenic avian influenza virus (HPAIV) by particle size categories (small, medium, and large) detected by the Andersen cascade impactor (ACI) and the Tisch cascade impactor (TCI) from inside and outside (5 m from exhaust fan) infected swine and poultry facilities.
A significantly lower concentration of RNA copies associated with particles of 1–3 µm was observed outside compared with inside and measured with the TCI sampler. HPAIV was the only virus detected with both air samplers in outside air samples, but we observed no significant differences between air collectors. All samples collected outside swine facilities with the TCI sampler were negative for PRRSV RNA.
Total airborne particles
The distribution of total particles in the air measured with the optical particle counter at the time of virus sampling inside (Fig. 4A) and outside (Fig. 4B) of the barns was different. There was higher concentration of total particles within the first submicrometer size (0.3–0.5 µm) compared with particles >0.5 µm (p < 0.001) for both locations; however, this difference was more evident in samples collected outside.
Distribution of total airborne particles (number of total particles/m3/∆og [diameter] and 95% confidence interval) inside (
Discussion
The cost-effective implementation of biosecurity interventions at the farm level to prevent herd-to-herd transmission of pathogens remains a significant challenge for farmers raising food animals. A particular challenge is presented by pathogens with attributes that enable airborne transmission, which facilitates rapid spread between farms and frustrates the efforts to contain transmission within a region. To meet this challenge, there is a need to obtain a better understanding of the aerobiology of pathogen transmission under field conditions and to develop better methods to study these phenomena. We found that the observed concentrations of viral RNA, and the observed associations between viruses and particles, were influenced by the choice of measurement device. Overall, both devices performed similarly, but the ACI sampler yielded higher RNA copies of PRRSV and HPAIV compared with the TCI sampler. Using 2 size-differentiating air samplers, we demonstrated that these important infectious agents can be associated with a wide range of particle sizes under field conditions and perhaps become spread by different modes from infected premises given that they could be detected in size ranges associated with both airborne and droplet transmission modes. Despite the information provided here on particle size, further studies are needed to investigate the component composition of particles of different size classes.
PEDV, primarily an enteric virus, was found in larger quantities in air samples than HPAIV (primarily a systemic infection in birds but with an important enteric component) and PRRSV (primarily a respiratory infection with a systemic component). Because different farms were tested for different viruses, there is a need to highlight possible differences between premises. The stage of clinical progression of outbreaks, the percentage of actively infected animals at the time of sampling, the difference in mechanisms of spread among the 3 viruses, and environmental conditions inside and outside of barns would be expected to influence the variability in positive yields observed between farms and viruses.
Determining the particle size associations for these viruses has important implications for the control of the diseases, and the use of droplet and airborne infection control measures. Although results from the ACI sampler from inside the facilities demonstrated the presence of virus across all size ranges studied, higher quantities were associated with larger sized particles (Fig. 2). Similar results on virus particle size distribution have been shown for swine influenza virus, PRRSV, and PEDV under experimental conditions. 3 Furthermore, our results showed that PEDV and HPAIV RNA copies are associated with the smallest particles (<0.4 μm) demonstrating a possible bimodal distribution of viral copies that could suggest potential for transmission via multiple mechanisms or under varying environmental conditions. Smaller particles tend to remain airborne for longer periods of time as compared to larger particles. For example, a 4 µm particle takes 33 min to settle 1 m in still air, whereas a 1 µm particle requires 8 h. 19 In addition, infectious particles with a small aerodynamic diameter tend to have more serious health implications than larger particles because they are inhaled into the lower respiratory tract. 26 We did not assess the viruses’ viability based on particle size, therefore the question of virus viability based on particle size found under field conditions still needs to be determined. Overall, our results indicate that PEDV, PRRSV, and HPAIV RNA can be associated with particles that are small enough to remain airborne for a long time and, in the cases of HPAIV and PEDV, disperse inside as well as outside the infected premises. Although a lack of PRRSV detection in smaller particles may simply reflect less overall shedding of PRRSV into the environment as compared to PEDV or HPAIV, our PRRSV data also suggest that long-distance airborne transmission is less likely, which is in contrast to previously published studies. 7 In addition, given the potential zoonotic risk of some of HPAI viruses, our results underline the importance of using proper personal protection equipment when working in or near affected flocks.
Differences in the air sampling process methods between the samplers could have an influence on sampling efficiency. For the ACI sampler, airborne particles were collected directly onto aluminum substrates placed on the impaction plates, whereas filter substrates were used with the TCI sampler and, as a result, airborne particles were captured in the filter medium when the airstream passed through the filter. However, the elution and separation of the captured virus from the crustal material is a critical step in pathogen detection. 14 Lower efficiency of the virus elution process from filter media, effects of environmental factors, and desiccation of the virus particles caused by the airstream passing around the particles are variables that warrant further investigation before considering the use of a filter-based large-volume air sampler under field conditions.
Despite a growing body of evidence, the relative con-tribution of the airborne route to the overall transmission of these viruses among farms remains controversial. Understanding the capabilities of available air samplers that can characterize aerosols of such important pathogens by size, specifically under field conditions, will fill a knowledge gap about airborne transmission that should greatly enhance understanding of this phenomenon.
Footnotes
Acknowledgements
We thank all the veterinarians and producers who participated in the study. The help of Dr. Robert Porter and Devi Patnayak at the University of Minnesota Veterinary Diagnostic Laboratory, the USDA/APHIS staff, in particular Dr. Brian McCluskey and Mia Torchetti, poultry industry veterinarians, and poultry companies in accessing the HPAI-affected flocks and information on the outbreaks is gratefully acknowledged. Special thanks to Martin Alonso and Mari Carmen Garcia-Mochales for their assistance on logistics; Marta Ferrandis, Cesar Prado, and the University of Minnesota Public Health residents for their assistance during the field sampling; and My Yang for her guidance during the laboratory procedures of this project.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This material was made possible, in part, by a Cooperative Agreement from the United States Department of Agriculture’s Animal and Plant Health Inspection Services (APHIS) and funding from the United States Department of Agriculture National Institute of Food and Agriculture (MIN-62-058; Agricultural Experimental Station).
