Abstract
Introduction
The most common diagnostic methods for canine infectious cyclic thrombocytopenia are the direct identification of the morulae in blood smears, serologic methods to detect antibodies, and DNA amplification by PCR.
17
The newer diagnostic tools are typically based on PCR, with advantages of greater sensitivity and specificity. Targets such as the heat-shock operon, 23S rDNA, and the citrate synthase gene have been used in diagnostic and phylogenetic studies, as well as in the characterization of
Materials and methods
Standardization of qPCR
Primers and probe were designed using commercial software
a
to examine the sequences of the
To optimize the primer and probe concentrations, concentration curves were generated using the positive control DNA as a template. The positive
After the standardization of the optimal concentrations of primers, the optimal probe concentration was determined. Single-probe assays were run in 4 replicates for each concentration (50 nM, 100 nM, 150 nM, 200 nM, and 250 nM). The PCR to determine the optimal concentration of the probe was performed in a final volume of 20 µL comprising 1× commercial PCR master mix,
g
600 nM of each primer, 3 µL (100 ng/µL) of
Analytical sensitivity and specificity of qPCR
The analytical sensitivity of qPCR was evaluated by using serial decimal dilutions of the amplicon cloned into plasmid.
h
Plasmid DNA concentration and purity were verified using a spectrophotometer.
i
The plasmid copy number versus Cq values were plotted to determine the analytical sensitivity of the qPCR based on commercial technology.
j
The number of copies ranged from 1 to 1 × 104 per µL, with 5 separate dilution series performed for each point of the curve in triplicate. This assay was performed using an exogenous internal positive control reagent (termed E-IPC)
k
that contains a probe marked with a reporter dye
l
and a quencher.
m
The linear regression, along with the determination coefficient (
The specificity of the assay was evaluated using DNA from the following related organisms and other pathogens transmitted by ticks:
Sampling
Between June 2011 and June 2012, 186 blood samples were collected from domestic dogs in the state of Rio de Janeiro, Brazil. Collection was performed by cephalic venipuncture, and the collected blood was placed in sterile tubes containing ethylenediaminetetra-acetic acid (EDTA). These samples were used to determine the performance of the new qPCR technique compared to both cytology (blood smear) and nPCR, which targeted 16S
DNA extraction
Genomic DNA was extracted from 300 μL of each blood sample using a commercial DNA purification kit, n according to the manufacturer’s recommendations. Next, the samples were quantified using a spectrophotometer i and diluted in aliquots of 100 ng/µL. As noted above, the positive blood control was obtained from a symptomatic dog with platelet inclusions. Nuclease-free water o was used as a negative control.
Evaluation of the qPCR performance
The E-IPC was added to the reaction to identify the presence of inhibitors. The experiment performed was a standard curve reaction carried out in duplicate using the same equipment used for qPCR.
p
The final volume was 20 µL comprising the following components: 1× E-IPC mix,
k
1× E-IPC DNA,
k
1× universal PCR master mix,
k
600 nM of each primer (
Nested PCR
To evaluate the performance of qPCR, the same DNA samples extracted from whole blood were also submitted to nPCR using primers targeting the 16S
Blood smear evaluation
Smears were prepared using the same whole blood samples used for DNA extraction and qPCR. The laminas were fixed in methanol, stained with Giemsa, and subjected to immersion microscopy (1,000×) to enable visualization of morulae in platelets. Approximately 100 fields were evaluated per lamina.
Statistical analysis
The McNemar test at a 5% significance level was used to measure the proportions of disagreement between qPCR and the other tests evaluated for the diagnosis of
Results
After designing and testing the probe and primers, it was found that the sequences were specific to the target after analysis in silico using a software tool as well as the in vitro test using DNA from other members of family
The detection limit of the qPCR was 1 copy of the plasmid per microliter in both tested curves. The determination coefficient of the 5 dilutions tested in the standard curve with the E-IPC was 99%, with a Cq range of 23.66 ± 0.17 cycles in the first dilution (1 × 104 plasmid copies) to 37.77 ± 0.35 cycles in the last dilution (1 plasmid copy; Fig. 1). The amplification curve with the E-IPC had the same layout during the exponential phase, and the reaction efficiency was 96.06%. This finding demonstrates that, even in reactions with few targets, the reaction was a success. Without the E-IPC, the reaction efficiency of the PCR was 101.80%. The determination coefficient of the 5 dilutions tested in the standard curve with the E-IPC was 99%, with a Cq range of 23.52 ± 0.10 cycles in the first dilution (1 × 104 plasmid copies) to 36.88 ± 0.40 cycles in the last dilution (1 plasmid copy; Fig. 2). As before, even in reactions with few targets, the reaction was efficient.
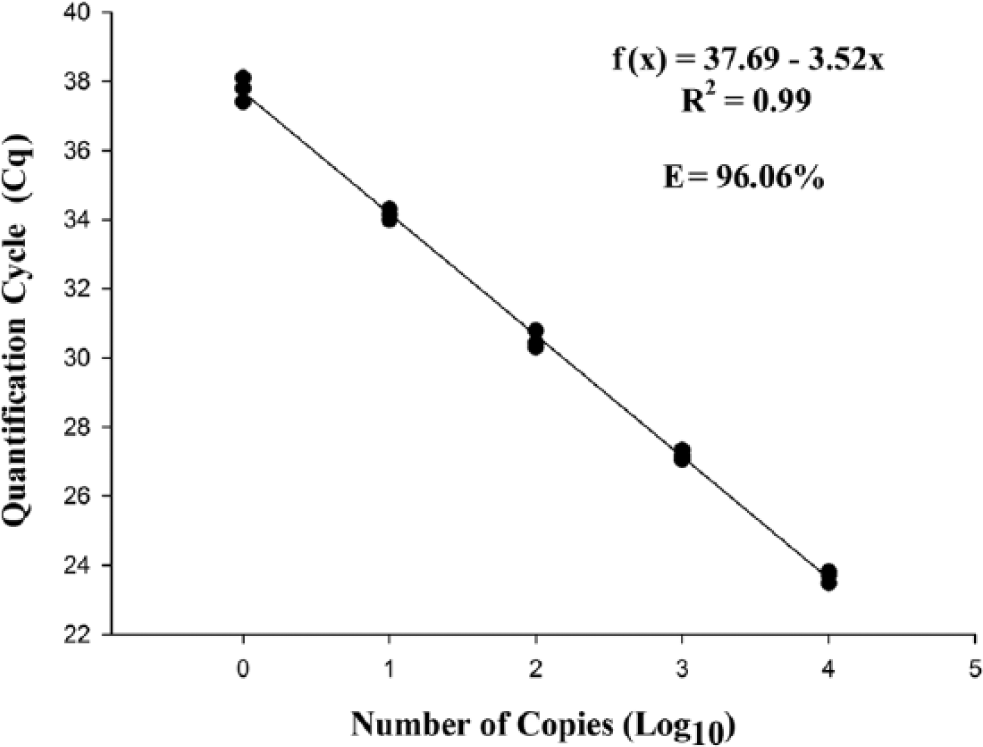
Standard curve plotted from serial decimal dilutions of plasmid DNA, containing the
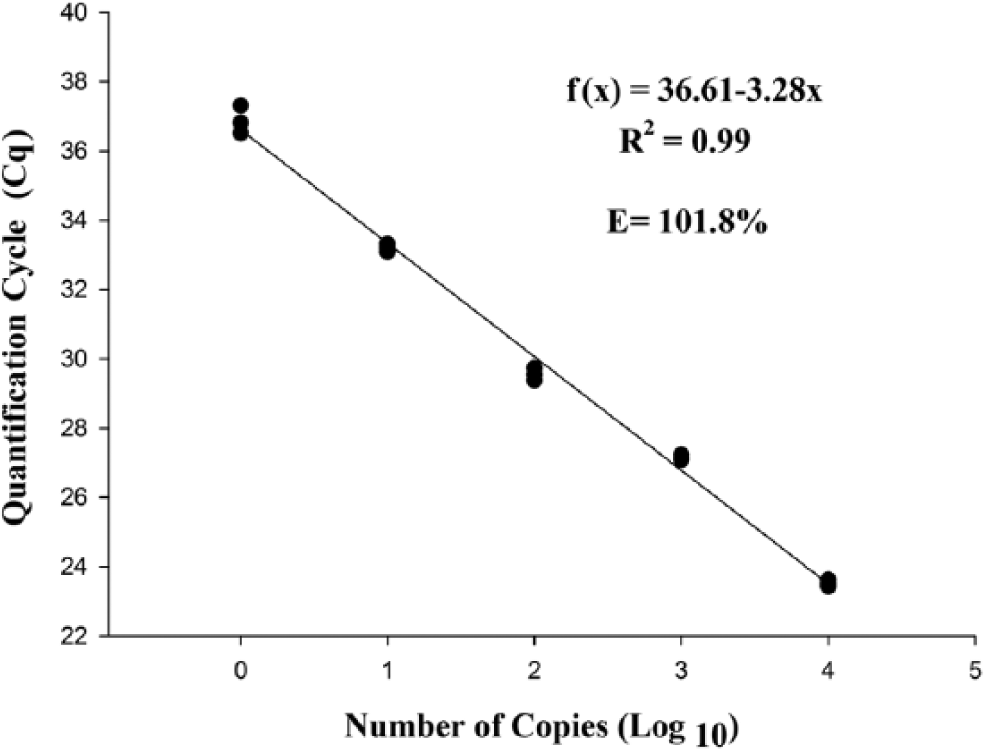
Standard curve plotted from serial decimal dilutions of plasmid DNA, containing the
Regarding qPCR specificity, only
Relationship between the quantification cycle (Cq) and the
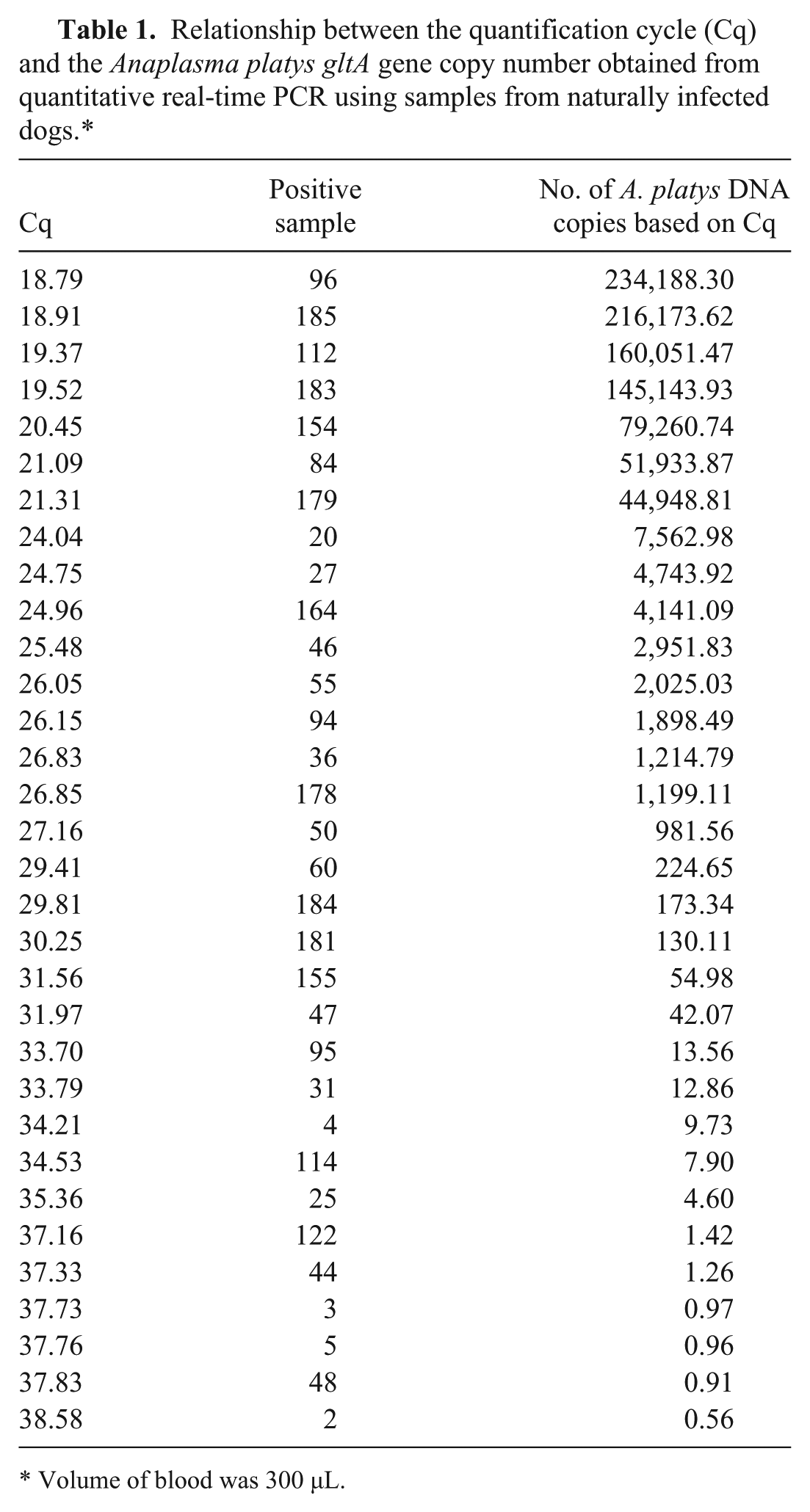
Volume of blood was 300 µL.
All positive nPCR samples were found to be positive when using qPCR. However, nPCR did not detect the target fragment in 18.75% (
Analysis of the disagreement between the results from the quantitative real-time (q)PCR and nested (n)PCR assays and blood smear tests.
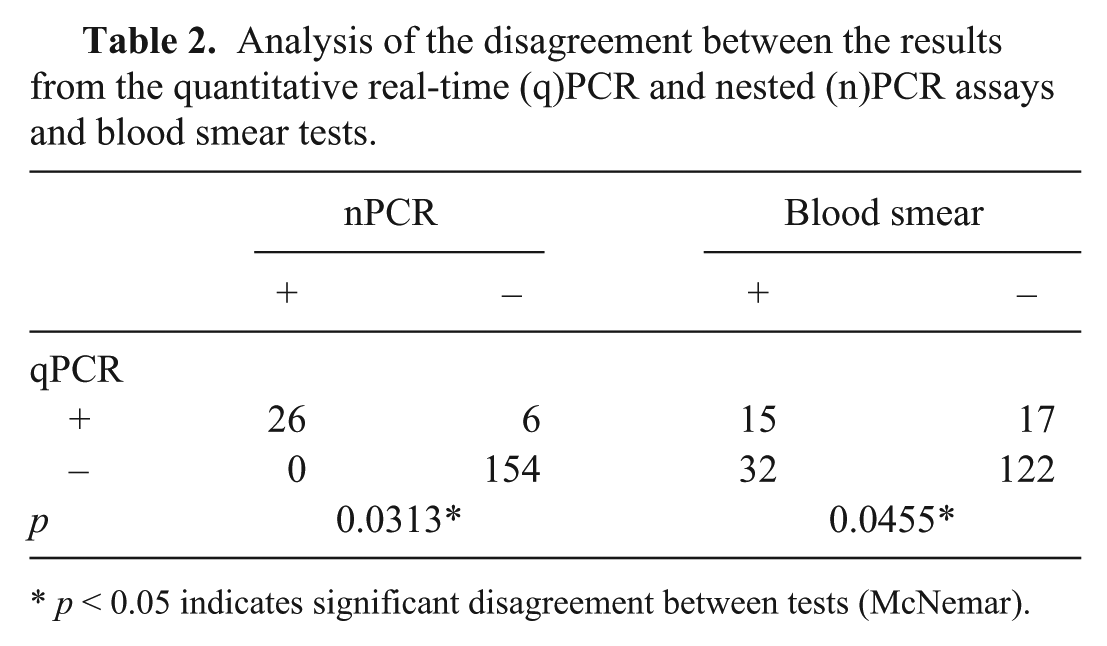
In the blood smears evaluation, 25.27% (
Discussion
This study presented the development of a hydrolysis probe qPCR assay targeting the
The quantitative reverse transcription (qRT)-PCR targeting 16S rRNA has been used previously as a molecular diagnostic tool.37,39,40 The qRT-PCR test is considered a robust diagnostic method because ribosomes and rRNA are present in a higher copy number, because of their role in the translation of genetic code, than genes on the chromosome. 37 However, a RNA-based technique is demanding because of the RNA degradation problem and the difficulty in handling RNA samples. 38 The use of a DNA-based method for the detection of hemoparasites, as used in this and other studies,12,14,19,22 is a less demanding approach. The qPCR technique can be used to monitor amplification products during the reaction and eliminate post-amplification processing, thus decreasing the turnaround time for the assay. 16 In addition to the operational advantages, the qPCR technique has a quantitative nature and is both sensitive and reproducible, potentially replacing direct PCR assays in diagnostic routines. 32
The primers and the probe used in our study were designed to provide high specificity and sensitivity. At the end of the probe, there is a purine (guanine), whereas in the
Our results showed that the potential inhibitors in the reaction were absent because of adequate amplification of E-IPC used in this assay. False-negative results in PCR may be related to the presence of inhibitors, problems with assay sensitivity, or unexpected pipetting errors. 10 The use of an internal positive control is common in reaction optimizations, and it is described as an appropriate tool because it determines whether the absence of specific amplification is actually a result of the absence of the target DNA. 36
Marked thrombocytopenia and low parasitemia occur in dogs during the initial period of
The adoption of a standard curve to evaluate qPCR efficiency is considered a quality benchmark for the assay, 43 and it is recommended in guidelines for qPCR experiments. 8 It has been shown that the optimum efficiency for qPCR assays range from 90% to 110%. 44 The efficiency for the qPCR developed in our study was 96.06%, which is considered desirable for adequate target detection in a qPCR reaction. 44 The curve without the E-IPC generated in this study had a higher efficiency (101.80%). This demonstrates that the presence of the E-IPC can interfere with the reaction because of the occurrence of competition, and thus lower the amplification efficiency of qPCR, a finding that has been previously reported. 21 However, this does not diminish the role of the E-IPC, which in our study was useful in confirming the absence of inhibitors in the reaction, maintaining the credibility of qPCR.
One of the
The false-positive cytology results may be explained by the occurrence of granulations in platelets a result of platelet activation,
17
dense granules, nuclear remnants of megakaryocytes,
29
technical artifacts, or inclusions of
Our study describes the development of a hydrolysis probe qPCR assay targeting the
Footnotes
Authors’ contributions
CB da Silva, MS Pires, M Peckle, HA Santos, and CL Massard contributed to conception and design of the study, and contributed to acquisition, analysis, and interpretation of data. JAR Vilela contributed to conception and design of the study, and contributed to acquisition of data. RL da Costa and GLV Vitari contributed to design of the study, and contributed to analysis and interpretation of data. LA Santos contributed to design of the study, and contributed to analysis of data. All authors drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Primer Express 3.0, Thermo Fisher Scientific, Waltham, MA.
b.
Oligo Explorer 1.2, Gene Link Inc., Hawthorne, NY.
c.
primerBLAST, NCBI, Bethesda, MD.
d.
SYBR Green assay, Applied Biosystems, Carlsbad, CA.
e.
Getting Started Guide of Primer Express Software 3.0, Applied Biosystems, Foster City, CA.
f.
SYBR Green PCR master mix, Applied Biosystems, Carlsbad, CA.
g.
TaqMan universal PCR master mix, Applied Biosystems, Foster City, CA.
h.
pGEM-T Easy vector system, Promega Corp., Madison, WI.
i.
Spectrophotometer Nanodrop ND-2000, Thermo Fisher Scientific, Wilmington, DE.
j.
TaqMan Technology, Applied Biosystems, Foster City, CA.
k.
TaqMan exogenous internal positive control reagents (VIC probe), Applied Biosystems, Foster City, CA.
l.
VIC, TaqMan Technology, Applied Biosystems, Foster City, CA
m.
TAMRA (carboxytetramethylrhodamine), TaqMan Technology, Applied Biosystems, Foster City, CA.
n.
Wizard genomic DNA purification kit, Promega Corp., Madison, WI.
o.
Ambion nuclease-free water, Thermo Fisher Scientific, Wilmington, DE.
p.
StepOnePlus real-time PCR system, Applied Biosystems, Foster City, CA.
q.
L-PIX Touch system, Loccus Biotecnologia, Cotia, São Paulo, Brazil.
Declaration of conflicting interests
The author(s) declared that there is no conflict of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. The National Council for Scientific and Technological Development and the “Carlos Chagas Filho” Foundation for Research Support of the State of Rio de Janeiro provided financial support.
