Abstract
Because many bird species are monomorphic or only sexually dimorphic in adult stages, it is difficult to determine their sexes, which may cause significant problems in population and conservation studies. DNA-based sexing relies on the chromodomain helicase DNA binding (CHD) gene located on the W chromosome and its homolog on the Z chromosome, giving distinct banding patterns on agarose gel as a result of length differences in intronic regions within this gene. We used 3 specific primer sets, CHD1F/CHD1R, 2550F/2718R, and P2/P8, for sex determination of 230 samples from 77 avian species. We report here the records for 70 of those species analyzed using the CHD1F/CHD1R primer set, and 49 species using 2550F/2718R, and 46 species using P2/P8. CHD1F/CHD1R PCR products on agarose gel generally showed an apparent single band in males and 2 bands in females, but the products of 2550F/2718R (61%) and P2/P8 (42%) showed distinct banding patterns for separate bird orders. However, when PCR products of these last 2 primer pairs labeled with fluorescent dye were run in a capillary gel and detected using a DNA analyzer, P2/P8 gave 2 distinguishable peaks in females, whereas 2550F/2718R results remained the same. DNA sexing with any of those 3 primer sets can be used for all sexually monomorphic avian taxa although the primer sets should be compared before choosing the most efficient one for molecular sexing of the studied species.
Introduction
Because many bird species are morphologically monomorphic or only sexually dimorphic in adult stages, it is difficult to determine their sexes accurately, a fact that causes significant problems in many biological studies. Indeed, the sex of individuals is an essential feature for field studies in the fields of ecology, behavior, genetics, breeding, and evolutionary and conservation biology.10,12,24,31 For instance, the sex of individuals is of critical importance in determining sex ratios in natural populations or in similar demographic studies because dispersal or mortality rate may be sex-specific. 8 Similarly, in some cases, foraging behavior,20,28 predation pressure, 15 dispersal,3,19 migration patterns, 11 and evolutionary pressures 16 incurred by sex may be divergent, which demonstrates the importance of sex identification in ecologic and demographic research. Knowing the sex of an individual is also a critical piece of information in captive breeding for conservation programs, for science education, and in terms of contributions to biological diversity. 26 For instance, when birds are exchanged by means of commerce or to prevent inbreeding, knowledge of sex is a prerequisite. 16 Moreover, determining sex in zoos is necessary in order to increase successful reproduction (1993 Meeting of the EEP Penguin TAG, Penguin Conserv 1993;6:10–12, available at: https://goo.gl/J8W6X2).
Although there is no perfect method, molecular techniques that provide accurate and rapid sex identification with noninvasive techniques are broadly used in veterinary clinical practices. 1 Given that male birds are homogametic (ZZ), whereas females are heterogametic (ZW), 32 several DNA-based sexing methods have been described. In DNA-based sexing using polymerase chain reaction (PCR), the highly conserved chromodomain helicase DNA binding (CHD) gene is amplified using specific primers that give distinct banding patterns on agarose gel as a result of intronic regions within this gene.9,17,19 One gene is located on the W chromosome (CHD-W), whereas its homolog is on the Z chromosome (CHD-Z). This leads to females generally displaying 2 bands of different sizes, 1 from CHD-Z and the other from the CHD-W gene fragment, whereas males have 2 identical sized copies.
There are currently several independently developed primer sets for sexing birds using DNA; particularly, 3 such sets are commonly utilized by researchers.12,18,23 However, comparisons between those 3 approaches have not been available, to our knowledge. Therefore, we compared the success rates of 3 different primer sets for sexing birds and sought to expand the DNA-based sexing database through inclusion of additional species.
Materials and methods
A total of 230 samples (feather, n = 57; blood, n = 173) were collected from wild birds captured during fieldwork, or from captive birds kept in Bursa, Antalya, and Ankara Zoos, and at a captive breeding facility in Birecik, Turkey, which covers 77 avian species belonging to 14 orders. A total of 29 species that belonged to Passeriformes were captured with mist nets during other fieldwork. After sampling was completed, all wild and captive specimens were released unharmed back into their natural or captive environments, respectively. Thirty-six species (both female and male specimens; Supplemental Table 1, available at http://vdi.sagepub.com/content/by/supplemental-data) were used only for the comparison of the success rates of 3 primer sets. Approximately 10–100 μL of blood samples were collected and kept in blood collection tubes (K3-EDTA). Alternatively, 1–8 plucked feathers were taken from a specimen’s upper tail coverts and greater wing coverts and kept dry in small paper envelopes. All samples were stored at 4°C until DNA isolation in the laboratory. A blood and tissue kit a was used to isolate genomic (g)DNA by following standard kit protocols.
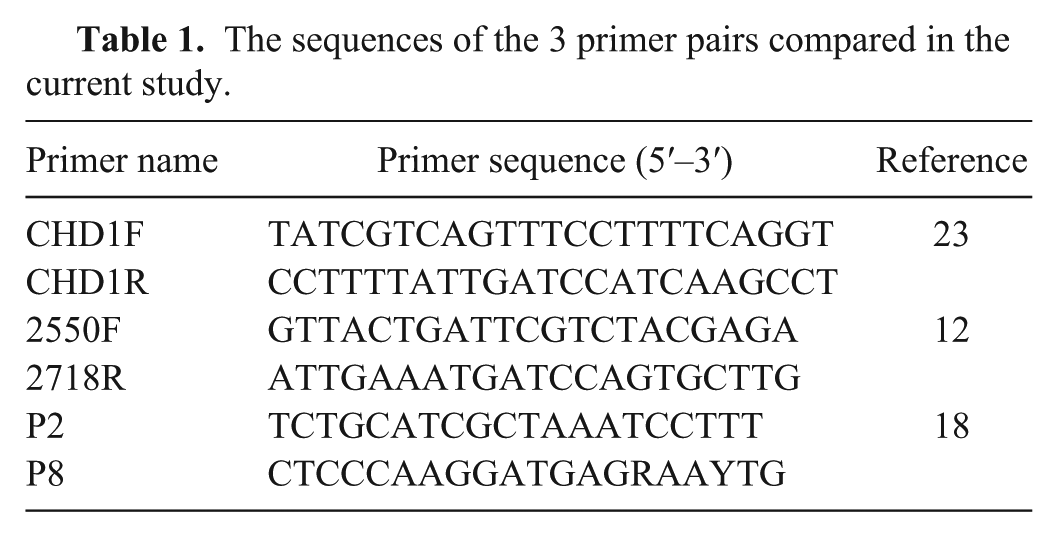
Three primer sets, CHD1F/CHD1R, 23 2550F/2718R, 12 and P2/P8, 18 were used to amplify fragments of the CHD gene (Table 1). The mixture of PCR reaction included 1× PCR buffer, b 1.0 mM MgCI2, 0.2 mM of each primer, 0.2 mM dNTP mix, 1 unit of Taq DNA polymerase, b and 0.1 μg of gDNA. Amplification for P2/P8 included an initial incubation at 94°C for 4 min, followed by 40 cycles at 94°C for 30 s, 51°C for 45 s, and 72°C for 45 s, and further extension at 72°C for 5 min. For amplification of 2250F/2718R and CHD1F/CHD1R, we followed a touchdown scheme 7 where annealing temperature was reduced 1°C per cycle, starting from 57°C, until it reached 50°C, followed by 30 cycles, and a final extension at 74°C for 5 min. PCR products were run on 3% agarose gel for 75 min at 90 V in standard Tris–borate–EDTA buffer. 30
The sequences of the 3 primer pairs compared in the current study.
To confirm the amplification specificity and determine certain differences between lengths of introns in the CHD-W and CHD-Z genes, fluorescent-labeled primers (6FAM) were used; their PCR products were separated by capillary electrophoresis and detected by a DNA analyzer. c Using this technique, it is possible to distinguish PCR bands 3 bp different in lengths, which was efficiently applied previously in several species. 23 Software d was used to perform fragment analysis.
Results
The fragment of CHD gene was amplified using CHD1F/CHD1R, 2550F/2718R, and P2/P8 primer sets to determine the sex of individuals from 77 avian species (Supplemental Table 1); Figure 1 shows their analysis on a simple 3% agarose gel. The analysis of CHD1F/CHD1R PCR products on 3% agarose gel generally showed a definite single band in males (CHD1-Z) and 2 bands in females (CHD1-Z and CHD1-W) except 2 Galliformes species, namely Meleagris gallopavo f. domestica (74F) and Coturnix coturnix (73F), 1 Psittaciformes species (Psittacula alexandri, 2F), and 10 Passeriform species, which all displayed only a single band of different sizes for both sexes. However, 5 specimens, namely 50F (Ficedula parva), 57F (Passer montanus), 54F (Fringilla coelebs), 59F (Lonchura striata domestica), and 60F (Taeniopygia guttata) produced 2 bands in females (Supplemental Table 1). In contrast, the amplification reactions performed with 2550F/2718R and P2/P8 primer sets showed distinct banding patterns for separate bird orders. In some cases, both males and females presented a single band either different or identical in size, or typically 2 bands in females and a single band in males (Supplemental Table 1).
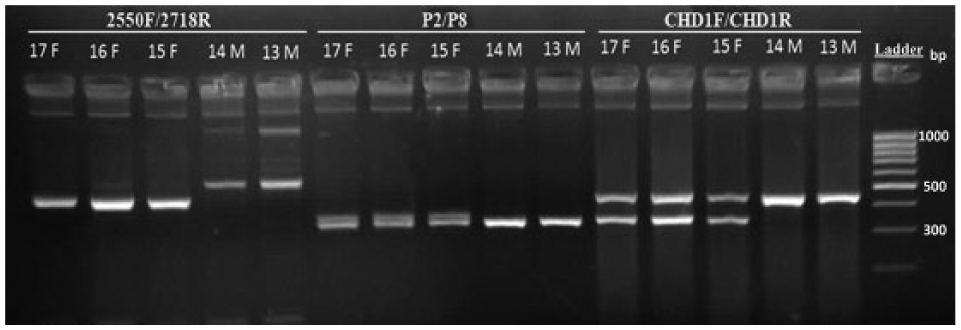
General banding patterns of 3 primer sets (CHD1F/CHD1R, P2/P8, 2550F/2718R) in Cygnus olor blood (lanes 13–17). M = male; F = female; ladder = 100 bp.
In the fragment analysis results, the amplified CHD1-W and CHD1-Z fragments with CHD1F/CHD1R and 2550F/2718R primer sets were 317–696 bp and 440–705 bp, respectively, whereas the product size of CHD1-Z and CHD1-W amplified by P2/P8 primer pairs was 316–419 bp. Remarkably, the size of the Z fragment was larger than the W fragment in CHD1F/CHD1R and 2550F/2718R amplifications, whereas the size of the W fragment was larger than the Z fragment in P2/P8 amplifications. Although the analysis of P2/P8 PCR products on agarose gel showed an apparent single band of equal size in both sexes in the orders Galliformes, Ciconiiformes, Phoenicopteriformes, Gruiformes, Accipitriformes, Columbiformes, and Piciformes, capillary electrophoresis analysis generated 1 peak for males and 2 peaks for females in these orders. However, in spite of the fragment analysis, 2 peaks were still not observed with the 2550F/2718R primer set in analyzed female passerines (n = 21; Supplemental Table 1).
Discussion
It was possible to compare the success rates of sexing 10 of 14 orders using 3 CHD-related primer pairs CHD1F/R, 23 2550F/2718R, 12 and P2/P8 18 (Table 2). Potential errors in interpreting sex-typing data, such as Z-polymorphism 6 and the formation of heteroduplex DNA molecules, 4 were avoided by sexing the same specimen with 3 different primer sets and analyzing the outcome through automated fragment analysis.6,14 The relatively high quality of DNA is directly related to both the number of feathers taken from birds and their sizes rather than freshness or type of feather samples. The success rates of 3 different primer sets in feather samples were 70.2%, 57.9%, and 64.9% for CHD1F/CHD1R, 2550F/2718R, and P2/P8, respectively. Similarly, according to agarose gel electrophoresis, the success rates of primer sets CHD1F/CHD1R, 2550F/2718R, and P2/P8 were 91.2% (n = 230), 56.2% (n = 230), and 50.9% (n = 230), respectively. This can be directly related to the fact that DNA in feather samples generally submit lower copy numbers than DNA obtained from blood, and perhaps DNA from feathers may get more easily degraded, so the quality and quantity of DNA extracted from feather material would be correspondingly low. 22 Although the results of the first primer set remained the same after fragment analysis, the success rate increased slightly with 2550F/2718R primers (57.8%) but increased markedly with the P2/P8 primers (98.4%).
Sexing success rates of the 3 primer sets in 10 avian orders.
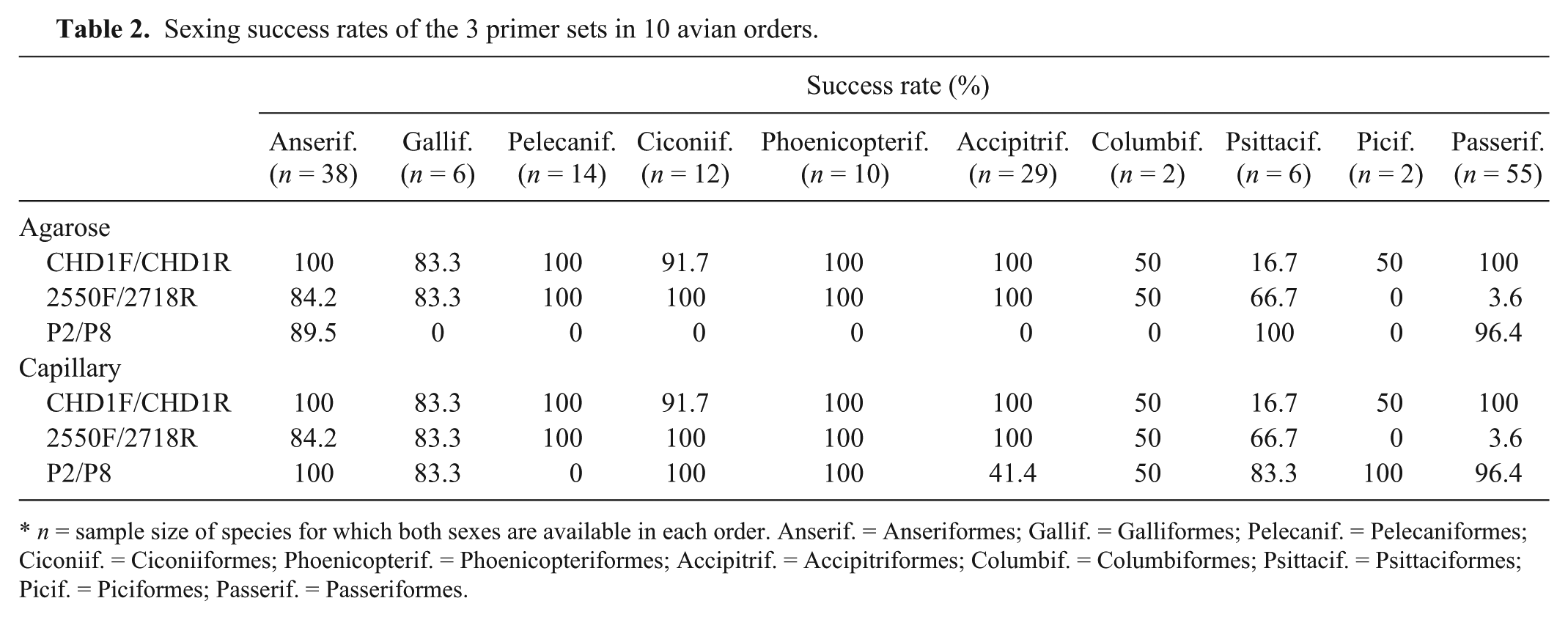
n = sample size of species for which both sexes are available in each order. Anserif. = Anseriformes; Gallif. = Galliformes; Pelecanif. = Pelecaniformes; Ciconiif. = Ciconiiformes; Phoenicopterif. = Phoenicopteriformes; Accipitrif. = Accipitriformes; Columbif. = Columbiformes; Psittacif. = Psittaciformes; Picif. = Piciformes; Passerif. = Passeriformes.
Previously, a single member of Anseriformes (Cygnus olor) was sexed 23 ; in our study, we sexed 8 additional Anseriformes, using the CHD1F/CHD1R primer set. When amplified with the 2550F/2718R primer set using the PCR program we adopted, complete success was not achieved for male specimens in this order, which may be caused by failure of PCR amplification of the CHD-Z fragment. 22 However, C. olor,18,23 Anser albifrons, 24 Tadorna tadorna, and Branta canadensis 2 were previously studied using the P2/P8 primer set. All galliform samples analyzed in our study with the CHD1F/CHD1R primer set were previously sexed, 23 and similar results were obtained. The 2550F/2718R primer set could not generate any PCR bands from Phasianus colchicus and C. coturnix, which may be caused by greater nucleotide diversity in the region to be amplified. 35 Likewise, the sex of C. coturnix could not be differentiated based on size differences,5,27 and, as such, an alternative PCR-based technique was used to determine sex (i.e., single strand conformation polymorphism). Yet, these 2 species were previously sexed 12 using the P2/P8 primers 23 ; C. coturnix did not produce any yields amplified in our study, which may be related to the low quantity of DNA obtained from feathers. Pavo cristatus was also previously sexed with P2/P8 and 2550F/2718R (NBAF-S bird sexing database, available at https://goo.gl/gt3eBn). Species belonging to Pelecaniformes, Ciconiiformes, and Phoenicopteriformes were analyzed using the CHD1F/CHD1R and P2/P8 primer sets in our study. Additionally, all studied species (n = 6) from Accipitriformes and Falco peregrinus were not previously sexed with the CHD1F/CHD1R primers, to our knowledge. Unlike our study, the 2550F/2718R primer set produced only a single band in females of Accipitriformes, 12 which is thought to be a result of different PCR profiles used to amplify the DNA of these species. Although Circus aeruginosus is a sexually dimorphic raptor in terms of plumage, our molecular result was in conflict with the morphology of the specimen. This discrepancy is likely caused by the fact that up to 40% of males of this species look like females. 33 Only half of the 12 Psittaciformes species were sexed with the CHD1F/CHD1R primer set, but there was no previous study on these species, to our knowledge, so we were not able to assess the reasons for this failure although polymorphism in the primer-binding site among distinct species might be an explanation. However, successful results were obtained for those species amplified with the 2550F/2718R and P2/P8 primer sets in both our study and others.13,21,26,29,34 The sex of Bubo bubo using the 2550F/2718R and P2/P8 primer sets could not be determined because of the lack of length polymorphism.34,36 However, using the ARMS (amplification refractory mutation system) technique, it was possible to sex the owl species B. bubo and Otus scops. 24 Our results also indicated success in sexing Dendrocopos syriacus and Jynx torquilla of the order Piciformes. Two species from Columbiformes, namely Streptopelia turtur and Streptopelia senegalensis, were also studied; however, no results could be obtained for either species because of the unsuccessful PCR amplification using CHD1F/CHD1R and 2550F/2718R primer sets, respectively. In the current study, all studied species (n = 29) from Passeriformes were correctly sexed using the CHD1F/CHD1R and P2/P8 primer sets without the necessity of capillary electrophoresis, but, unlike successful results reported (https://goo.gl/gt3eBn), 12 when different PCR profiles were used, sex identification was not possible in our study for this order with the 2550F/2718R primer set. This may be caused by mutations in the primer binding site that leads to a failure of the PCR amplification of the CHD-W fragment, 12 or caused by the effect of the annealing temperature used in the PCR. 25 Twenty-eight species of Passeriformes were sexed in our study using the CHD1F/CHD1R primers; Passer montanus had been previously sexed. 23 Although both males and females amplified with the CHD1F/CHD1R primers produced only single bands of different sizes in our study in almost all passeriform birds, the species from the same order typically generated 2 bands in females and 1 band in males 23 when a different PCR program was used. The P2/P8 primer set also produced typically 2 bands in females and 1 band in males; nevertheless, 1 species, Taeniopygia guttata, generated only 1 male band for a female specimen, which can be explained in 2 possible ways: nucleotide variations on 1 site of primer binding or degraded DNA. 23
CHD gene–based PCR amplification proved to be a reliable, generally accurate, and satisfactory technique for sex identification of avian species of diverse phylogenetic backgrounds. Among the 3 primer sets studied, CHD1F/CHD1R and P2/P8 were able to determine the sex of a wide variety of bird species of different orders, whereas 2550F/2718R was only partially successful. However, it might be necessary to carry out additional fragment analysis (e.g., capillary electrophoresis) with the P2/P8 primer set, thus increasing costs associated with the procedure. Overall, DNA sexing with 3 primer sets can be used for all monomorphic avian species, but we recommend a preliminary comparison of those 3 primer sets for efficiency prior to molecular sexing of large numbers of specimens.
Footnotes
Acknowledgements
We thank the staff of Bursa, Antalya, and Ankara Zoos for help with collecting samples from captive bird specimens. We also thank Soner Oruç for help with sampling wild specimens in the field studies. Burak Doğan helped with both collecting and analysis.
Authors’ contributions
CC Bilgin contributed to conception of the study. CC Bilgin, Ç Akın Pekşen, and E Çakmak contributed to design of the study, acquisition of samples, and analysis and interpretation of results. E Çakmak and ÇA Pekşen did the molecular work. All authors drafted the manuscript, critically revised the manuscript, gave final approval, and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Qiagen, Hilden, Germany.
b.
Fermentas, Thermo Fisher Scientific, Waltham, MA.
d.
Peak Scanner software, Applied Biosystems, Foster City, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by grant BAP-08-11-DPT.2011K121010 at the Middle East Technical University.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
