Abstract
The swine industry currently lacks validated antemortem methods of detecting baseline herd prevalence of Mycoplasma hyopneumoniae. The focus of our study was to evaluate alternative antemortem detection techniques and to determine baseline litter prevalence in preweaned pig populations utilizing the selected technique and a real-time polymerase chain reaction (qPCR) assay. Preliminary data was analyzed on weaned piglets with evidence of respiratory disease (n = 32). Five sample types (antemortem nasal swab, tracheobronchial mucus, postmortem deep airway swab, bronchoalveolar lavage, and lung tissue) were collected from each pig. Individual samples were tested for M. hyopneumoniae using qPCR. Compared to nasal swabs, tracheobronchial mucus demonstrated higher test sensitivity (P < 0.0001). Tracheobronchial mucus was collected from apparently healthy preweaned piglets (n = 1,759; sow farms 1–3) and preweaned piglets exhibiting signs of respiratory disease (n = 32; sow farm 4), ranging in age from 12 to 25 days. Samples from sow farms 1–3 were pooled into 2 groups of 5 per litter (n = 360 pools from 180 litters), and qPCR was utilized to analyze each pool. A qPCR-positive result, threshold cycle <37, from either pool designated the litter positive for M. hyopneumoniae. Two out of 180 litters revealed a positive result (1.1%). Individual qPCR assays were run on the samples collected from sow farm 4. Five out of 30 samples revealed a positive result (16.7%). Tracheobronchial mucus collection in combination with qPCR is a sensitive antemortem sampling technique that can be used to estimate the prevalence of M. hyopneumoniae in preweaned pigs, thus providing insight into the infection dynamics across the entire farrow-to-finish process.
Mycoplasma hyopneumoniae, the causative agent for porcine enzootic pneumonia (EP) and a key component in the porcine respiratory disease complex, remains a major threat to the world’s swine industry.8,15 As an example, M. hyopneumoniae is considered to be one of the top sources of economic losses in U.S. swine production swine production. 4 Although vaccination protocols are widely implemented in swine herds, mycoplasmal pneumonia remains a threat to swine production.3,13 However, it is apparent that not all herds are affected equally, and the difference has been linked to the prevalence of M. hyopneumoniae in weaned pig populations. 6 Research suggests that the main risk factor for the development of disease is vertical transmission from sow to piglet during lactation and that the severity of the disease at slaughter may be predicted by the initial prevalence at weaning. 6
A 2013 study 16 suggested vaccination as the most cost-effective control measure for EP. However, previous work12,18 indicated that a concurrent Porcine reproductive and respiratory syndrome virus (PRRSV) infection, either natural or from a modified live vaccine, reduces the efficacy of a M. hyopneumoniae vaccine. Because PRRSV is endemic in the United States, such data support the necessity for effective antimicrobial treatment options. There is research to demonstrate that antimicrobial therapy is an effective method of control of clinical signs associated with EP. 2 Studies suggested reducing prevalence in the breeding herds which in turn may reduce spread of disease to piglets. 8 However, antemortem estimation of herd prevalence and the ability to obtain herd-specific isolates for sensitivity testing is required to establish baseline data to support future research. In the recent past, several studies comparing antemortem diagnostic techniques to detect M. hyopneumoniae have been published.5,9,10 Comparison of lung tissue and bronchoalveolar lavage fluid (BALF) of M. hyopneumoniae–affected piglets revealed a higher sensitivity for lung tissue. 10 In another study, 4 different antemortem sampling techniques, namely tracheobronchial mucus collection (TBMC), BALF, nasal swabs, and tonsil scrapings were compared for their ability to detect M. hyopneumoniae–infected piglets using real-time polymerase chain reaction (qPCR). 5 The study revealed a 3.5 times higher sensitivity for TBMC samples as compared to the other sample types. 5 However, these studies have each focused on individual analysis of collected samples, which can be quite expensive for screening large swine operations with low M. hyopneumoniae prevalences.
In the present study, 2 diagnostic investigations are reported. The first investigation identified a suitable protocol for both sample collection and diagnostic testing for M. hyopneumoniae, whereas the second investigation applied the selected protocol to 4 sow farms across the state of Iowa to assess litter prevalence of M. hyopneumoniae in the preweaned piglet population.
Thirty-two weaned pigs, from a commercial sow farm in Iowa, showing clinical signs of respiratory disease were submitted for diagnostic evaluation. a The TBMC method was utilized in our study as previously described5,9 with some modifications. 19 Nasal swabs were collected, and TBMC was performed antemortem, while deep airway swabs (down to the tracheobronchial split), BALF, and lung tissue, from the cranial lung lobes, were collected postmortem from each pig. All samples were stored on ice after collection until analysis. a In an effort to identify the optimal sample site and DNA extraction method for detection of M. hyopneumoniae, a qPCR assay, using 3 separate extraction methods (viral, total nucleic acid [TNA], and high volume), was run on each sample (32 pigs × 5 samples × 3 extraction methods), for a total of 480 qPCR reactions. The viral and high volume extractions were performed using an RNA/DNA nucleic acid commercial extraction kit b according to the manufacturer’s recommendations. The third extraction was a total nucleic acid extraction that was performed using a different commercial kit c following the manufacturer’s recommended protocol for oral fluids. Each extraction method was carried out on a particle processor d using the processor program specific for each extraction method. e Following extraction, the nucleic acid templates were used in a qPCR f reaction according to manufacturer’s recommendations. g Based on the initial results, pooling potential was assessed for the TBMC samples. Nine individual samples, all of which had originally tested positive, were pooled at a 1:5 and a 1:10 dilution, where 1 known positive sample was pooled with 4 or 9 known negative samples, respectively. The pooled threshold cycle (Ct) values using the 3 extraction methods were compared with the original individual Ct values.
For the second investigation, minimum inclusion criteria were as follows: no use of any antibiotics effective against Mycoplasma species in either piglets or sows for at least 3 weeks prior to sampling, and the presence of at least 20 gilt litters, 20 litters from parity 2 or parity 3 sows, and 20 litters from sows currently in their fourth parity or greater on the farm in a single weaning group. Lactating sows from each of the nonclinical case farms (sow farms 1–3) were assigned a group number based on their current parity (gilts, group 1; parity 2 or 3 sows, group 2; and fourth parity or above sows, group 3). Up to 10 piglets from 20 litters per group from the 3 sow farms were sampled using the TBMC technique for a possible total of 600 piglets per farm sampled (10 piglets × 20 litters × 3 groups). In our study, samples were pooled in groups of 5 per litter so that all litters had 2 pools of samples and the litter was designated the experimental unit. Ten piglets were sampled at random if >10 piglets per litter were present. If <10 piglets per litter were present, all piglets in the litter were sampled, and the first 5 samples in the litter were pooled together and the remaining samples were then pooled. If >20 eligible litters per group were present on the farm, litters were sampled at random. All sampled piglets were between 12 and 25 days of age. From sow farm 4, 32 individual piglets exhibiting signs of respiratory disease were chosen for sample collection with the TBMC technique between 13 and 20 days of age. All samples were tested individually.
Each sample or sample pool was analyzed using a validated M. hyopneumoniae p183 real-time PCR.a,17 Briefly, nucleic acid was extracted from TBMC using a high-volume extraction technique as described above. For the pooled samples, 150 µL from each sample was collected from the individual tube and pooled with the 4 other samples (or the remaining samples) from the same litter. The results of both PCR tests were recorded for each litter (n = 180) from sow farms 1–3. If either pooled PCR identified a positive sample, the entire litter was considered positive for M. hyopneumoniae. The results of the individual PCR tests from sow farm 4 were also recorded for each piglet (n = 32).
Statistical analysis was completed using commercial software. h A linear mixed model was conducted with the Ct values as the response variable, the method of PCR extraction and sample collection technique as the fixed effects, and the individual pig selected as the random effect. Pairwise comparisons were performed among methods of PCR extraction and among sample collection technique.
The results from the first diagnostic investigation are summarized in Table 1. Nasal swabs gave the lowest frequency (6.3% positive), whereas the TBMC samples provided the highest frequency with 59.3% of the pigs being qPCR positive. Results showed that sample collection method also had a significant effect on the Ct value (P < 0.0001; Table 2). Pairwise comparisons revealed the markedly lower sensitivity of using nasal swabs and that the predicted least squares mean Ct value for the TBMC technique was lowest compared with the other 4 sample collection methods (Table 2). Based on these observations, TBMC was chosen for the field protocol. Using all extractions, 17 piglets (53.1%) with a negative nasal swab tested positive utilizing the TBMC sample (Table 1). The linear mixed model also showed that extraction method had an overall effect on the Ct value (P < 0.01). The interaction between extraction technique and sample collection method was not significant (P > 0.05). Pairwise comparisons of extraction technique revealed a significant difference between the Ct values of the viral (Ct = 32.9) and TNA (Ct = 29.5) extraction methods (P < 0.05), whereas pairwise comparisons between viral and high-volume (Ct = 32.1) extraction methods and high-volume and TNA extraction methods were not statistically significant (P > 0.05). The results of the pooling statistics indicated that when pooling the samples at a 1:5 and 1:10 dilution, the overall categorical outcome of the Ct value (as positive or negative) changed the least times using the high-volume extraction method. Based on a Ct cutoff value of 37, the highest sensitivity was obtained using the 1:5 dilution step. Based on all of these results, high-volume extraction and the 1:5 dilution were decided as the preferential approach for the field study.
Percent Mycoplasma hyopneumoniae polymerase chain reaction (PCR) positive by extraction method and sample type (diagnostic investigation no. 1).*
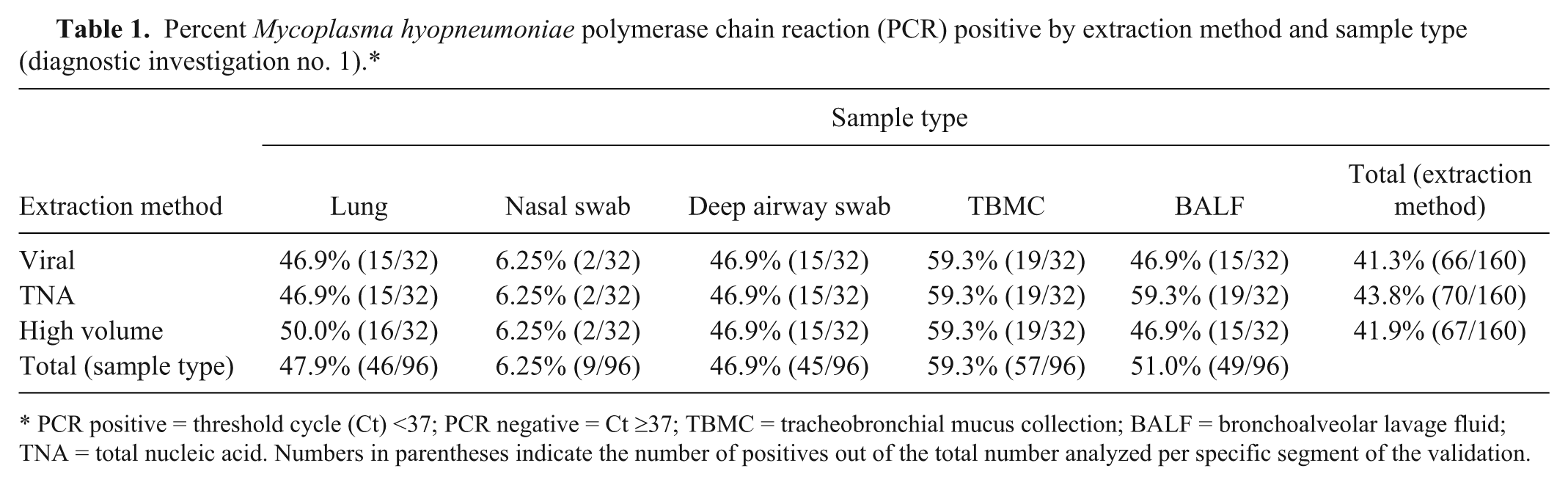
PCR positive = threshold cycle (Ct) <37; PCR negative = Ct ≥37; TBMC = tracheobronchial mucus collection; BALF = bronchoalveolar lavage fluid; TNA = total nucleic acid. Numbers in parentheses indicate the number of positives out of the total number analyzed per specific segment of the validation.
Predicted least squares mean of threshold cycle (Ct) values for 5 sample collection techniques for Mycoplasma hyopneumoniae.*
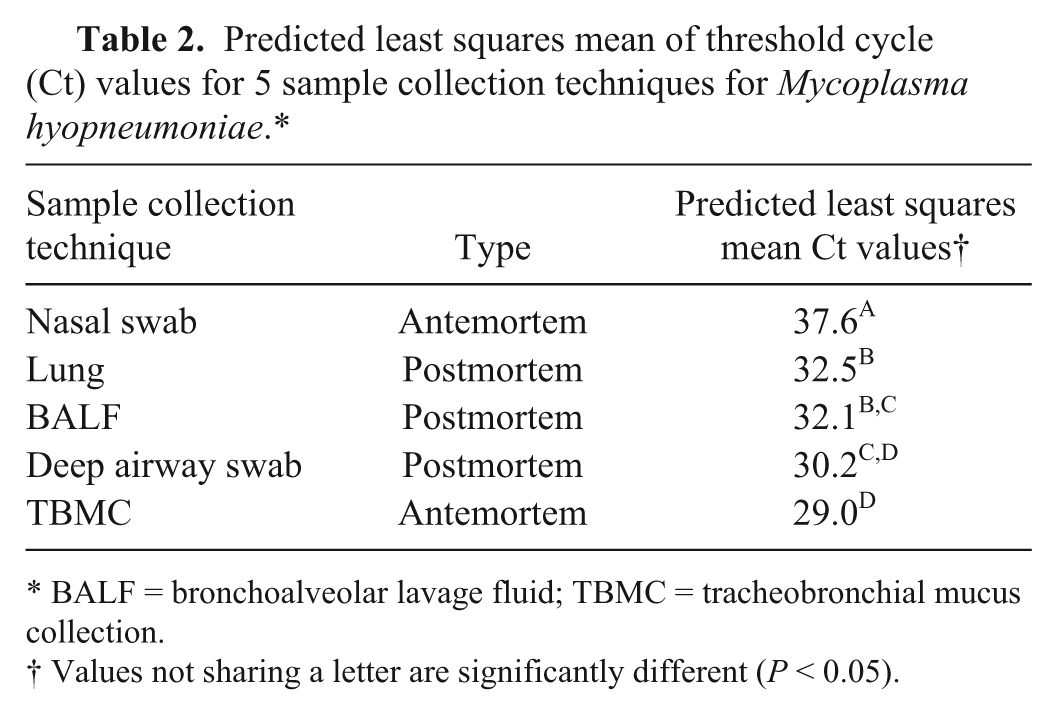
BALF = bronchoalveolar lavage fluid; TBMC = tracheobronchial mucus collection.
Values not sharing a letter are significantly different (P < 0.05).
The M. hyopneumoniae qPCR results from the pooled litters from sow farms 1–3 are shown in Table 3. Of the 180 litters sampled, 2 litters (1.1%; 95% confidence interval: 0%, +2.62%) returned positive qPCR results. Of the 32 piglets sampled from sow farm 4, 5 individuals (16.7%) returned positive qPCR results (data not shown).
Percent Mycoplasma hyopneumoniae positive by pooled real-time polymerase chain reaction (qPCR) assays by group and sow farm.*
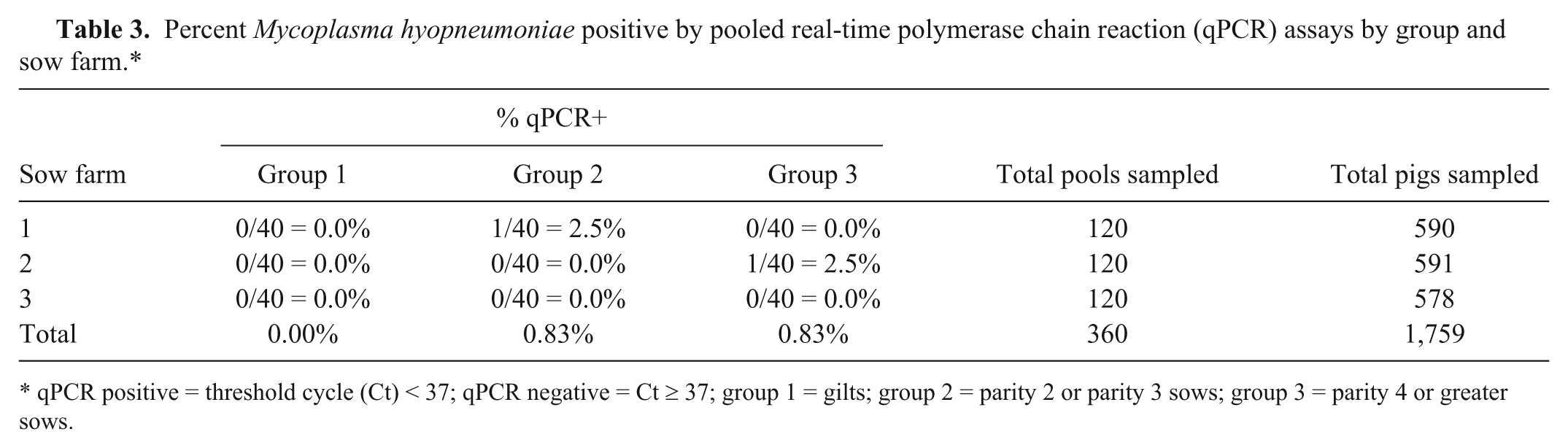
qPCR positive = threshold cycle (Ct) < 37; qPCR negative = Ct ≥ 37; group 1 = gilts; group 2 = parity 2 or parity 3 sows; group 3 = parity 4 or greater sows.
Understanding the prevalence of M. hyopneumoniae in the swine population is important information when considering management of herd outbreaks and routine herd health monitoring. Nasal swabs have been the method of sample collection in the past for antemortem herd detection. 1 However, there have been several studies7,9,14 reporting the use of more sensitive sampling techniques for detecting M. hyopneumoniae by PCR. These sampling methods, although more sensitive, are conducted postmortem. Other reports5,10,20 evaluating antemortem testing in combination with PCR have indicated that the ideal sampling sites are located in the lower respiratory tract of pigs, thus samples targeting the trachea and bronchi are more likely to yield M. hyopneumoniae organisms. A previous study 5 reported a technique that combines both antemortem testing capabilities with increased sensitivity; tracheobronchial swabbing proved to be 3.5 times more sensitive than nasal swabbing. 5 To the authors’ knowledge, no studies in the research literature can be found to apply this novel sampling technique to assess prevalence of M. hyopneumoniae in United States’ swine herds.
The results of both diagnostic investigations presented in this research suggest that TBMC is the most sensitive antemortem test available to assess true prevalence of M. hyopneumoniae in the pig population. The overall litter prevalence in the nonclinical preweaned pig population from our study was low (1.1%) and in agreement with a 2013 study 11 that reported a prevalence of 3.6% in the weaned pig population. However, the 2013 study 11 assessed the prevalence at the piglet level, whereas in our study, prevalence was determined at the litter level. A 2015 study was conducted using this TBMC technique in Benelux pig herds without specific respiratory disease. 19 The study showed a higher M. hyopneumoniae prevalence of at least 7.1% at 3–5 weeks of age, increasing to 10.9% at 6–11 weeks of age. 19 A potential explanation for this high prevalence might be that piglets in the youngest group were between 21 and 35 days of age, whereas in our study, the age at sampling was between 12 and 25 days of age. Although it is suggested that M. hyopneumoniae can be isolated in pigs as early as 1 week old, it is shown that prevalence increases with age, peaking at around the late nursery to early finishing stage. 19 Moreover, seasonal variations 19 and batch-to-batch variations 6 also have a significant impact on the prevalence of M. hyopneumoniae in early postweaning piglets. Therefore, it is possible that the piglets sampled in our study were truly negative for M. hyopneumoniae. Instead of targeting piglets at the age of weaning, it may be more fitting to assess the prevalence of M. hyopneumoniae in nursery-age piglets. In this scenario, it is likely that the combination of recent stressful events (weaning, movement, commingling) may initiate higher bacterial loads in the herd.
The low disease prevalence reported in our study is consistent, to a degree, with previous research.6,11,14 Using nasal swabs, the prevalence of M. hyopneumoniae in the weaned pig population was found to be low with quite a large batch-to-batch variation. 6 A previous study 11 reported a M. hyopneumoniae prevalence of 3.6% in the weaned pig population utilizing nasal swabs for sample collection and nested PCR for detection in clinically affected herds, while a 2007 study 14 reported a prevalence of 3.8% in 3-week-old piglets from sows in a herd with EP issues. However, the study design of these studies was different from the current study, as all samples were collected in M. hyopneumoniae clinical herds, whereas in our study, piglets were sampled from nonclinical farms. Another report, 21 using nasal swabs and nested PCR, found a disease prevalence of 16% in 6-week-old pigs clinically infected with EP. This value agrees with the prevalence estimate from the clinically affected sow farm 4. The increase in prevalence between the nonclinical and clinical sow farms suggest that this TBMC technique may be most judiciously applied to diseased herds to detect changes in the prevalence of M. hyopneumoniae that might have relevance for predicting performance of offspring or timing application of interventions.
Footnotes
Acknowledgements
We would like to thank the participating herd veterinarians, managers, and staff for their assistance during the study.
Authors’ contributions
F Vangroenweghe, L Karriker, K Crawford, E Christianson, and T Marsteller contributed to conception and design of the study. K Crawford, L Karriker, P Thomas, K Hammen, J Ellingson, and J Bates contributed to acquisition, analysis, and interpretation of data. F Vangroenweghe and R Main contributed to analysis and interpretation of data. K Harmon and S Abate contributed to analysis of data. F Vangroenweghe, L Karriker, and K Crawford drafted the manuscript. All authors critically revised the manuscript; gave final approval; and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately.
a.
Iowa State University Veterinary Diagnostic Laboratory, Ames, IA.
b.
MagMAX Pathogen RNA/DNA extraction kit, Life Technologies, Carlsbad, CA.
c.
MagMAX Total Nucleic Acid Isolation kit, Life Technologies, Carlsbad, CA.
d.
Kingfisher 96 magnetic particle processor, Thermo Fisher Scientific, Waltham, MA.
e.
f.
Applied Biosystems 7500 Fast real time PCR system, Life Technologies, Carlsbad, CA.
g.
VetMAX–Plus qPCR master mix and VetMAX M. hyopneumoniae reagents, Life Technologies, Carlsbad, CA.
h.
JMP10, SAS Institute, Cary, NC.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Three authors (F. Vangroenweghe, E. Christianson, T. Marsteller) are employees of Elanco Animal Health, a Division of Eli Lilly and Company, Greenfield, Indiana. While Eli Lilly and Company financed this study, the company played no role in the analysis and interpretation of data. None of the other authors has any other financial or personal relationships that could inappropriately influence or bias the content of the article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding was provided by Eli Lilly and Company, Greenfield, Indiana.
