Abstract
Although Fusarium sp. infections have been reported in aquatic invertebrates, studies of Fusarium spp. as fish pathogens remain very limited. In our study, a fungus was isolated from diseased zebrafish (Danio rerio). DNA sequence analysis of the fungus, based on a partial region of the translation elongation factor 1α gene (EF-1α), the internal transcribed spacer region and domains D1 and D2 of the large subunit of the ribosomal RNA gene (ITS plus LSU), and the RNA polymerase II subunit gene (RPB2), showed 99.9–100% homology to Fusarium solani species complex sequences. Multilocus sequence typing analysis based on 3-locus haplotypes (EF-1α, ITS plus LSU, and RPB2) suggests that the isolated strain was type 3+4-P. Challenge experiments showed that this organism could be pathogenic to zebrafish, but usually does not infect healthy subjects under normal circumstances.
Introduction
Fusarium spp., a large genus of filamentous fungi in the family Nectriaceae, class Sordariomycetes, can be pathogenic to animals and plants, and can exist as saprophagous organisms on decomposed organic matter. 29 Most members of this genus are mycotoxigenic pathogens and are distributed worldwide. 19 Fusarium spp. have been reported as etiologic agents in various human diseases, such as keratitis, endophthalmitis, mycetoma, and endocarditis, and can result in life-threatening systemic infections.3,20,22,23 Over the past 30 years, numerous Fusarium spp. infections have been reported in aquatic invertebrates.14,15,28 However, reports of Fusarium as fish pathogens remain very limited.8,18,27
Zebrafish (Danio rerio) have been widely used as a laboratory model for the investigation of basic mechanisms of vertebrate developmental genetics. 17 Many common zebrafish disease agents have been reported, such as mycobacteria, 25 Pseudoloma neurophilia, 13 and others. 26 However, studies on fungal infections remain very limited.
In the current study, Fusarium solani species complex (FSSC) was isolated from zebrafish, and identified by morphologic and molecular methods. The pathogenicity of the isolated organism was also evaluated in infected zebrafish.
Materials and methods
Diseased fish, fungus isolation, and morphologic identification
From April to September in 2010, a disease outbreak occurred in cultivated zebrafish at the Institute of Hydrobiology, Chinese Academy of Science (Wuhan, China). A 20–30% mortality rate was recorded during the outbreak. The major signs were skin ulceration, belly distension, and mottled liver. Short hyphae were found in ulcerated tissues by light microscopy. Fungus isolation was carried out as described previously. 11 Briefly, the body surface necrotic tissues, liver, kidney, and intestines were aseptically examined by means of Gram stains and 20% KOH stains. Necrotic tissues were washed with distilled water, then in 70% (v/v) alcohol for 3 s, then again in distilled water. The tissues were inoculated on corn meal agar (CMA) supplemented with penicillin–streptomycin (100 μg/mL), and incubated at 25°C. Pure cultures were obtained by the single spore culture method. 10 The morphologic characteristics of the isolates were studied by scanning a and transmission b electron microscopy.
Molecular and multilocus sequence typing analysis
Genomic DNA extraction was carried out as described previously. 11 Primers ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) 30 were used to amplify the intergenic transcribed spacer (ITS) region. Polymerase chain reaction (PCR) conditions were as follows: 1 cycle at 94°C for 5 min; followed by 35 cycles of 94°C for 30 s; 58°C for 30 s and 72°C for 1 min; and an extension cycle at 72°C for 5 min. Amplified fragments were cloned into a commercial vector. c The recombinant plasmids were transformed to Escherichia coli (DH5α) competent cells and sequenced using M13 forward and reverse primers on a DNA sequencer. d The PCR for elongation factor 1α gene (EF-1α) was performed using primers ef1 (forward primer: 5′-ATGGGTAAGGA(A/G)GACAAGAC-3′) and ef2 (reverse primer: 5′-GGA(G/A)GTACCAGT(G/C)ATCATGTT-3′) 21 as described above for the ITS PCR except that the annealing temperature was 53°C. The sequences of products from both PCR assays’ production were compared with those of FSSC sequences from GenBank and Fusarium-ID (http://goo.gl/YZt6Ia).
Three nuclear gene regions were chosen for multi-locus sequence typing (MLST) examination using published phylogenetic analyses: the ITS plus domains D1 and D2 of the nuclear large subunit (LSU) of the ribosomal (r)RNA, EF-1α, and the second largest subunit RNA polymerase II (RPB2).3,23,31 The PCR conditions and primers for amplification of LSU and RPB2 were described in a previous study. 24 The sequence type of the strain was identified by multiple sequences analysis against the Fusarium MLST database (http://goo.gl/W21ztN).
Infectivity experiment
A total of 400 healthy zebrafish (length: 51 ± 2 mm) from the Institute of Hydrobiology were exposed to a conidia suspension of isolate BMY1 through intraperitoneal, wound, and bath infections. The fish were kept in tanks (15/tank) with flowing water (volume: 10 L; water temperature: 25–26°C) and were acclimatized for 15 days before challenge. The experiment was performed in 3 replicates. The fish were fed twice a day with 3% (w/w) common commercial small pellet feed e throughout the experiments. All of the experiments were approved by the Institutional Animal Care and Use Committee at the research facility.
In order to obtain sufficient sporulation, inocula were prepared from cultures of the isolate on CMA slants for 7 days at 25°C. Conidia were harvested by washing the agar surface with sterile saline. Suspensions of conidia were counted in a hemocytometer and adjusted to the desired concentration. Each conidial suspension was shaken vigorously before application to ensure uniform dosage delivery.
In the intraperitoneal infection model, 60 zebrafish from 4 tanks were injected with 10 μL of a spore suspension of BMY1 at the concentrations of 102, 103, 104, and 105 spores per fish (one concentration in one tank), respectively. The fish in the negative control tank were injected with 10 μL of sterile saline. In the wound infection model, 15 fish were artificially grazed by wire netting and then immersed for 2 h in the BMY1 spores suspension (1.0 × 107 spore/mL) in 8-L tanks. The exposed fish were placed in a new tank with aeration (volume: 10 L; water temperature: 25–26°C). Fifteen fish were exposed to saline as controls. In the bath challenge model, 15 fish were immersed for 2 h in the BMY1 spores suspension (1.0 × 107 spore/mL) in 8-L tanks. The exposed fish were placed in a new tank with aeration. Fifteen fish were exposed to saline as controls.
Clinical signs and mortality were monitored and recorded daily. All dead fish were autopsied, and necrotic tissues, liver, and kidney were aseptically streaked on CMA agar to confirm the cause of death. After incubation at 25°C for 48–72 h, fungal identification was carried out as described above. Mortalities were valid only if BMY1 was re-isolated as a pure culture and confirmed by molecular sequence analyses.
Statistical treatment of data
Statistical analysis was performed using commercial software. f Survival curves were calculated by Kaplan–Meier analysis.
Results
Fungus isolation and morphologic characteristics
Five Fusarium isolates obtained from the infected fish had similar morphology, showing colonies that are woolly to cottony with cream-to-white aerial mycelium and a cream reverse on agar. Hyphae were septate. Conidiophores produced from the hyphae were simple (nonbranched) or branched monophialides (Fig. 1A). Conidia varied in size with culture age. Microconidia were ovate, singly formed, and abundant. Macroconidia were produced terminally, normally on short lateral branches. Most of the macroconidia had 1–3 septa at an average size of 15.8–29.2 × 2.8–3.5 μm (Fig. 1B, 1C). In young cultures, conidia were aseptate and measured 7.9–12.8 × 2.4–3.2 μm, and usually occurred at the tip of the phialide. Chlamydospores were also detected (Fig. 1D).
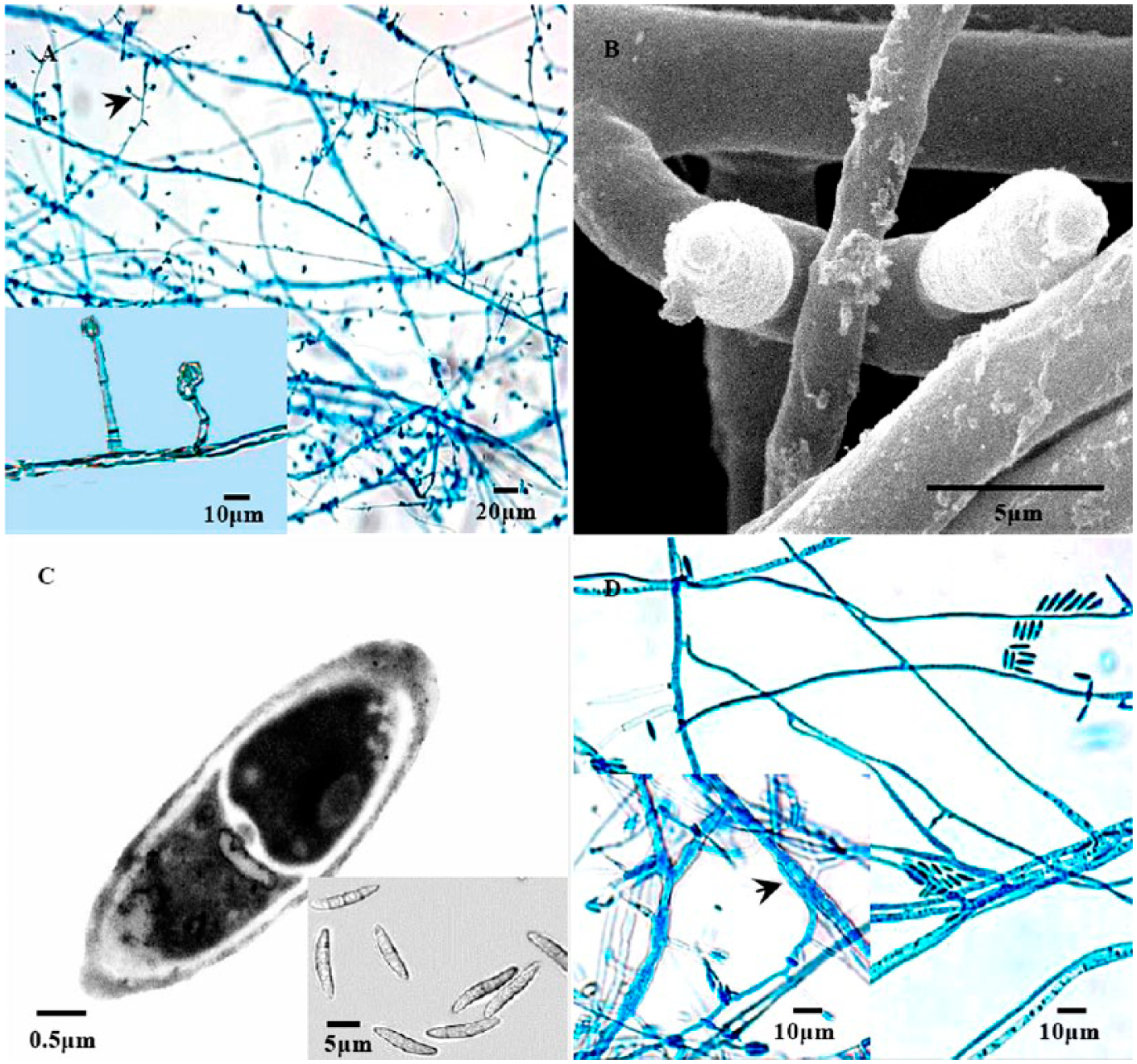
The conidiophore, conidia, and chlamydospore of Fusarium isolate BMY1.
Gene cloning and MLST analysis
Fragments of the ITS rDNA of ~568 base pairs (bp) were obtained by PCR amplification from the 5 Fusarium isolates. The sequences of these 5 fragments were identical (100% similarity). Isolated BMY1 was selected as the representative isolate, and the ITS rDNA sequence of this isolate was deposited in GenBank (accession JF975615). The EF-1α gene fragments (~666 bp) were also amplified from the 5 Fusarium isolates. All the sequences were identical (100% similarity). The amplified nucleotide sequence showed 99–100% homology to other EF-1α gene sequences of FSSC from GenBank and Fusarium ID.
Analysis of the 3 loci (EF-1α, ITS plus LSU, and RPB2) against the Fusarium MLST database identified the isolate as the haplotype 3+4-p. The sequences were deposited in GenBank as accessions KM851046 (EF-1α), KJ653811 (LSU), and KJ653812 (RPB2).
Infectivity experiment
A cumulative mortality curve (Fig. 2) was used to illustrate the results of the challenge experiment. With intraperitoneal challenge of 105 spores per fish, there was 100% mortality within 13 days after challenge. A lower level of mortality was seen with the concentrations of 104, 103, and 102 spores per fish (86.7%, 60%, and 53.3%, respectively). The first dead fish was observed on the 2nd day in the tank with 105 spores per fish. On the 3rd and 4th day, dead fish were also observed in tanks with 104, 103, and 102 spores per fish (Fig. 2). Before death, most infected fish showed behavioral changes, such as loss of appetite, lethargy, and spiral swimming. Other clinical signs include severe congestion in the head, fins, and perianal region (external), belly distension, and mottled liver (internal). Some infected fish displayed perforated abdomen and pale gills.
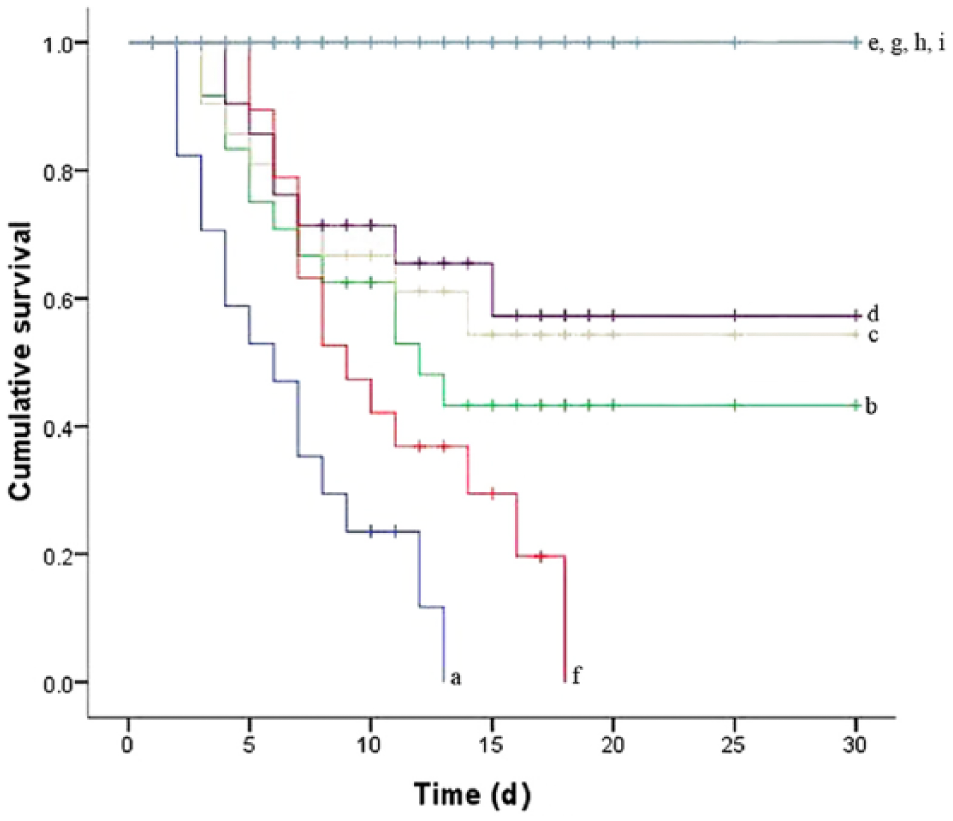
Cumulative survival curve:
In the wound challenge study, all of the fish were dead by the 18th day, with the first fish death on the 5th day (Fig. 2). Between the 2nd and 3rd days, clinical signs similar to the intraperitoneal challenge were observed, in addition to skin ulcerations. A few fish showed short hyphae in and around the wound areas when examined microscopically. Typically, the diseased fish floated weakly in the water prior to death. Isolate BMY1 was recovered from dead fish and sampled live fish. No mortality or visible changes were observed in the bath challenge and in any of the control groups (Fig. 2).
Discussion
Based on morphologic analysis, the BMY1 isolate was assigned to the genus Fusarium in the current study. However, identification within the genus Fusarium is difficult if only based on highly variable morphologic characteristics.4,7 A variety of DNA-based molecular approaches have been developed for the identification of fungal pathogens in recent years.2,7,21,22 Analysis of the EF-1α gene is an effective molecular identification tool in Fusarium, 7 and a combination of this gene with the ITS region has been used for phylogenetic analysis of FSSC. 4 In addition, MLST based on 3 loci (ITS plus LSU, EF-1α, RPB2) has been widely used to identify phylogenetic relationships at the interspecific level. 24 These markers in phylogenetic and population studies have led to novel species delimitations within the FSSC and haplotype nomenclature system.24,31 In our study, the isolated strain was identified as a member of the FSSC comprising >45 phylogenetic or biological species. 31 Interestingly, no other Fusarium species was recovered from the diseased fish in the study. These findings suggest that this common environmental saprophyte, 6 responsible for approximately two-thirds of all human and animal fusariosis worldwide, 7 may be responsible for a zebrafish disease outbreak.
In the challenge experiments, the cumulative mortalities from intraperitoneal and wound infection models indicated that F. solani was pathogenic to zebrafish. However, in the intraperitoneal challenge, the fish died more quickly without any body surface ulceration, which was different from the natural infection. The intraperitoneal inoculation appears to reduce the period before disease onset and result in a higher mortality rate. In the wound infection, similar clinical signs to those seen in the natural outbreak were present. There was no mortality or disease in fish exposed by bath infection. These results suggest that the FSSC isolate BMY1 can be highly pathogenic only after invading the epidermis, which is similar to infections of Fusarium fusarioides in Micropterus salmoides, 9 and Mucor circinelloides in Pelteobagrus fulvidraco. 12
Some metabolic products of FSSC, including javanicin, fusarubin, anhydrofusarubin, and fusaric acid, have been demonstrated to have toxic and enzymatic activities.1,5,8,16 The cartilage of bonnethead sharks 18 and the exoskeleton of insects and crustaceans 14 can be penetrated by F. solani. Trauma and stress may increase the susceptibility of fish to fungal infections, especially by providing easier access to tissues that could be substrates for fungal enzymes, which play an essential role in the infection process. Therefore, such infections may not be confined to the wound area but could become systemic and lead to mortality.
Footnotes
Authors’ contributions
X Ke contributed to conception and design of the study, and drafted the manuscript. M Lu contributed to conception and design of the study. J Wang contributed to design of the study. All authors contributed to acquisition, analysis, or interpretation of data; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Quanta 200 SEM, FEI, Eindhoven, The Netherlands.
b.
HT-7700, Hitachi Ltd., Tokyo, Japan.
c.
pMD-19T vector, Takara Biotechnology Co. Ltd., Dalian, China.
d.
Applied Biosystems, Carlsbad, CA.
e.
Tongwei Group Co. Ltd., Chengdu city, Sichuan Province, China.
f.
SPSS 17.0 software, SPSS Inc., Chicago, IL.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by China Agriculture Research System (CARS-49) and Exploration Program of the Institute Field (095A03).
