Abstract
Many methicillin-resistant Staphylococcus aureus (MRSA) strains are multidrug-resistant; consequently, infectious diseases involving MRSA are recognized as troublesome diseases not only in human health care but also in animal health care. A bovine mastitis case caused by MRSA isolates of the New York/Japan clone (NJC), which occurred in Japan in 2005, was monitored in the current study. Isolates of the NJC are typical of hospital-acquired MRSA in Japan. The genetic backgrounds of these strains differ from those of bovine-associated S. aureus, which are typically of clonal complex (CC)97, CC705, and CC133. Moreover, the NJC isolates in this bovine outbreak possessed a β-hemolysin–converting bacteriophage and an immune evasion cluster, as found in the NJC isolates from humans, so it is possible that this clone was introduced into the dairy herd by a human carrier. Most bovine intramammary infections (IMIs) caused by the NJC isolates in our study were asymptomatic, and obvious clinical signs were recognized in only the first 3 infected cows. Of a total of 78 cows, 31 cows were MRSA carriers, and these carrier cows were detected by testing the milk of all lactating cows at 1-month intervals. These S. aureus carrier cows were culled or the infected quarter was dried off and no longer milked. Both IMI and mastitis caused by MRSA were completely eradicated after 5 months. Genotyping data suggested that exchanging of the staphylococcal cassette chromosome mec (the determining factor in methicillin resistance) occurred easily between MRSA and methicillin-sensitive S. aureus in the udders of carrier cows. This case study demonstrates an effective procedure against the spread of MRSA in a dairy herd, and highlights the risk of emergence of new MRSA strains in a dairy herd.
Introduction
Staphylococcus aureus is an important causal bacterium of bovine mastitis, but it is also associated with various infectious diseases in humans. 28 Bovine intramammary infection (IMI) caused by S. aureus is characterized by subclinical mastitis and long-term persistent infection, and the curative effect of antimicrobial therapy is low. 17 Therefore, IMI due to S. aureus spreads insidiously and is very difficult to prevent and cure. 17 Moreover, bovine mastitis due to methicillin-resistant S. aureus (MRSA), well known as a typical multidrug-resistant organism, has been reported in all parts of the world and has attracted a great deal of attention.12,19
Notably, MRSA possesses the mecA gene encoding PBP2a, which is a modified penicillin (PC)-binding protein that plays a role in methicillin resistance. The mecA gene is located on a mobile genetic element, designated the staphylococcal cassette chromosome mec (SCCmec). 5 To date, 11 main types of SCCmec have been distinguished, and some of them possess additional resistance genes.5,21,31 Importantly, SCCmec types I, II, and III, some subtypes of IV, and type VIII determine multi-resistance, as these cassettes contain additional drug-resistance genes on integrated plasmids (pUB110, pI258, and pT181), and transposons (Tn554, Ψ Tn554, and Tn4001). Moreover, additional drug-resistance genes (ant(4′), tet, ermA, ble, aad9/spc, cad, and aacA-aphD) are also possessed by some MRSA strains.5,21,31 Thus, some MRSA strains exhibit not only resistance to β-lactams but also show multidrug resistance.5,21,31
MRSA is a typical causal agent of nosocomial infectious diseases in human hospitals worldwide, and some clonal groups have achieved pandemic spread as hospital-acquired MRSA (HA-MRSA). 5 The genetic background of MRSA is usually described both in terms of sequence type (ST), as determined by multilocus sequence typing (MLST), and SCCmec type. Twelve HA-MRSA clones have been previously reported as pandemic clones, of which the New York/Japan clone (NJC; ST5-SCCmecII) is well known. 5 The NJC is also the major HA-MRSA in Japan, and nosocomial infection caused by NJC has been reported widely in Japanese hospitals.5,16
MRSA has become a notable problem as a causal pathogen in domestic animal diseases including bovine mastitis, but most cases of this disease are caused by S. aureus ST398 (SCCmecIV or V), well known as livestock-associated (LA-) S. aureus (LASA).12,19 Cases of domestic animal illness caused by HA-MRSA are uncommon; consequently, these disease symptoms and actual situations are uncertain.12,19 The current study describes an outbreak of bovine mastitis caused by the isolates of the NJC as well as the use of molecular epidemiological methods to monitor the outbreak. Details that include epidemiological data, bacterial profiles, and countermeasures taken against the outbreak could serve as useful references for understanding the characteristics of the disease and for discussing countermeasures against new outbreaks.
Materials and methods
Monitored farm
The cases of bovine mastitis and IMI that were monitored in this study occurred between March and August 2005. All of the lactating cows were checked by bacterial examination with or without clinical symptoms, and various samples were collected as described later. Cows from which MRSA was isolated were culled or the infected quarter dried off as soon as possible. This outbreak was followed for 5 months from the last isolation of MRSA, and termination of the outbreak was established.
The dairy farm monitored in this study houses ~60 milking cows in a tie stall barn, and the cows are milked twice daily using 4 milking machines. The arithmetic mean of the somatic cell counts in milk bulk tank was 114,000/mL, and the arithmetic mean of the gross amount of milk from each individual cow was 8,500 kg during 2004. Although this farm had purchased both cows and heifers, cows from this farm were not sent out to any other farms.
Staphylococcus aureus isolates
Staphylococcus aureus isolates were mainly collected from individual milk and bulk milk through the testing of milk from all lactating cows at 1-month intervals from March to December 2005. Individual milk was first collected as composite milk. When S. aureus was isolated from composite milk, quarter milk was re-collected. Swabs of each cow’s body and surroundings and of the equipment and antiseptics (dipping) used in milking were also collected at the same time. After 2006, abnormal milk samples (i.e., mastitic milk) were checked as needed until March 2009. Samples of bovine milk were collected aseptically. In addition, sampling of the milking machines and antiseptics was carried out after milking, and samples from the bodies of the cows and their surroundings were collected before milking. Samples other than milk were collected by wiping the area with a sterile cotton swab dampened with phosphate buffered saline. 15
A 100-μL aliquot of all samples was cultured on 5% sheep blood agar at 37°C with 5% CO2 for 18 h, and smooth colonies showing a zone of β-hemolysis or double-hemolysis were interpreted as S. aureus–like colonies, and subcultured. Moreover, S. aureus–like colonies were counted when milk samples were cultured. Bacterial isolates that were coagulase, a catalase, and Voges–Proskauer test a positive and showed coccal cell morphology were selected. The selected isolates were identified as S. aureus by PCR testing for the species-specific 442-bp fragment 15 within the Sa 442 gene 15 or by sequencing of the 16S ribosomal RNA gene (Supplemental Table 1, available at http://vdi.sagepub.com/content/by/supplemental-data). 8 The amplification of the target within the Sa 442 gene was performed using a commercial kit. b The composition of the PCR mixture was 10–100 ng template DNA, 0.2 μM (each) primers, 200 μM (each) deoxyribonucleotide triphosphates, 1× buffer, b and 1.25 U of DNA polymerase. b Filter-sterilized (0.22-μm pore size) PCR-quality water was added to a final volume of 50 μL. The PCR amplification conditions comprised denaturation at 98°C for 10 s, annealing at reported temperature (see Supplemental Table 1) for 15 s, and extension at 68°C for 1 min/kb; these steps were performed for 30 cycles. A total of 97 S. aureus isolates were consequently collected from samples in this outbreak. As well, a total of 40 isolates, confirmed as NJC and collected from human samples in Japan, 14 were used as a basis for investigation into the relationship between MRSA isolates in this outbreak and the typical human isolates in Japan.
Pulsed-field gel electrophoresis of S. aureus isolates
Preparation of genomic DNA plugs was performed as described previously. 20 The digestion of genomic DNA plugs by restriction enzyme SmaI, and the conditions for electrophoresis, were as previously described. 15 The pulsed-field gel electrophoresis (PFGE) pulsotypes (PTs) were analyzed by visual inspection using commercial software c according to the criteria described previously.26,30 Briefly, a dendrogram was created using the Dice coefficient with optimization and a position tolerance setting of 1% based on the banding patterns of each PT. The PTs that showed >80% similarity in this dendrogram were defined as the same cluster. 26
Bacteriological and genetic characteristics of S. aureus isolates
Genomic DNA of the S. aureus strains was extracted with a commercial kit per the manufacturer’s instructions. d The various typing methods (MLST, 9 SCCmec typing, 32 and X region of protein A gene (spa) typing 13 ) were performed as described in previous studies and on the SCCmec website (http://goo.gl/PCqMLO; Supplemental Table 1). The allele number and ST for MLST were determined using the MLST website (http://www.mlst.net/). The founder and clonal complex (CC) of each ST were determined by the enhanced version of the Based Upon Related Sequence Types (eBURST) program, 10 while numeric spa repeats and type codes were determined using the Ridom SpaServer website (http://www.ridom.de/spaserver/).
The coagulase serotype was confirmed using a commercial kit. e Capsules were serotyped using rabbit antisera against S. aureus capsule serotypes 1, 2, 5, and 8 following a published procedure. 25 Detection of the epidermal cell differentiation inhibitor gene (edin), 22 hemolysin genes (hla, hlb, hld, and hlg), 22 leucocidin genes (lukE/lukD, lukM/lukF’-PV, and pvl), 22 streptolysin-associated protein SagD- and SagB-like protein genes (sagD and sagB, respectively), 16 staphylococcal enterotoxin and enterotoxin-like toxin genes (sea to sei, and selj to selr), 27 and toxic shock syndrome toxin (TSST-1) gene (tst) 27 was performed with the primers described previously using a commercial kit b (see details above for PCR conditions and Supplemental Table 1). Staphylococcal enterotoxin, enterotoxin-like toxin, and TSST-1 are termed pyrogenic toxin superantigens (PTSAgs). To help assess the relationship between the NJC strains isolated from humans and the bovine outbreak isolates, the detection of β-hemolysin–converting bacteriophage (int) and immune evasion cluster (IEC)–encoding virulence factor genes was performed as described previously (Supplemental Table 1). 18 These virulence factor genes include staphylococcal complement inhibitor (scn), chemotaxis inhibitory protein (chp), staphylokinase (sak), staphylococcal enterotoxin A (sea), and staphylococcal enterotoxin P (sep).
Antimicrobial susceptibility test of S. aureus isolates
The minimum inhibitory concentrations (MICs) of the S. aureus isolates were determined according to the Clinical and Laboratory Standards Institute (CLSI) method of performing the agar dilution antimicrobial susceptibility test for bacteria that grow aerobically. 3 The tested antibiotics were PC, ampicillin (ABPC), dicloxacillin (DCX), nafcillin (NF), cefazolin (CEZ), kanamycin (KM), gentamicin (GM), erythromycin (EM), oxytetracycline (OTC), tylosin (TS), oxolinic acid (OA), chloramphenicol (CP), enrofloxacin (ERFX), oxacillin (OX), and vancomycin (VCM). Except in the cases of OX and VCM, the test was performed by inoculating S. aureus isolates onto Mueller–Hinton agar f supplemented with diluted antibiotics. Muller–Hinton agar supplemented with 4% (w/v) NaCl was used for the OX tests, and brain–heart infusion agar f was used for the VCM tests. The breakpoints for each antimicrobial were based on CLSI guidelines.2,4 As for antimicrobials for which the breakpoints are not established (DCX, NF, OTC, TS, and OA), the bullet value of the bimodal distribution of each antimicrobial was defined as the breakpoint. 24 Reference strain S. aureus ATCC 29213 served as the assay control. Of tested antimicrobials, acceptable quality control ranges of 4 antimicrobials (DCX, NF, OTC, and OA) are not established, 4 therefore MICs of all tested antimicrobials for S. aureus ATCC 29213 in our study are listed in Table 1 for subsequent judgment of quality control. The presence of the mecA gene was tested by PCR as previously described (Supplemental Table 1). 23
The relationship between genotypes and minimal inhibitory concentration (MIC) ranges in Staphylococcus aureus isolates from a bovine mastitis outbreak in Japan.*
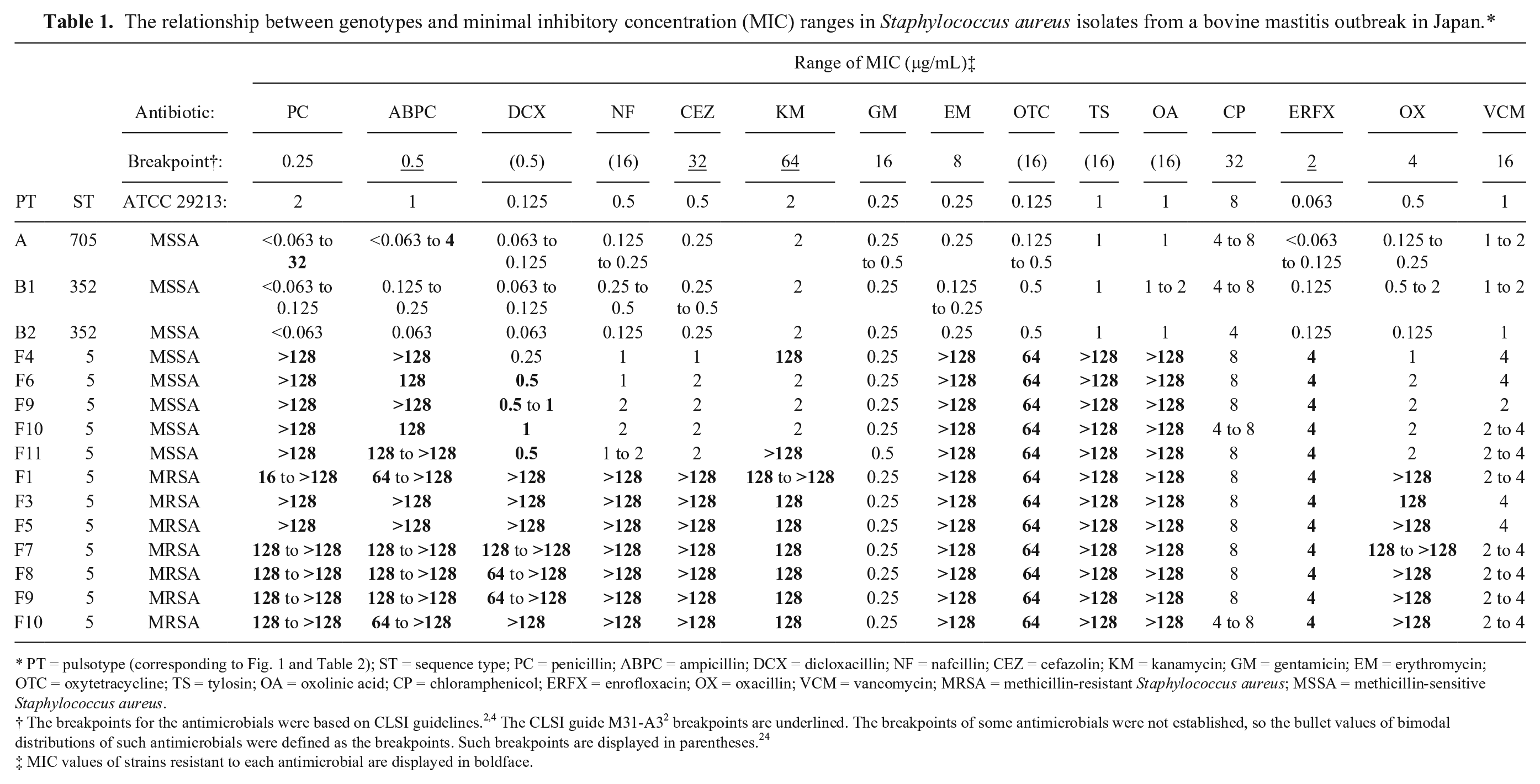
PT = pulsotype (corresponding to Fig. 1 and Table 2); ST = sequence type; PC = penicillin; ABPC = ampicillin; DCX = dicloxacillin; NF = nafcillin; CEZ = cefazolin; KM = kanamycin; GM = gentamicin; EM = erythromycin; OTC = oxytetracycline; TS = tylosin; OA = oxolinic acid; CP = chloramphenicol; ERFX = enrofloxacin; OX = oxacillin; VCM = vancomycin; MRSA = methicillin-resistant Staphylococcus aureus; MSSA = methicillin-sensitive Staphylococcus aureus.
The breakpoints for the antimicrobials were based on CLSI guidelines.2,4 The CLSI guide M31-A3 2 breakpoints are underlined. The breakpoints of some antimicrobials were not established, so the bullet values of bimodal distributions of such antimicrobials were defined as the breakpoints. Such breakpoints are displayed in parentheses. 24
MIC values of strains resistant to each antimicrobial are displayed in boldface.
Results
In this mastitis outbreak, a total of 78 cows were examined, and S. aureus was isolated from 37 animals, with MRSA isolated from 31 of the 37 cows. Clinical symptoms (i.e., sclerosis and swelling of the quarters, and clotted milk) were observed in only 3 cows that were diagnosed with mastitis in March and April 2005. These clinical mastitis cases were caused by MRSA. On the other hand, algia of the quarters was not observed in these cows. The remaining IMIs did not show obvious mastitic symptoms. Infection by MRSA in multiple quarters was confirmed in 13 cows, and these cows were culled as soon as possible. If only 1 quarter was infected, the infected quarter was treated by infusion of a commercial disinfectant g for suppressing the excretion of MRSA and milk was not collected from the treated quarter (i.e., drying off). Eventually, a total of 15 cows were culled in this case. In this herd, IMI due to MRSA ended after 5 months as determined by testing all of the cows’ milk and slaughtering or individual quarter treatment (drying off) of infected cows every month. Consequently, MRSA has not been isolated from cows or the environment in this farm to date.
A total of 97 S. aureus isolates were collected from bovine milk, teats, milking equipment, and towels; all of the isolates were distinguished by PFGE into 13 PTs belonging to 3 clusters (Table 2; Fig. 1). All of the isolates belonging to clusters A and B were methicillin-sensitive S. aureus (MSSA), and all but 1 of these isolates were sensitive to all tested antimicrobials (Table 1). Isolates belonging to cluster A were ST705 by MLST and t529 by spa typing (Table 2). Moreover, the following isolates were detected and continuously isolated from bovine milk during the investigation period: capsule serotype 8; 8 exotoxin genes (lukE/lukD, lukM/lukF’-PV, hla, hlb, hld, hlg, sagB, and sagD); and 6 PTSAg genes (sec, seg, sei, sell seln, and tst; Table 2).
Bacterial characteristics of Staphylococcus aureus isolates from a bovine mastitis outbreak in Japan.*
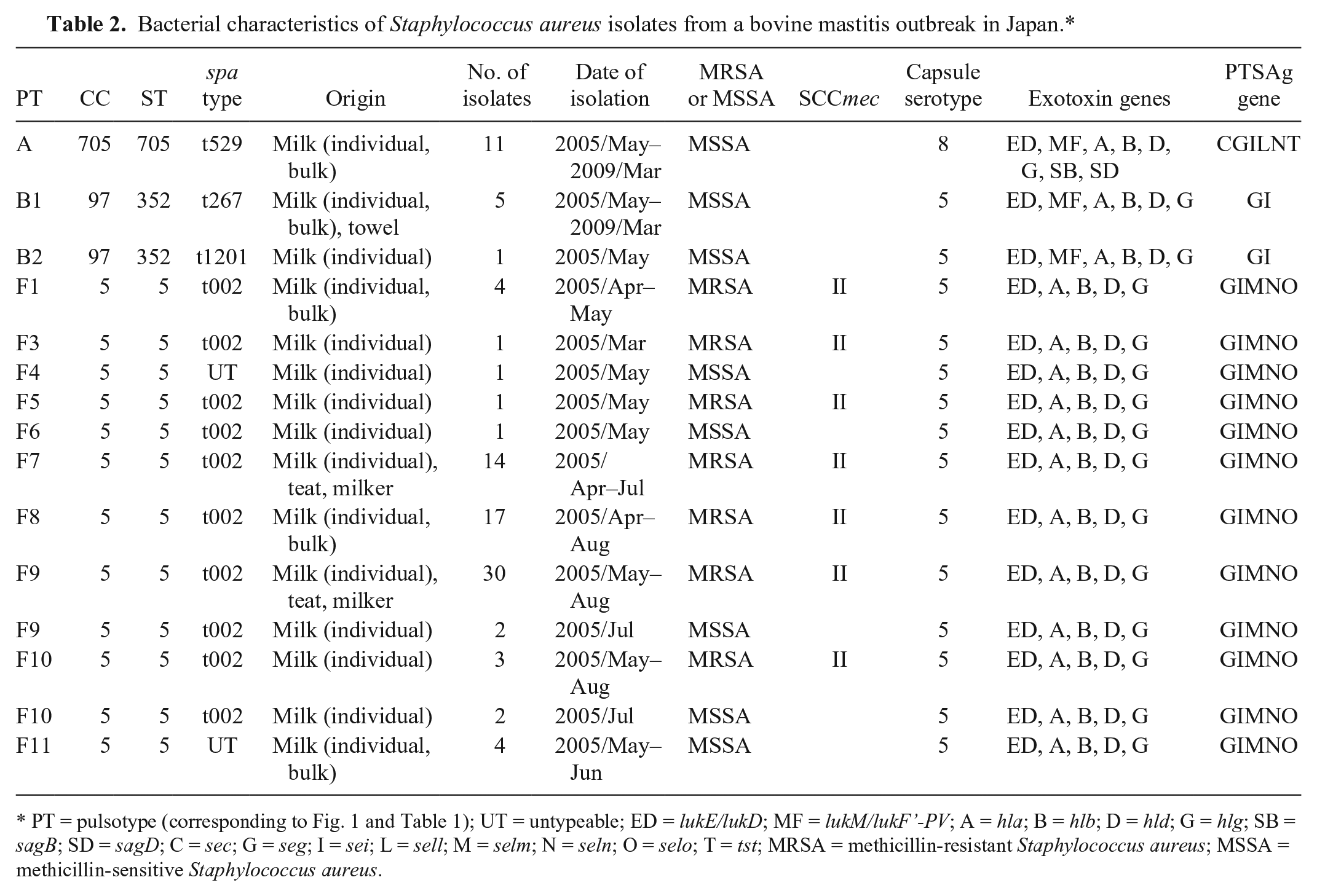
PT = pulsotype (corresponding to Fig. 1 and Table 1); UT = untypeable; ED = lukE/lukD; MF = lukM/lukF’-PV; A = hla; B = hlb; D = hld; G = hlg; SB = sagB; SD = sagD; C = sec; G = seg; I = sei; L = sell; M = selm; N = seln; O = selo; T = tst; MRSA = methicillin-resistant Staphylococcus aureus; MSSA = methicillin-sensitive Staphylococcus aureus.
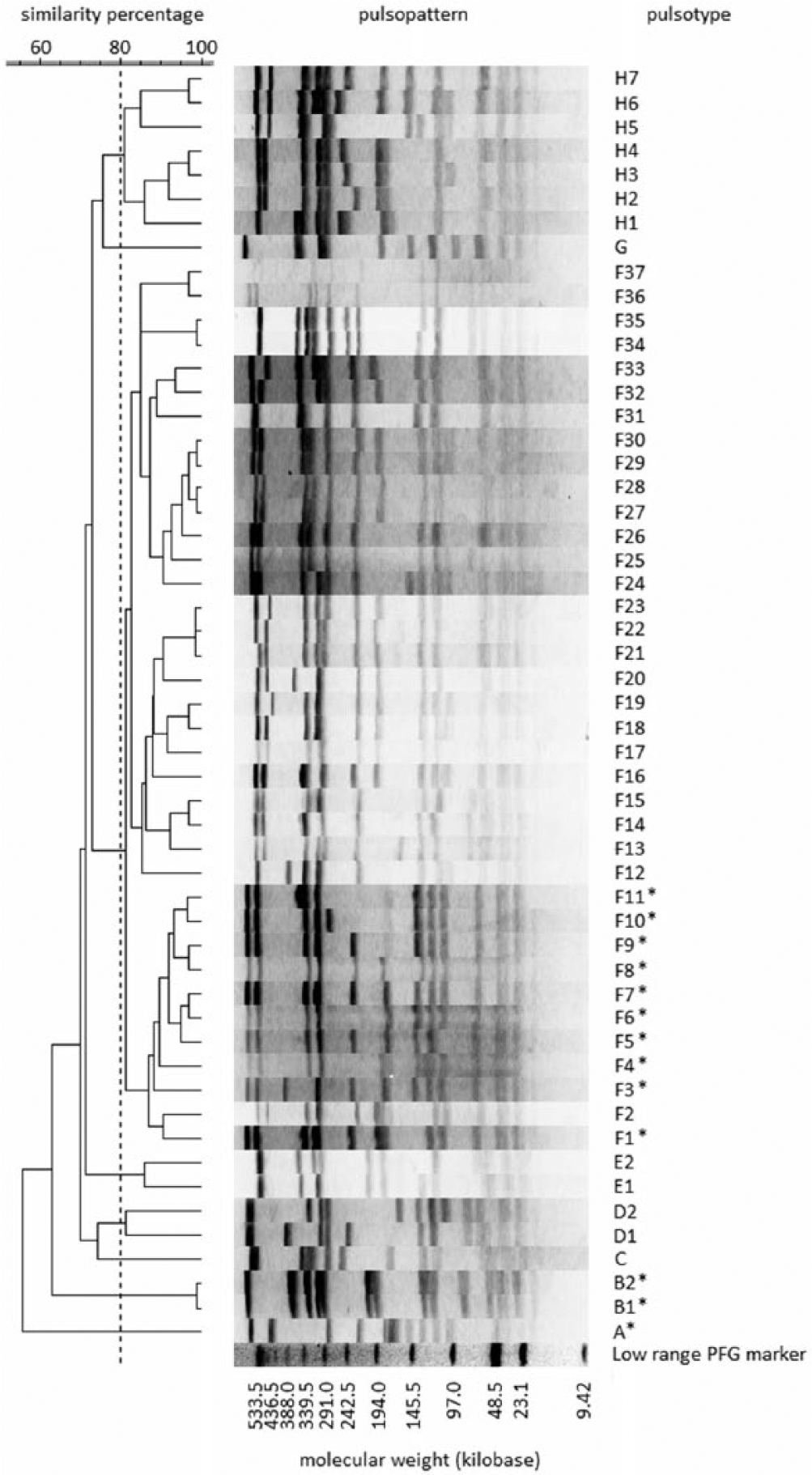

Dendrogram based on pulsopatterns of SmaI-digested total DNA from Staphylococcus aureus isolates in a bovine mastitis outbreak and the methicillin-resistant S. aureus New York/Japan clone from humans in Japan. Pulsotypes in this outbreak are marked with asterisks.
Isolates assigned to cluster B belonged to ST352 by MLST and t267 or t1201 by spa typing (Table 2; Fig. 1). Moreover, the following isolates, obtained from bovine milk and towels during the investigation period, were detected: capsule serotype 5; 6 exotoxin genes (lukE/lukD, lukM/lukF′-PV, hla, hlb, hld, and hlg), and 2 PTSAg genes (seg and sei; Table 2).
All of the MRSA isolates were present in cluster F, and were resistant to 12 antimicrobials (PC, ABPC, DCX, NF, CEZ, KM, EM, OTC, TS, OA, ERFX, and OX) and susceptible to GM, CP, and VCM (Table 1; Fig. 1). All of the MRSA isolates were ST5 by MLST, t002 by spa typing, and SCCmecII (Table 2). These genotyping results suggest that MRSA isolates in this outbreak were HA-MRSA NJC. Moreover, the following isolates, obtained from bovine milk, teats, and milking machines between March and August 2005, were detected: capsule serotype 5; 5 exotoxin genes (lukE/lukD, hla, hlb, hld, and hlg); and 5 PTSAg genes (seg, sei, selm, seln, and selo; Table 2). The concentration of MRSA in milk did not correlate with clinical symptoms (i.e., there were no differences in the concentration between cases with clinical symptoms and cases without clinical symptoms).
Although many isolates belonging to cluster F were MRSA, some of the isolates were MSSA (Tables 1, 2). These MSSA isolates were identified as PT F4, F6, F9, F10, and F11 by PFGE (Table 2; Fig. 1). Some of the isolates within PT F9 and F10 were MRSA. The MSSA isolates belonging to PT F6, F9, and F10 showed the same features in terms of bacterial genotypes and phenotypes regardless of their sensitivity to methicillin. Additionally, MSSA belonging to cluster F showed multidrug resistance against 8 or 9 antimicrobials, but were susceptible to GM, CP, and VCM (Table 1). Interestingly, all of the MSSA strains of PT F9 and F10 were isolated from the milk of cows that had IMI that also yielded MRSA strains of the same PT.
A total of 27 isolates of NJC obtained from humans were also allocated to cluster F by PFGE (Fig. 1). As for IEC, most of the NJC isolates from humans possessed scn (90%), chp (87.5%), and sak (92.5%; Table 3). Moreover, most of these isolates did not possess intact hlb (7.5%), and int was detected in 92.5% (Table 3). As with the NJC isolates from humans, all MRSA other than PT F9 in this outbreak possessed scn, chp, sak, and int, and truncated hlb (Table 3). On the other hand, all MRSA showing PT F9 did not possess IEC genes, and their hlb was intact (Table 3).
Immune-evasion complex and β-hemolysin–converting bacteriophage screening results for methicillin-resistant Staphylococcus aureus (MRSA) isolates in a bovine mastitis outbreak and in New York/Japan clone human isolates in Japan.*
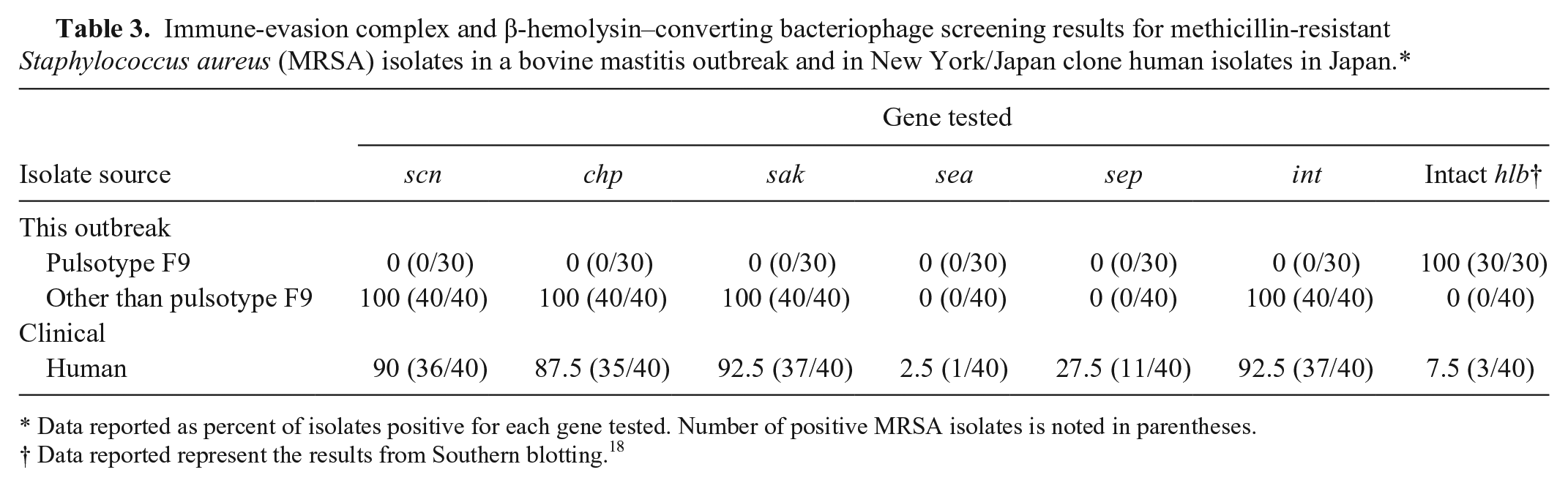
Data reported as percent of isolates positive for each gene tested. Number of positive MRSA isolates is noted in parentheses.
Data reported represent the results from Southern blotting. 18
Discussion
Although bovine mastitis due to MRSA has been observed sporadically throughout the world since 1972, 6 most of S. aureus isolates from bovine milk in Japan were susceptible to all antibiotics, unlike the human isolates. 14 On Japanese dairy farms, S. aureus isolates belonging to CC97 and CC705 are spreading widely as bovine mastitis causal bacteria, but the emergence of MRSA has fortunately not been confirmed in these CCs as of now.14,16 Moreover, S. aureus belonging to CC133 has been consistently isolated from bovine IMI cases in Europe, 16 but MRSA belonging to CC133 has not been reported to date. Most IMI cases caused by MSSA belonging to CC97 and CC705 are asymptomatic and are characterized by persistent infection, and thus they are difficult to eliminate from herds. 17 The MSSA of ST352 and ST705 in the present case were also consistently isolated from bovine milk without obvious mastitic symptoms throughout the investigation period as has been conventionally reported. 17
The reason that MSSA belonging to CC97 and CC705 can tenaciously infect the bovine udder may be that these strains possess a toxin with strong cytotoxicity against bovine immune cells (i.e., LukM/LukF′-PV). 1 The S. aureus isolates from this outbreak that were ST352 and ST705 possessed LukM/LukF′-PV, as well. As LukM/LukF′-PV can immediately remove bovine immune cells, these strains are likely to be able to easily maintain infection in the bovine udder. Staphylococcal exotoxins other than LukM/LukF′-PV also show cytotoxicity to bovine neutrophils, and although they are much weaker than LukM/LukF′-PV, these exotoxins may also lead to some pathogenicity. 1 Moreover, all of the isolates in this outbreak possessed some PTSAg genes. These PTSAgs are important proinflammatory factors much like lipoteichoic acid, and the superantigen activity that PTSAgs show disturbs the immune system of the host. 7 Staphylococcal enterotoxin C, which is one of the major PTSAgs among bovine isolates, is suggested to facilitate immunosuppression to maintain persistent infection in bovine mammary glands. 11 The exotoxin gene and PTSAg gene profiles of MRSA isolates in this outbreak were very similar to those of MRSA isolates from human samples in Japan. 14 In other words, seg, sei, selm, seln, selo, lukE/lukD, hla, hld, and hlg were possessed by the majority of both types of isolates. 14 Moreover, the resistance pattern of MRSA isolates in this outbreak was also similar to that of the MRSA isolates from human samples. Although the majority of both types of MRSA isolates were resistant to β-lactams, cephems, and macrolides, many of them were susceptible to CP and VCM. 14 Such a genetic and antimicrobial resistance profile suggests a close relation between the 2 types of isolates.
Bovine IMI cases in which MRSA was isolated are very rare. 16 One reason that bovine-associated S. aureus remains susceptible to antibiotics may be that dairy farmers and veterinarians are discreet about the use of antibiotics that would result in shipment restrictions for milk. However, reports of bovine IMI by MRSA have increased rapidly since the 2000s. In particular, bovine mastitis caused by the MRSA exhibiting ST398 has conspicuously increased in Europe since 2006. 19 S. aureus of ST398, which emerged in pigs in 2005, is rapidly spreading around the world as LASA and has been isolated from livestock and humans in Europe, South America, and China (http://saureus.mlst.net/). 19 The combination of toxin genes that S. aureus ST398 possesses differs from the combinations present in bovine-associated S. aureus (CC97, CC705, and CC133), and lukM/lukF′-PV are not present in S. aureus ST398. 29 In nature, S. aureus ST398 may not be suitable for infection in cows. Moreover, bovine mastitis caused by S. aureus ST398 has not yet been reported in Japan.
Although the author has observed bovine mastitis caused by MRSA ST8 and ST461 and has isolated MRSA ST5 and ST89 from bovine milk in Japan, 16 bovine IMI caused by MRSA is very rare in Japanese dairy herds. Because the genotypes of these MRSA are common among human isolates, it is speculated that they were brought into dairy herds by human carriers. The MRSA isolates in this outbreak and the NJC isolated from human samples also showed similar pulsopatterns. Additionally, the MRSA isolates (other than PT F9) from this outbreak and the NJC isolates from human samples showed common genetic characteristics in IEC and hlb. Most clinical MRSA ST5 from humans possess IEC genes and truncated hlb, 18 and the present study obtained comparable results. Therefore, it can be speculated that the MRSA isolates (other than PT F9) in this outbreak were introduced by humans. On the other hand, most LA-MRSA ST5 strains did not possess IEC genes, and their hlb are intact, 17 as in the MRSA of PT F9 in the current study. Although MRSA other than PT F9 and MRSA PT F9 in this outbreak showed different genetic characteristics in IEC and hlb, their pulsopatterns were very similar. Therefore, both seem to have been derived from the same origin. Changes in these genetic characteristics (i.e., the omission of IEC genes and bacteriophages in hlb) may occur easily. If this speculation is correct, it may be that the NJC was in the process of adapting to livestock colonization. The results suggest that MRSA PT F9 was most likely becoming the predominant strain in this outbreak.
In this study, bovine mastitis and IMI caused by the NJC showed obvious mastitic symptoms in only the first 3 infected cows, but the remaining infected cows were asymptomatic. Moreover, no correlation between the concentration of MRSA in milk and disease severity was confirmed. Such symptomless infections are similar to those in common bovine mastitis and IMI caused by bovine-associated S. aureus, 17 and these results may suggest that mastitis and IMI cases caused by NJC seem non–life-threatening. Therefore, the efficient detection of asymptomatic carrier cows, as occurred in this outbreak, is believed to be the most important measure for eliminating MRSA from dairy herds. The PTs and genotypes of the MRSA isolates from milking machines and towels used in milking corresponded to those of the MRSA isolates from milk samples, therefore the process of milking is believed to be the main route of the spread of MRSA, as with MSSA. 17 The main infection sources of MRSA are also believed to be the milk and the bodies of carrier cows, as with MSSA. 17
Finally, the presence of MSSA and MRSA with identical genotypes was confirmed in this outbreak. Furthermore, these MSSA and MRSA genotypes were isolated from the same cows. This phenomenon suggests that the exchange of SCCmec occurred easily among S. aureus strains in the cows’ udders. This suggestion and the indication that the omission of IEC genes and bacteriophage in hlb may also occur provide a basis for development of LA-MRSA strains though separate or combined pathways. Although these hypotheses may be overly speculative, measures to prevent the invasion of MRSA into dairy herds in order to prevent emergence of bovine-adapted MRSA are clearly important.
Footnotes
Acknowledgements
I thank the staff of the Federation of Agricultural Mutual Aid Associations and the Livestock Hygiene Service Center for their help with the collection of bovine milk samples and bacterial isolates.
Author’s contribution
E Hata contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Eiken Chemical Co. Ltd., Tokyo, Japan.
b.
PrimeSTAR GXL DNA polymerase, Takara-Bio Co. Ltd., Kusatsu, Japan.
c.
BioNumerics software version 5.10, Infocom Co. Ltd., Tokyo, Japan.
d.
InstaGene matrix, Bio-Rad Laboratories, Hercules, CA.
e.
Denka Seiken Co. Ltd., Tokyo, Japan.
f.
Difco Laboratories Inc., Sparks, MD
g.
2% isodine for animal, Meiji Seika Pharma Co. Ltd., Tokyo, Japan.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received the following financial support for the research, authorship, and/or publication of this article: Grants were provided by the Ministry of Agriculture, Forestry and Fisheries of Japan and the National Agriculture and Food Research Organization of Japan.
