Abstract
Our study describes a newly developed mini-array test for the rapid detection of poxviruses in animals and humans. The method is based on detection that combines target nucleic acid amplification by polymerase chain reaction and specific hybridization, using enzyme-linked antibodies, allowing identification of zoonotic orthopoxviruses and parapoxviruses in animal and human biological samples. With 100% specificity, the test rules out the possibility of cross-reactions with viral agents causing look-alike diseases. The assay was employed in the field to investigate the causes of several outbreaks of a malignant proliferative skin disease that affected domestic ruminants in Sicily during 2011–2014. Due to specific aspects of the lesions, the animals were clinically diagnosed with papillomatosis. The mini-array test allowed the identification of coinfections caused by more than 1 viral species belonging to the Parapoxvirus and Orthopoxvirus genera, either in goats or in cattle. Our study suggests that the so-called “papillomatosis” can be the result of multiple infections with epitheliotropic viruses, including zoonotic poxviruses that cannot be properly identified with classical diagnostic techniques.
The families Papillomaviridae and Poxviridae include a number of viral species affecting different mammalians. The Papillomaviridae family now comprises 32 genera,20,21 several of which infect ruminants. In ruminant species, papillomaviruses induce hyperplastic benign lesions of both cutaneous and mucosal epithelia that generally regress, but may also occasionally persist with the possibility to evolve into cancer in the presence of environmental carcinogenic cofactors. In the Poxviridae family, the genera Parapoxvirus (PPV) and Orthopoxvirus (OPV) include viruses that cause skin lesions in domestic and wild ruminants as well as in humans. The parapoxviruses Orf virus (ORFV), Bovine papular stomatitis virus (BPSV), Pseudocowpox virus (PCPV), and the orthopoxviruses Vaccinia virus (VACV) and Cowpox virus (CPXV) infect via damaged skin and give rise to proliferative and/or ulcerative lesions of the skin and mucosae.
Papillomavirus and poxvirus infections recognize the same risk factors caused by highly resistant agents, and they do not necessarily require direct contact between infected and susceptible host to be transmitted. The shed scabs, or necrotic skin or mucosal tissue, contain high titers of stable viruses that can remain infectious long enough to sustain indirect transmission. Both papillomaviruses and poxviruses infections in animals are usually diagnosed symptomatically and, only on occasion, a laboratory confirmation is requested. In fact, the clinical signs are considered typical; however, the lesions can be easily confused with those shown by other diseases of ruminants such as those caused by Foot-and-mouth disease virus, Bluetongue virus, Sheeppox virus, and CPXV. Early recognition of the etiologic agents allows proper control measures and disease management strategies, to prevent disease spread among animals and humans in the case of zoonotic viruses. At present, there are no routine diagnostic tests for poxviruses, and this can lead to the possibility of misdiagnosis with other epitheliotropic pathogens. 14 Poxvirus zoonoses fall into the category of neglected zoonoses, and their incidence in animals and humans is often unknown or greatly underestimated. In humans, underreporting leads to underestimating the true number of disability-adjusted life years, which can be averted by an effective control. The availability of highly specific and sensitive diagnostic assays is considered critical to perform an accurate differential diagnosis and to understand the real incidence and prevalence of zoonotic poxviruses among mammals. 14
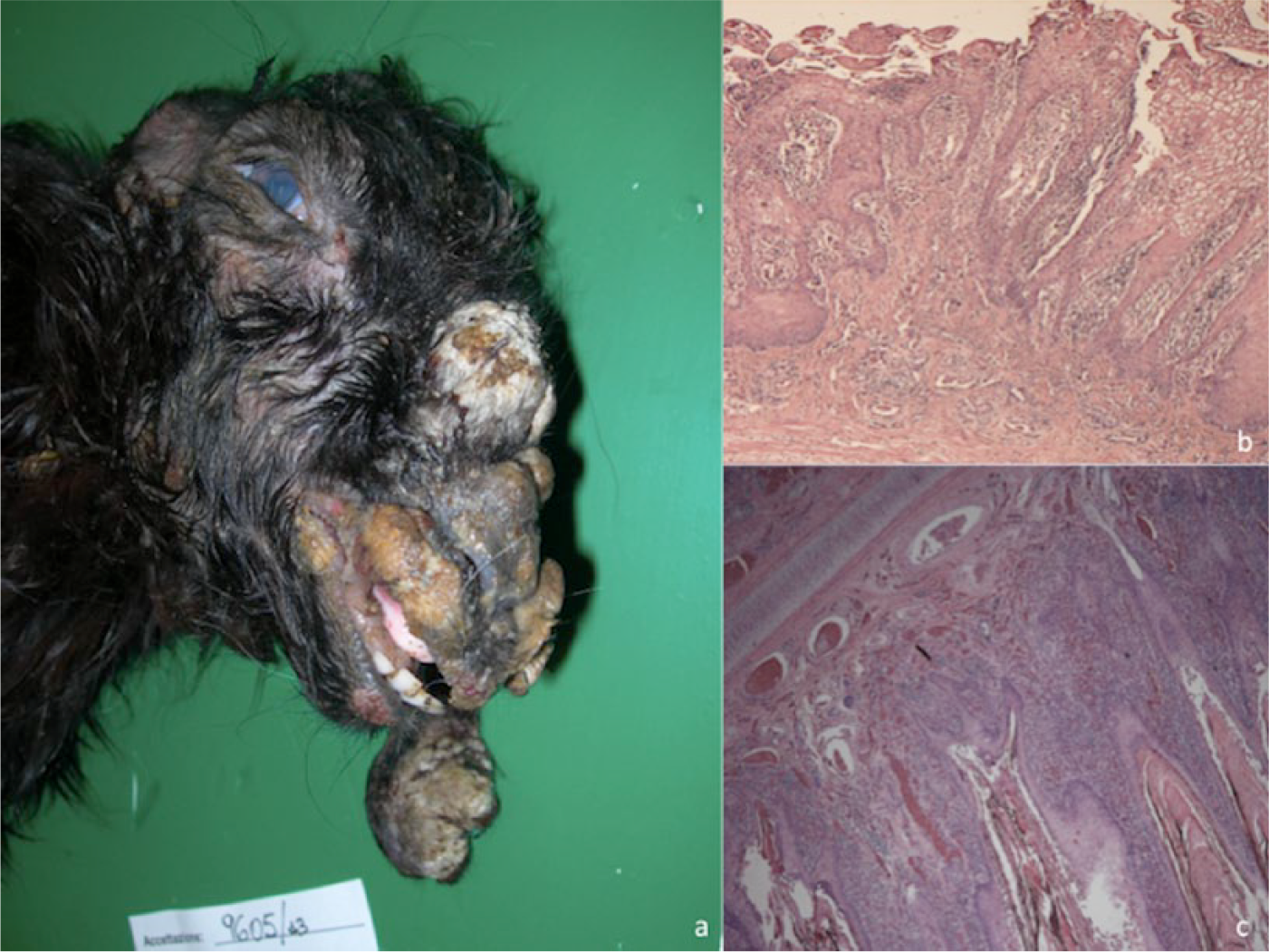
The aim of our study was to identify, through classical and innovative diagnostic methods, the viruses responsible for a number of outbreaks of a disease characterized by massive cutaneous lesions in domestic ruminants in Sicily. The clinical diagnosis was generically defined as papillomatosis because the animals were showing multiple tumor-like lesions disseminated throughout the entire body (Fig. 1a). In many cases, the symptoms were characteristic of a systemic disease, with anorexia, lethargy, and death by starvation in the most severely affected animals. Pathological samples (Table 1) consisting of scabs and proliferative lesions, were submitted for standard histological examinations and negative-staining electron microscopy (EM). 12 Histology mainly identified hyperkeratosis and acanthosis of the epidermis and proliferation of the fibroblasts, consistent with fibropapillomatosis, in cows, while degenerative changes in the stratus spinosum, epidermal hyperplasia, hyperkeratosis, ballooning and degeneration of keratinocytes with eosinophilic inclusion bodies, were mainly evidenced in the lesions of sheep and goats (Fig. 1b, 1c). Negative-staining EM lead to the identification of Papillomavirus particles in the samples collected from diseased cows and Parapoxvirus virions in affected small ruminants.
Epitheliotropic viruses detected in the pathological lesions of diseased animals. The sequences of orthopoxviruses detected in cows and goats were submitted to GenBank, and their accession numbers are indicated.*

DPV = Deltapapillomavirus; XPV = Xipapillomavirus; EPV = Epsilonpapillomavirus; BPSV = Bovine papular stomatitis virus; PCPV = Pseudocowpox virus; ORFV = Orf virus; – = negative result.
Viral DNA was extracted from the pathological specimens a and analyzed by rolling circle amplifications b followed by restriction enzyme digestions, as previously described, 19 to characterize the papillomavirus circular DNA. Different sets of primers3,4 have been used to further characterize the identified Papillomaviridae types. The DNA samples, purified from the lesions of all the diseased ruminants, were further analyzed using a newly developed mini-array assay developed within the FP7 POC4PETS project (http://www.poc4pets-fp7.eu/project/default.aspx). This assay allows the identification of different zoonotic poxviruses, belonging to the genera Parapoxvirus and Orthopoxvirus, within the same sample. The test is based on a detection approach that combines target nucleic acid amplification by polymerase chain reaction (PCR) and specific hybridization probes spotted on a mini-array. The assay, versus standard PCRs, showed 100% specificity, 87% sensitivity, and a limit of detection of 1 fg of viral DNA. The straightforward interpretation of the mini-array through 1-step multiple detection of the hybrids, which yields colored spots on the mini-array that are visible to the naked eye, is a distinct advantage over other molecular diagnostic tools.
Briefly, for multiple detection of OPVs and PPVs, 2 target genes (HA, encoding the hemagglutinin, and B2L, encoding the major envelope protein, respectively) were selected. The cytochrome C oxidase subunit I (COX I) gene, of different animal species including sheep, goat, cow, horse, dog, cat, and human, was also selected as internal control. Oligonucleotide primers for targets and probe amplifications were designed using the Primer3Plus c web tool 25 and OligoAnalyzer d 3.1 software. To allow multiplex amplifications for each target gene, the best primers candidates for probes and targets have been preliminarily defined as well as the optimal PCR conditions (based on amplicon yield, specificity, sensibility, no false positive). Specific probes of OPV, PPV, and positive control were obtained by PCR amplification with DNA polymerase. e Mini-arrays were then generated by dotting the double-stranded DNA probes onto a positively charged nylon membrane f using an arrayer g and denaturing the DNA probes by incubation of the membrane in NaOH solution. The membrane was pre-hybridized in hybridization buffer containing 1% skim milk and then dried. Targets are thus labeled during multiplex PCR by incorporating digoxigenin-11-dUTP h with DNA polymerase. e The subsequent detection assay consists of a target-probe hybridization step followed by a colorimetric revelation of the hybrids using enzyme-conjugated anti-digoxigenin antibody, i producing a dark blue precipitate in the presence of appropriate substrate (Fig. 2). The mini-array analyses, performed on the DNA purified from all the diseased ruminants, lead us to confirm that all of the sheep and goats were infected with PPV. Seven out of 12 cows, previously found positive for different Bovine papillomavirus genotypes using rolling circle amplification analysis, were also found coinfected with parapoxviruses (Table 1). Surprisingly, 2 goats and 2 cows from Messina Province and Ragusa Province (Italy), respectively, showed a coinfection with Parapoxvirus and Orthopoxvirus (Table 1).
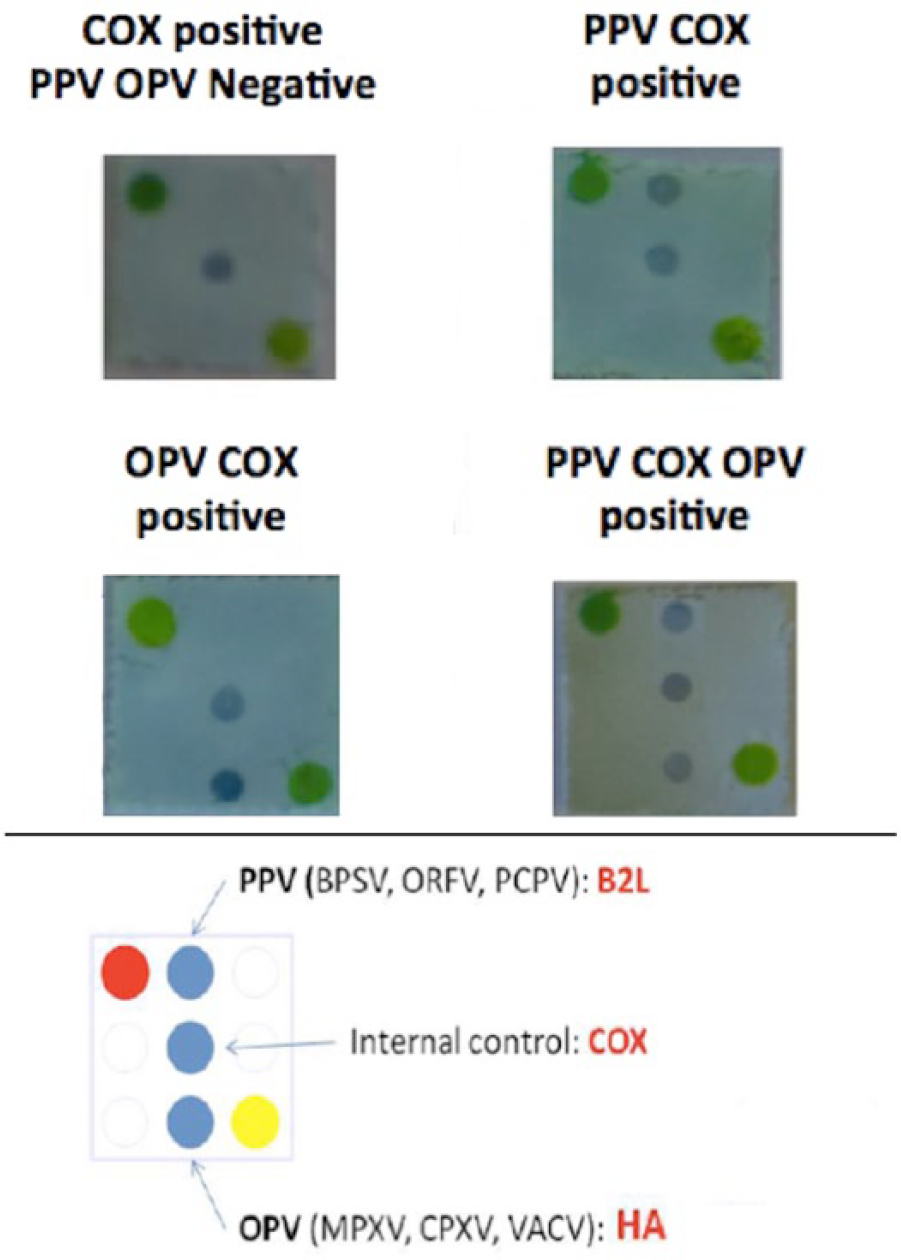
Mini-array results allowing the simultaneous detection of orthopoxviruses and parapoxviruses. PPV = genus Parapoxvirus; OPV = genus Orthopoxvirus; BPSV = Bovine papular stomatitis virus; ORFV = Orf virus; PCPV = Pseudocowpox virus; MPXV = Monkeypox virus; CPXV = Cowpox virus; VACV = Vaccinia virus.
To further characterize the poxviruses identified by the mini-array test, the viral DNA was subsequently amplified j by PPPI and PPPIV primers targeting the Parapoxvirus B2L gene as previously described. 15 PCR amplification of the OPV DNA was performed using the primers rpo f2 (5′-ATGTCGAGCTTTGTTACCAAT-3′) and rpo r2 (5′-TGCCATAGTATTCGTGTTTATACT-3′) l targeting the highly conserved Rpo 18 gene that encodes the DNA-dependent RNA polymerase of OPVs. 2 Thermal cycler conditions were set with an initial denaturation at 94°C for 5 min, followed by 40 cycles of denaturation at 94°C for 30 sec, annealing at 53°C for 30 sec and extension at 72°C for 45 sec, with a final extension at 72°C for 10 min. The PCR, performed on the mini-array–positive samples, lead to the amplification of specific fragments of 594 bp and 526 bp, respectively, for the B2L gene of PPVs and Rpo 18 gene of OPVs. The amplified products were subsequently purified k and sequenced. l Sequence analysis of the PCR products lead us to identify BPSV in 3 cows and PCPV in 4 animals, while sheep and goats were shown to be all infected with ORFV. The Rpo 18 nucleotide sequences were aligned using ClustalW m with previously published OPV sequences from GenBank, showing an identity ranging from 97.3% to 100% among the 4 OPVs detected in the Sicilian outbreaks. The alignments performed, with published OPV sequences, showed a nucleotide identity ranging from 99.6% to 100% between strains 9605-425, 14647-429, and 33225-498, and CPXV strain UK2000_K2984 (GenBank accession HQ420900). At the amino acid level, a complete identity was detected with multiple CPXV and VACV strains. A similarity of 98.1% and 98.8%, at nucleotide and amino acid level, respectively, was detected between strain 32063-501 and CPXV EleGri07/01 (id. KC813507). Phylogenetic trees were generated by the neighbor-joining method 21 with 1,000 bootstrap replicates 10 using the Tamura 3-parameter model.n,22,23 These analyses showed that the 4 OPV strains cluster into 2 distinct groups (Fig. 3).
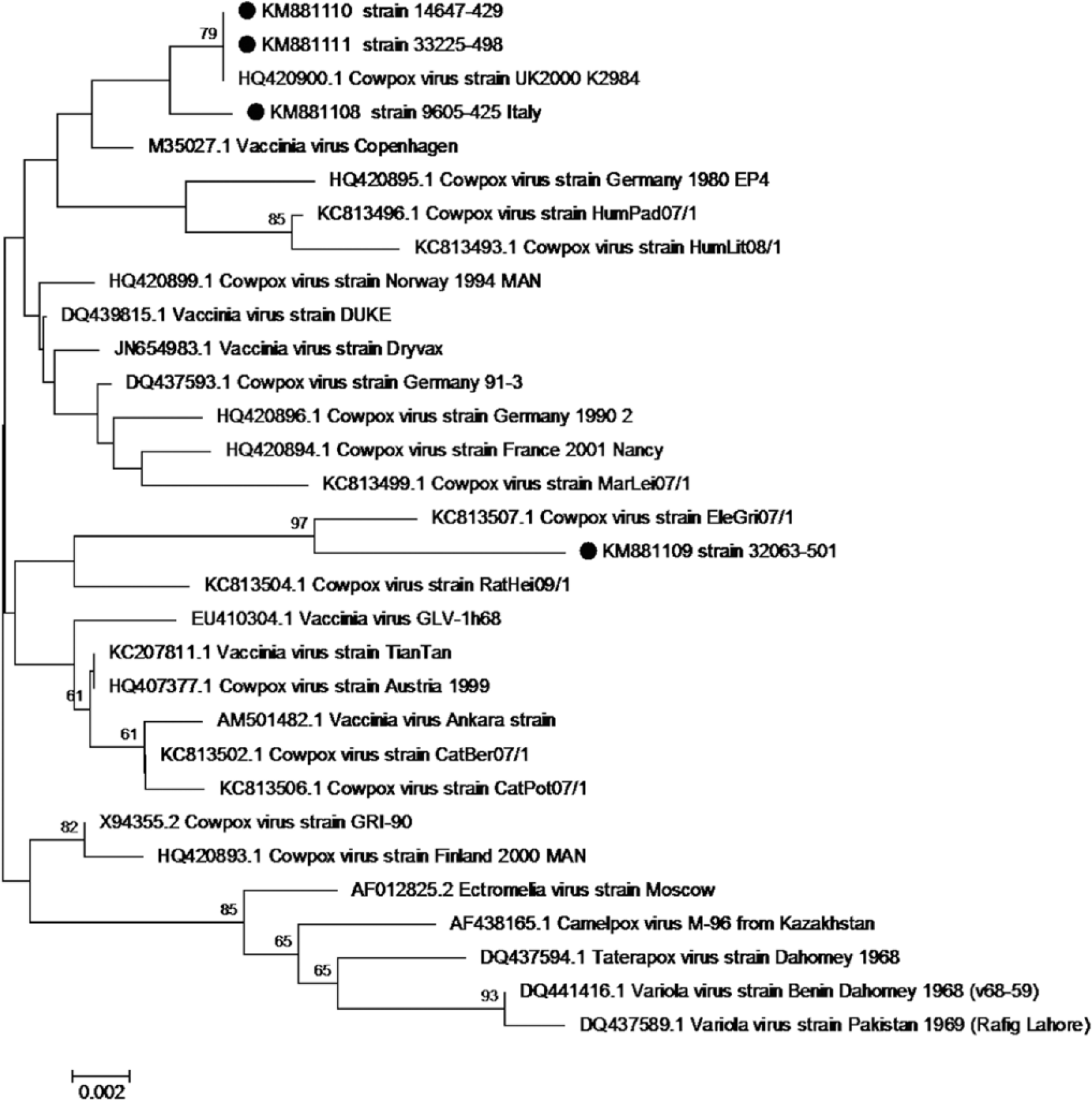
Phylogenetic tree generated from alignment of complete orthopoxvirus (OPV) Rpo 18 gene sequence using the neighbor-joining method and the Tamura 3-parameter model of nucleotide substitution, implemented in MEGA 6. n The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) is shown above the branches (cutoff 60%). Dots indicate the viral strains described in our study.
To determine the viability of viruses in samples, 60% confluent Vero (African green monkey kidney cell) monolayers were inoculated with 10-fold dilutions of sonicated tissue extract. 6 Infected cells were incubated at 37°C in a 5% CO2 atmosphere in Dulbecco modified Eagle medium. Cell infection was monitored microscopically by observation of OPV-specific cytopathic effect (CPE). In order to quantify the OPV DNA in the pathological samples, a real-time SYBR Green PCR was performed.j,o The absolute quantification of the viral DNA was obtained by plotting 10-fold dilutions (1.7 × 101 to 1.7 × 105 copies/µL) of pUC19 plasmid containing the cloned target fragment. No CPE was observed in inoculated cell culture, while the quantitative real-time PCR revealed an OPV load ranging from 101 to 103 DNA copies/µL, suggesting that DNA-positive samples might not contain viable virus.
Our data leads us to demonstrate that classical techniques, such as histology and EM, may not be sufficiently effective and reliable for the diagnosis. The mini-array assay used in our study revealed that some of the animals affected by severe proliferative lesions, clinically defined as papillomas, were in fact coinfected with zoonotic PCPV, BPSV, and OPV. It is important to emphasize that zoonotic poxviruses share risk factors and clinical features with other life-threatening diseases in humans 18 ; for this reason, rapid and reliable diagnostic methods, like the one used in our study, have the potential to allow the rapid assessment of potentially fatal infections in humans. With the aid of this new method, it was possible to demonstrate that 2 goats and 2 cows, from different farms, were coinfected with different epitheliotropic viruses. In particular, the cattle identified as 14647-429 and 33225-498 that showed the presence of papillomavirus particles by EM were also coinfected with zoonotic poxviruses (BPSV and PCPV, respectively, and a virus belonging to genus Orthopoxvirus). A previous study reported coinfections of Parapoxvirus and Orthopoxvirus in cattle in Brazil. 9 The description of the clinical cases seen in Brazil were consistent with those found in the Italian outbreaks. Despite the large number of diseased animals, the authors 9 were able to demonstrate the simultaneous presence of Parapoxvirus and Orthopoxvirus particles by EM in the pathological sample of only 1 animal. As in our study, no CPE was detected in inoculated cell cultures. Further trials, using alternative cell substrates, will be necessary to rule out the possibly that the pathological samples contained only DNA remnants or viral particles with compromised integrity by either the animal immune system or by external factors during the sample collection or processing. 11
The results of our study indicate that tissue culture isolation is less sensitive than molecular methods to detect zoonotic poxvirus in coinfected animals while the mini-array method shows to be reliable, robust, and suitable for areas where OPV and PPV cocirculate. Coinfections with ORFV-like parapoxvirus and VACV have been previously reported in dairy cows 9 while we identified the simultaneous presence of ORFV and OPV in 2 goats. In a 2010 study, coinfections with VACV and PCPV in a rural worker and in dairy cows were described. 1 Our study reinforces the fact that coinfections with OPV and PPVs may occur in cattle not only involving PCPV but also BPSV. The high sequence identity and the circulation of CPXV in Italy in different animal species5–7 lead us to conclude that the OPV DNA detected in domestic ruminants might be CPXV. This virus used to be enzootic in cattle; however, such infections have not been diagnosed over the last decades, while individual cases of cowpox have been repeatedly found in cats or exotic zoo animals that were responsible for human transmission.8,13,16,17,24,26 It is not possible to conclude that CPXV was the causal agent of the lesions reported in the domestic ruminants in our study; in fact, its low genomic load can be the result of interference caused by multiple viral agents replicating at the same time in the same target cell. On the other hand, the presence of CPXV DNA in the pathological samples of the domestic ruminants lead us to speculate that the virus may use these animal species as reservoirs to spread and persist in the environment through the scab material produced by other epitheliotropic viruses.
Footnotes
Authors’ contributions
A Scagliarini contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved. G Casà, F Savini, M Morent, R Puleio, C Buttaci, V Cannella, G Purpari, and P Di Marco contributed to acquisition, analysis, and interpretation of data, and gave final approval. B Trentin contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; critically revised the manuscript; and gave final approval. L Gallina contributed to conception and design of the study, and gave final approval. A Lavazza contributed to acquisition, analysis, and interpretation of data; critically revised the manuscript; and gave final approval. D Piquemal contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and gave final approval. A Guercio contributed to conception and design of the study; critically revised the manuscript; and gave final approval.
a.
NucleoSpin tissue kit, Macherey Nagel GmbH & Co. KG, Düren, Germany.
b.
Templiphi 100, GE Healthcare Europe GmbH, Milan, Italy.
e.
FastStart Taq DNA polymerase, Roche Diagnostics GmbH, Mannheim, Germany.
f.
Amersham Hybond-N, GE Healthcare Europe GmbH, Milan, Italy.
g.
Biomek 2000, Beckman Coulter s.r.l., Milan, Italy.
h.
Digoxigenin-11-dUTP, Roche Diagnostics GmbH, Mannheim, Germany.
i.
Anti-digoxigenin-POD, Fab fragments; Roche Diagnostics GmbH, Mannheim, Germany.
j.
PCR reagents, Qiagen GmbH, Hilden, Germany.
k.
High Pure PCR product purification kit, Roche Molecular Diagnostics, Mannheim, Germany.
l.
ABI PRISM 3730, Applied Biosystems, Foster City, CA.
m.
DNASTAR Inc., Madison WI.
n.
MEGA 6 software, Center for Evolutionary Medicine and Informatics, Tempe, AZ.
o.
Rotor Gene 3000, Corbett Research, Sydney, New South Wales, Australia.
Declaration of conflicting interests
The author(s) declared no potential conflict of interest with respect to the research, authorship, and publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research leading to these results has received funding from the Italian Ministry of Health Ricerca corrente (SI 14-11) and the European Union, Seventh Framework Programme, FP7/2007-2013 (grant agreement 315653).
