Abstract
In 2015, 6.7–60% mortality was observed in black seabream (Acanthopagrus schlegelii), rock bream (Oplegnathus fasciatus), and gray mullet (Mugil cephalus) farmed in the southern coast of Korea. On examination, numerous amoebae were found on the gills of these 3 fish species with detection rate of 100%. Some rock bream and gray mullet were coinfected with bacteria (Pseudomonas anguilliseptica, Vibrio tapetis, or Vibrio anguillarum). Histologic examination revealed extensive hyperplastic epithelium and lamellar fusion in the gills. Numerous amoebae were seen between gill filaments. The amoebae collected from the 3 fish species had specific 630 bp of a partial 18S rRNA gene fragment for Neoparamoeba perurans. Phylogenetic analysis based on partial 18S rRNA gene nucleotide sequences revealed that these Korean amoeba isolates belonged to the N. perurans group. Based on our results, black seabream, rock bream, and gray mullet were added as new hosts for N. perurans.
Amoebic gill disease (AGD) is a serious parasitic disease that affects marine cultured salmonids including Atlantic salmon (Salmo salar), coho salmon (Oncorhynchus kisutch), and rainbow trout (Oncorhynchus mykiss) in Australia, the United States, and several European countries.7–10 AGD also affects other fish species cultured in marine environments, including turbot (Psetta maxima; syn. Scophthalmus maximus), European seabass (Dicentrarchus labrax), sharpsnout seabream (Diplodus puntazzo), ballan wrasse (Labrus bergylta), and ayu (Plecoglossus altivelis) in Spain, South Africa, the Mediterranean, Norway, and Japan.1 –3,7 –9 Thus, AGD threatens the marine farming industry in many countries.
The clinical signs of AGD include multiple white-to-gray patches on swollen gills, with excess mucus surrounding gill arches. 7 The main histologic feature of AGD is prominent epithelial hyperplasia in the gills, resulting in complete lamellar fusion.7,10 Neoparamoeba perurans, Neoparamoeba pemaquidensis, and Neoparamoeba branchiphila have been isolated from or detected in marine fish with AGD.7,9 Of these amoebae, N. perurans is known as a cosmopolitan etiologic agent of AGD. 14
AGD had not been reported in marine fish in far east Asia until 1994. In 1995, amoebic infection was reported in cultured olive flounder (syn. Japanese flounder; Paralichthys olivaceus) in Korea. 4 Since then, AGD has been found in cultured ayu in 2008 in Japan 1 and coho salmon in 2014 in Korea. 6 We report herein amoebic infection in the following 3 new marine fish species that are cultured in Korea: black seabream (syn. black porgy; Acanthopagrus schlegelii), rock bream (syn. barred knifejaw; Oplegnathus fasciatus), and gray mullet (syn. flathead mullet; Mugil cephalus).
In 2015, 6.7–60% mortality was observed in black seabream, rock bream, and mullet farmed on the southern coast of Korea. Black seabream and rock bream were cultured in a farm (land-based system) located in Goheung County. Gray mullet were cultured in a farm (net cage system) located in Hadong County. A total of 21 diseased fish (10 black seabream, weight: 119 ± 29 g; 7 rock bream, 62 ± 23 g; and 4 mullet, 128 ± 23 g) were collected and individually subjected to parasitologic, bacteriologic, and virologic examinations. Gills and body surface were microscopically examined for parasites. Kidney, spleen, and liver were cultured on brain–heart infusion agar a plates supplemented with 1% NaCl and incubated at 15°C for 14 d. Kidney and spleen samples were homogenized in 10 volumes of Hanks balanced salt solution b and centrifuged at 3,000 × g for 20 min. The supernatant was filtered through a 450-nm membrane filter. One-hundred microliters of the filtrate was inoculated onto Chinook salmon embryo (CHSE-214) and fathead minnow (FHM) cells in 24-well tissue culture plates. c After inoculation, cells were incubated at 15°C for 14 d to examine cytopathic effects (CPE). For histologic studies, gills were removed from diseased fish and immediately fixed in 10% neutral buffered formalin. After fixation, standard histologic procedures were followed for tissue dehydration and paraffin embedding. Tissue sections were stained with hematoxylin and eosin. Bacteria and parasites (amoebae) from diseased fish were subjected to polymerase chain reaction (PCR) analysis. Bacterial and amoebic genomic (g)DNAs were isolated using a gDNA extraction kit d following the manufacturer’s instructions. DNA templates from bacteria were amplified using universal primers (63F: 5′-CAGGCCTAACACATGCAAGTC-3′; 1406R: 5′-ACGGGCGGTGTGTRC-3′) to amplify a 1,343-bp region of the 16S rRNA gene on a thermocycler. e PCR parameters were an initial denaturation at 94°C for 5 min, 35 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s, extension at 72°C for 1.5 min, and a final extension at 72°C for 5 min. The amplified DNA fragments were directly sequenced using a DNA analyzer. f All sequences were submitted for similarity searches with BLAST. PCR amplifications for amoebae were performed using primers F (5′-ATCTTGACYGGTTCTTTCGRGA-3′) and R (5′-ATAGGTCTGCTTATCACTYATTCT-3′) to amplify a 636-bp region of N. perurans 18S rRNA gene. 13 PCR cycle conditions were 94°C for 3 min, 35 cycles of 94°C for 30 s, 50°C for 30 s, and 72°C for 45 s, and 72°C for 10 min. PCR products were purified and cloned into pCR 2.1 TOPO (topoisomerase I) vector g and transformed into Escherichia coli strain TOP10. g Sequencing was performed using M13 reverse and T7 promoter primers. The resulting sequences were assembled with the MEGA6 program. 11 Sequences at 588–590 bp without primer sequences were submitted to GenBank (accessions KU985056–KU985058) and compared with those of 49 worldwide amoebae. 6 Multiple alignment was conducted using Clustal X 12 to infer genetic relationships among these sequences with neighbor-joining algorithm. The final phylogenetic tree was drawn with MEGA6.
Black seabream and rock bream were maintained in square tanks (6 × 6 × 1 m) with partial recirculation using coastal seawater (salinity: 33%) at the Goheung farm. Water temperature was kept at 13–14°C. Fish deaths continued for up to 3 mo, ultimately reaching a cumulative mortality of ~60% (6,000 of 10,000 black seabream) and 10% (600 of 6,000 rock bream). All tested black seabream exhibited no external or internal signs of the disease except gray gills (4 of 10 fish) or pale patches on the gills (6 of 10 fish; Fig. 1A). Most rock bream (6 of 7 fish) showed pale patches on the gills and enlarged spleens (Fig. 1C). Three of 7 fish showed hemorrhage of skin and opercula. The gills of black seabream and rock bream were infected with numerous amoebae with a detection rate of 100% (Fig. 1B, 1D). No other parasites except amoebae were found on the gills or on the body surface of diseased fish. No viruses were isolated from the kidney–spleen tissue homogenates from diseased fish. Bacteria (>100 of the same colonies) were isolated from 2 of 5 rock bream tested, but not from black seabream. The 16S rRNA gene sequences of the bacteria isolated from rock bream shared 100% sequence identities with the 16S rRNA of Pseudomonas anguilliseptica (GenBank accession X99541) known to cause disease in rock bream cultured in Korea. 5 Histologic examination of 2 fish revealed extensive hyperplastic epithelium and lamellar fusion in the gills, with many amoebae between gill filaments (Fig. 2A, 2B). The clinical and histologic signs of diseased fish were similar to those of AGD in other species reported previously.7,10 In PCR assays, all samples (3 fish) showed specific N. perurans 636-bp fragments. When these PCR products of N. perurans–positive samples were sequenced, nucleotide sequences of these isolates from black seabream (BSBGoH15) and rock bream (RBGoH15) showed 98.3% sequence identities to each other. The nucleotide sequences of a black seabream isolate shared 99.3% (584 of 588 bp) sequence identities with the 18S rRNA sequence of N. perurans (GenBank EF216900) from Atlantic salmon in Australia. The nucleotide sequences of the rock bream isolate shared 99.2% (585 of 590 bp) sequence identities with the 18S rRNA sequence of N. perurans (EF474478) from Atlantic salmon in Norway (data not shown). A phylogenetic tree based on the 18S rRNA sequences of 52 worldwide amoebae revealed that the Korean amoeba (BSBGoH15 and RBGoH15 isolates) belonged to the N. perurans group instead of the N. eilhardi, N. pemaquidensis, N. aestuarina, or N. branchiphila group (Fig. 3). These results confirmed severe N. perurans infection in diseased black seabream and rock bream, although some rock bream were also coinfected with P. anguilliseptica.
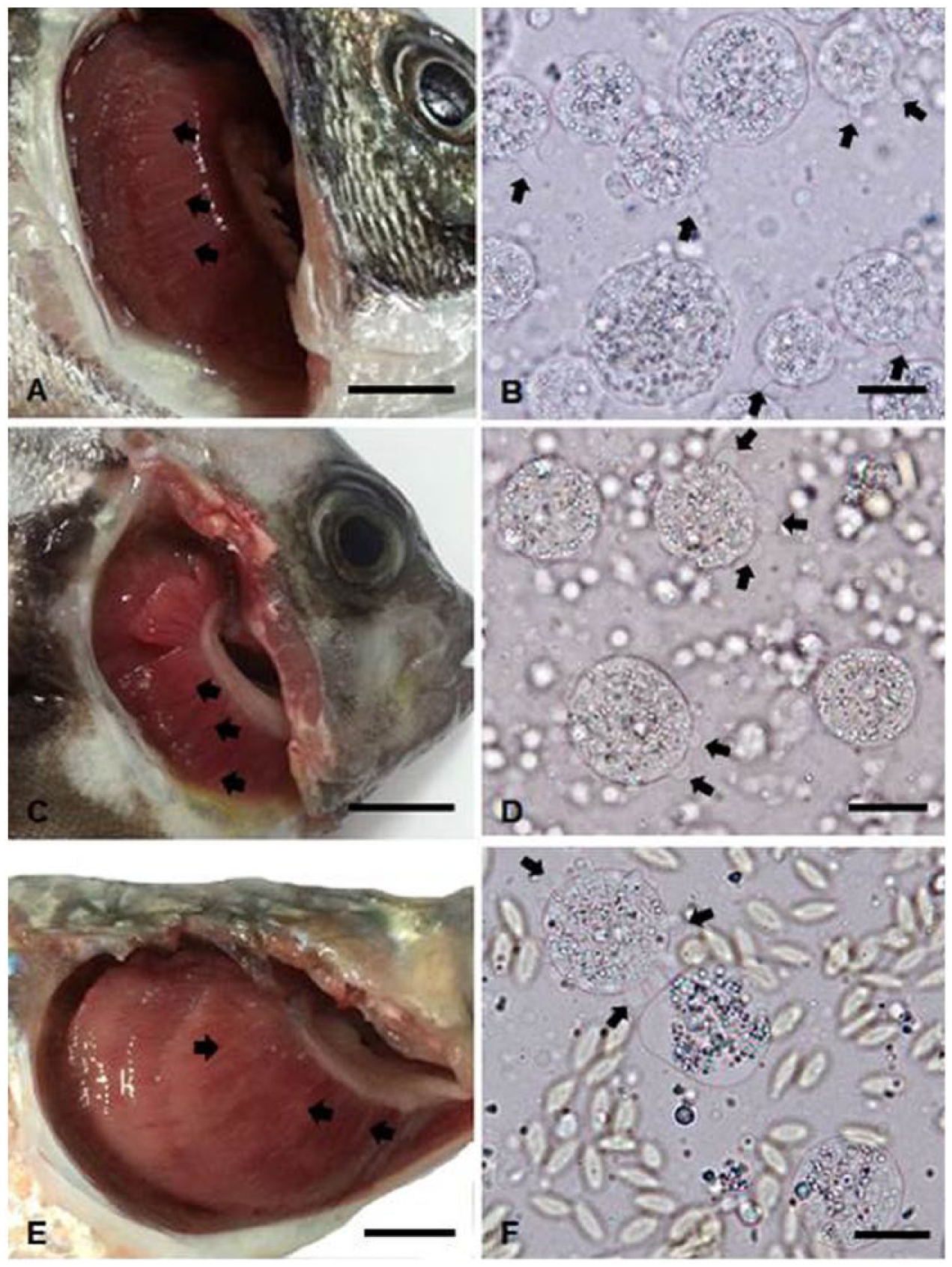
Amoebae infections in cultured black seabream (Acanthopagrus schlegelii;
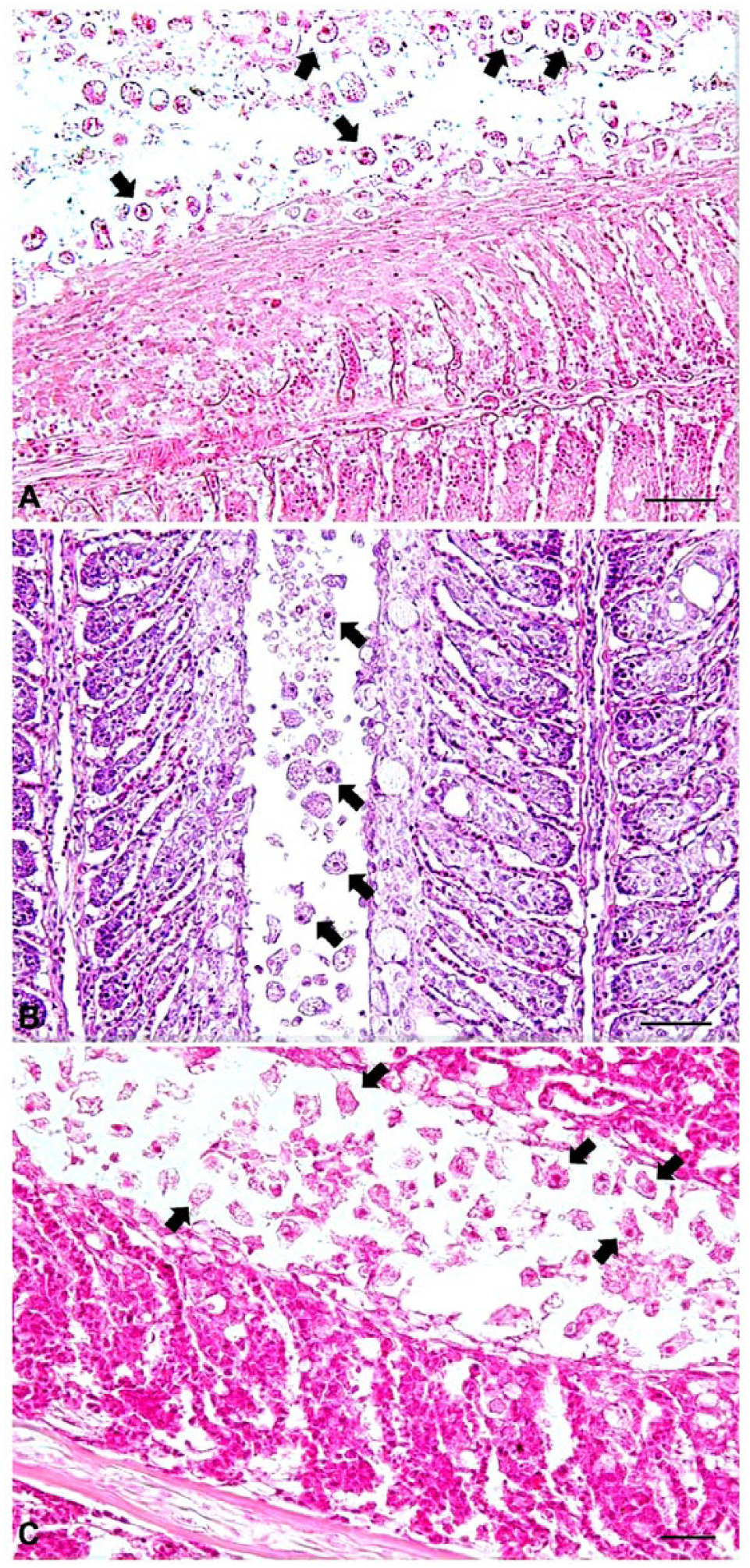
Histopathology of the gills of black seabream (Acanthopagrus schlegelii;
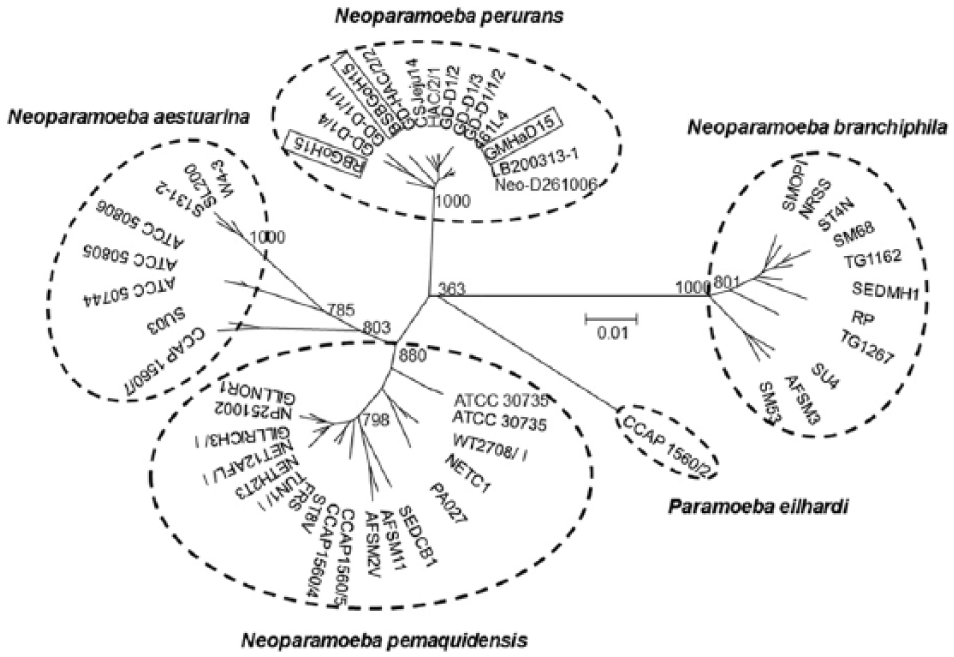
Molecular phylogenetic tree showing the genetic relationships of partial 18S rRNA gene nucleotide sequences among 52 worldwide amoebae. Bootstrap values for 1,000 replicates are shown at major tree nodes. The distance marker refers to the expected number of substitutions per site. GenBank accessions for amoebae isolates are as follows: Neoparamoeba perurans, GD-D1/1/2 (EF216903), GD-D1/3 (EF216900), GD-D1/2 (EF216899), GD-HAC/2/1 (EF216904), CSJeju14 (KU985055), BSBGoH15 (KU985056), RBGoH15 (KU985057), GMHaD15 (KU985058), 461L4 (EF474478), Neo-D261006 (EU326494), LB200313-1 (KF179520), GD-HAC/2/2 (EF216905), GD-D1/1/1 (EF216902), and GD-D1/4 (EF216901); Neoparamoeba branchiphila, AMOPI (EF675600), NRSS (AY714367), ST4N (AY714365), SM68 (AY193725), TG1162 (EF675601), SEDMH1 (AY714366), RP (EF675603), TG1267 (EF675602), SU4 (EF675599), AFSM3 (AY193724), and SM53 (AY193726); Paramoeba eilhardi, CCAP1560/2 (AY686575); Neoparamoeba pemaquidensis, GILLNOR1 (AY714352), NP251002 (AY714351), GILLRICH3/I (EF675606), NET12AFL/I (EF675604), NETH2T3 (AY714350), TUN1/I (EF675607), FRS (AY714356), ST8V (AY714355), CCAP1560/4 (AF371969), CCAP1560/5 (AF371970), AFSM2V (AY193722), AFSM11 (AY193723), SEDCB1 (AY714357), PA027 (AY714358), NETC1 (AY714363), WT2708/I (EF675605), ATCC30735 (AF371972), and ATCC30735 (AY183887); and Neoparamoeba aestuarina, SU03 (EU331035), CCAP1560/7 (AY686574), W4-3 (DQ229957), SL200 (DQ229959), S131-2 (DQ229958), ATCC50806 (AY121852), ATCC50805 (AY121851), and ATCC50744 (AY121848).
Mullet were maintained in net cages (12 × 12 × 8 m) at the Hadong farm. Mortality was first noticed in March 2015 and continued for 2 mo (water temperature: 11–17°C), ultimately reaching a cumulative mortality of ~6.7% (6 of 90 tonnes). Four examined fish showed pale patches on the gills, skin ulcers, and hemorrhage of the skin, fins, opercula, and fat (Fig. 1E). Two of 4 fish had enlarged spleens. No viruses were isolated from the kidney–spleen tissue homogenates. Numerous amoebae were found on the gills with a detection rate of 100% (Fig. 1F). Bacteria were isolated from skin ulcers (3 of 4 fish) and kidney or spleen (1 of 4 fish). The 16S rRNA gene sequences of these bacteria isolated from skin ulcers and kidney–spleen shared 100% sequence identities with the 16S rRNA gene sequence of Vibrio tapetis (LN555640) and Vibrio anguillarum (KC884650), respectively. Histologic examination (2 fish) revealed extensive fusion of secondary lamellae with epithelium hyperplasia (Fig. 2C), consistent with the lesions in diseased black seabream and rock bream. The 18S rRNA gene sequences of amoebae isolated from mullet (GMHaD15) shared 99.3–98.8% sequence identities with the 18S rRNA gene sequences of Korean N. perurans isolates (BSBGoH15, RBGoH15, and CSJeju14) and 99.8% (587 of 588 bp) sequence identities with the 18S rRNA gene sequences of N. perurans isolated from ballan wrasse in Norway (KF179520; data not shown). A phylogenetic tree based on 18S rRNA gene sequences of 52 worldwide amoebae revealed that the Korean amoeba (GMHaD15 isolate) belonged to the N. perurans group instead of the N. eilhardi, N. pemaquidensis, N. aestuarina, or N. branchiphila group (Fig. 3). These results confirmed severe N. perurans infection in diseased mullet, although some fish were coinfected with Vibrio sp.
Footnotes
Authors’ contributions
W-S Kim contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. K-H Kong contributed to acquisition and analysis of data and drafted the manuscript. J-O Kim contributed to acquisition and analysis of data. S-J Jung and J-H Kim critically revised the manuscript. M-J Oh contributed to conception and design of the study; critically revised the manuscript; and gave final approval. W-S Kim, S-J Jung, J-H Kim, and M-J Oh agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
BD Biosciences, Sparks, MD.
b.
Gibco, New York, NY.
c.
Nunc, Uden, The Netherlands.
d.
Bioneer, Daejeon, Korea.
e.
MyGenie 96 thermal block, Bioneer, Daejeon, Korea.
f.
ABI PRISM 3730 XL, Applied Biosystems, Foster City, CA.
g.
Invitrogen, Carlsbad, CA.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was a part of the project titled “Studies on the occurrence status of infectious disease in rainbow trout mariculture and its reduction plans” funded by the Ministry of Oceans and Fisheries, Korea.
