Abstract
Bluetongue virus (BTV) is a vector-transmitted pathogen that typically infects and causes disease in domestic and wild ruminants. BTV is also known to infect domestic canines as discovered when dogs were vaccinated with a BTV-contaminated vaccine. Canine BTV infections have been documented through serological surveys, and natural infection by the Culicoides vector has been suggested. The report of isolation of BTV serotype 11 (BTV-11) from 2 separate domestic canine abortion cases in the states of Texas in 2011 and Kansas in 2012, were apparently unrelated to BTV-contaminated vaccination or consumption of BTV-contaminated raw meat as had been previously speculated. To elucidate the origin and relationship of these 2 domestic canine BTV-11 isolates, whole genome sequencing was performed. Six additional BTV-11 field isolates from Texas, Florida, and Washington, submitted for diagnostic investigation during 2011 and 2013, were also fully sequenced and analyzed. The phylogenetic analysis indicates that the BTV-11 domestic canine isolates are virtually identical, and both share high identity with 2 BTV-11 isolates identified from white-tailed deer in Texas in 2011. The results of the current study further support the hypothesis that a BTV-11 strain circulating in the Midwestern states could have been transmitted to the dogs by the infected Culicoides vector. Our study also expands the short list of available BTV-11 sequences, which may aid BTV surveillance and epidemiology.
Introduction
Bluetongue virus (BTV; family Reoviridae; subfamily Sedoreovirinae; genus Orbivirus) is the causative agent of the arthropod-transmitted disease bluetongue, an economically important disease that affects a wide range of domestic and wild ruminants.9,12 Although severe disease caused by BTV is most common in sheep and deer, subclinical infection typically occurs in goats and cattle. To date, 26 distinct serotypes of BTV are recognized worldwide, and a potential additional serotype has been identified in France.4,20 Virulence is not determined by serotype alone, and variation in disease severity reflects differences in virulence among specific viral strains, as well as host, vector, and environmental factors.4,12 The serotypes 10, 11, 13, and 17 occur throughout much of the United States, with their prevalence varying markedly between transmission seasons. 11 Generally BTV is not contagious; rather it is primarily vector-transmitted by biting midges of the genus Culicoides. 13
Abortion and death due to BTV infection in domestic dogs has also been reported. Earlier cases were linked to vaccination of dogs with a BTV-11–contaminated, multicomponent, modified live canine vaccine.1,8,18 Subsequent experimental infection of dogs (pregnant and not pregnant) with the isolated contaminant BTV-11strain led to disease manifestation in the infected pregnant females only, which included abortions, and pulmonary edema that resulted in death or euthanasia. 3 Consumption of contaminated raw meat has been proposed to be another potential route of infection.2,12 However, in other cases that suggest vector transmission of BTV to dogs may have occurred, there is serologic evidence of infection without a prior history of exposure due to a BTV-contaminated vaccine or raw meat diets. 14 In 2 incidences of domestic canine abortions associated with BTV infection, 6 contaminated raw meat and vaccination were ruled out as potential sources and thus the authors concluded that the infections were most likely due to direct transmission of BTV by the vector. Unfortunately, no official ongoing animal or entomological surveillance for BTV exists, and very little full-sequence data is available for BTV-11 in particular. Therefore, in order to further characterize and elucidate the relationship and origin of the 2 domestic canine BTV-11 isolates, whole genome sequencing was performed. Six additional BTV-11 field isolates from 4 states including Texas during 2011, where and when the first canine abortion case was identified, were also sequenced. The phylogenetic relationships among these BTV-11 sequenced genomes are discussed.
Materials and methods
BTV-11 isolates
The 2 isolates from the domestic canine aborted fetuses were provided by Edward Dubovi at Cornell University. Detailed case reports of the 2 canine isolates are described previously. 6 Briefly, the first case was a pregnant Rottweiler that aborted a litter of pups and later died in Texas, October 2011; the second was identified in September 2012 from a Bulldog breeding kennel located in Kansas where 2 abortion episodes occurred and 3 dogs were scored as strong positives by serum neutralization testing. Two transmission seasons were possible in the period between these canine abortion cases, at sites nearly 500 miles apart. Six additional BTV-11 field isolates were provided by the National Veterinary Services Laboratory in Ames, Iowa. These samples were from field specimens originally submitted for diagnostic or export testing purposes. Further information and accession numbers for all of the isolates are listed in Table 1.
Newly sequenced Bluetongue virus serotype 11 strains included in the current study.*
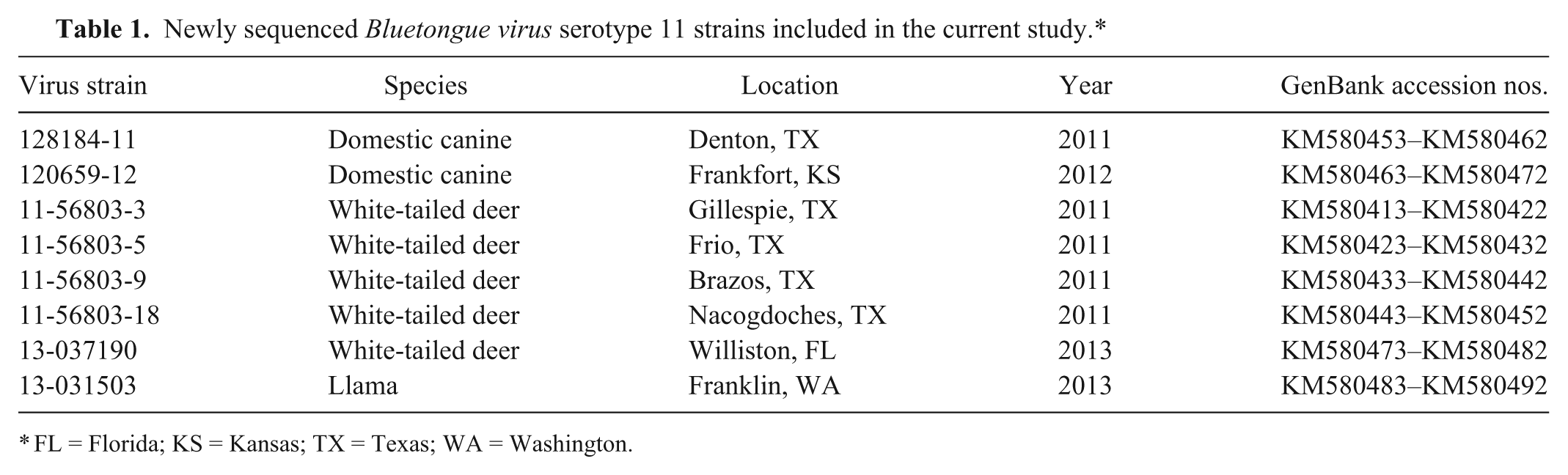

FL = Florida; KS = Kansas; TX = Texas; WA = Washington.
Whole genome sequencing and phylogenetic analysis
Virus strains were isolated and propagated in embryonated chicken eggs, followed by passage in baby hamster kidney (CCL 10) a or bovine pulmonary artery endothelial (CCL 209) a cells, for a total of 3 passages outside the original tissue. Viral double-stranded RNA was then purified from the extracted total RNA by a previously described method. 19 A whole genome sequence–independent amplification procedure was performed as previously described. 15 Briefly, restriction sites were removed from the anchor primer PC3-T7restrict (5′p-GTTCAGCCTGACCACGTTAATACGACTCACTATATTTTTATAGTGAGTCGTATTA-OH3′) to facilitate sequencing and then purified by high-performance liquid chromatography. The PC3-T7restrict primer (600 ng) was ligated to the double-stranded RNA (500 ng) in the ligation reaction mix as described previously 15 for a final volume of 90 μL. Ligation was performed at 37°C for 16 hr, and then purified using a commercial kit b following the protocol recommended by the manufacturer. The cleaned and ligated product (8 μL) was supplemented with 25 pmol PC2restrict primer (5′p-ACGTGGTCAGGCTGAAC-3′) and 1 μL of 300 mM methyl mercury hydroxide and allowed to denature for 10 min. The mixture was placed on ice, and 1.5 μL of 1 M β-mercaptoethanol was added and allowed to neutralize for 1 min. The copy DNA was then gener-ated using cloned Avian myeloblastosis virus reverse transcriptase, c annealed at 65°C for 90 min, and the remaining products neutralized as described previously. 15 Amplification of copy DNA was performed using 25 pmol PC2restrict primer and Taq enzyme. d Amplification products were confirmed by agarose gel electrophoresis and purified using the commercially available kit. b
Library preparation and sequencing was performed using a commercial platform e with the corresponding commercially available kits, f following the protocols recommended by the manufacturer. Briefly, approximately 1 μg of viral copy DNA was fragmented, barcoded, and quantitated. f Template generation, enrichment, and sequencing of the template were performed on the appropriate instruments. e
Standard flow gram format files were imported to commercial softwareg for contig creation. Partial contigs were assembled and blasted to determine reference sequences that were used for reference assemblies. The depth of sequence coverage ranged from 76 to 1,068 per isolate with a median coverage of 916. Multiple sequence alignments of the nucleotide sequences were performed using the multiple alignment fast Fourier transformation method followed by in silico translation of the open reading frames and alignment of the predicted proteins for each of the 10 genomic segments. Phylogenetic analysis was performed using the neighbor-joining method and 1,000 replicates to generate bootstrap confidence values. Distance relationships were calculated from both the nucleotide and protein alignments. Although a comprehensive analysis was performed with all full-length gene sequences available from GenBank, for clarity, only the most relevant, closely related strains and U.S. prototype BTV strains are shown.
Results
Phylogenetic analysis revealed that the 2 domestic canine isolates are nearly identical, although slight genetic drift was observed among almost all of the gene segments with the exception of M6 (VP5), which were 100% identical (Table 2, column A). The overall genetic identity shared between the genome segments of the 2 domestic canine isolates ranged from 96 to 100%, and the predicted proteins ranged from 99 to 100%. Above 79% genetic identity and above 86% predicted protein identity was shared between the domestic canine isolates and the other U.S. BTV-11 isolates sequenced (Table 2, column B).
Comparison of genetic (and predicted protein) sequences of Bluetongue virus serotype 11 (BTV-11) and U.S. prototype BTV strains.*
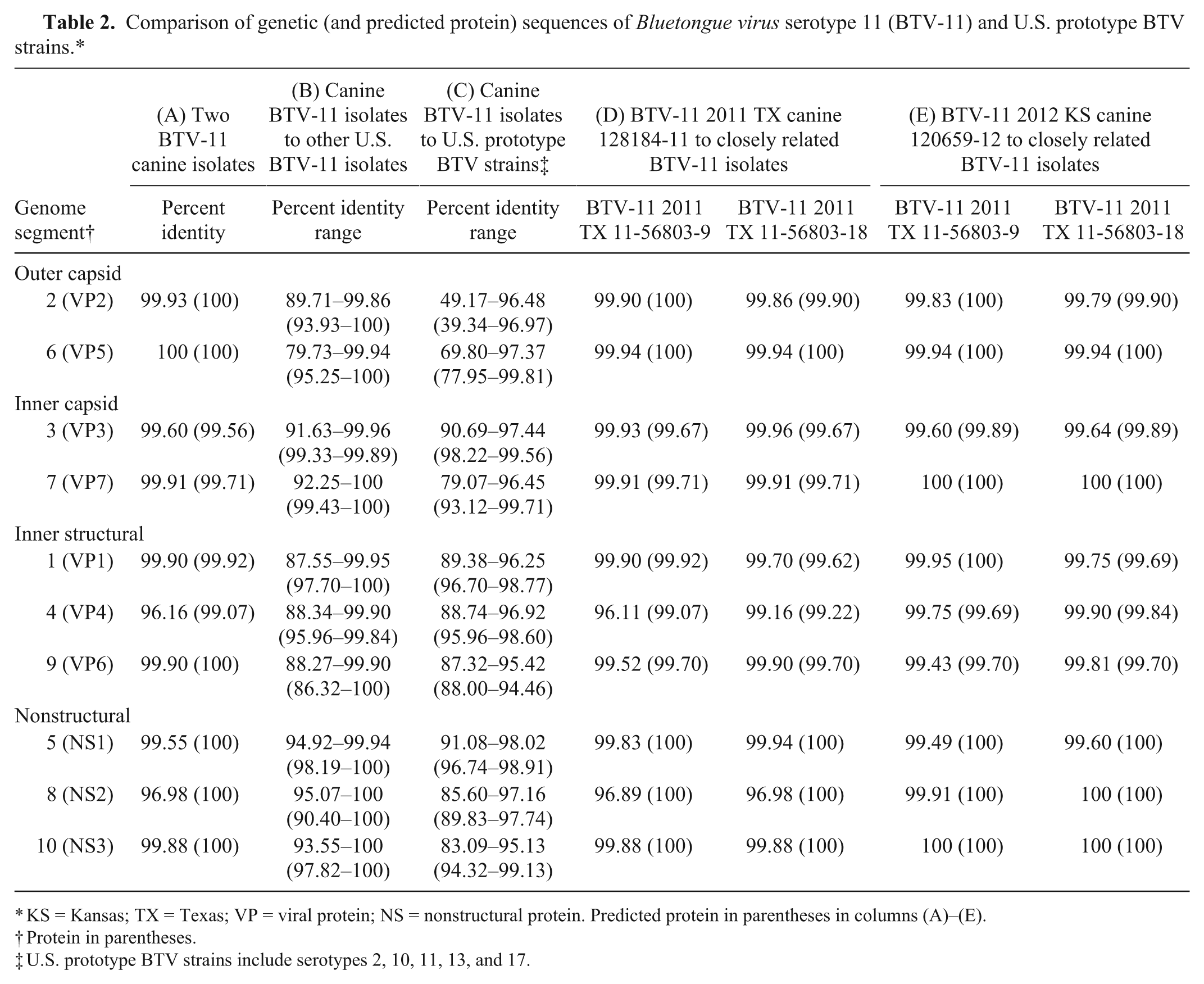
KS = Kansas; TX = Texas; VP = viral protein; NS = nonstructural protein. Predicted protein in parentheses in columns (A)–(E).
Protein in parentheses.
U.S. prototype BTV strains include serotypes 2, 10, 11, 13, and 17.
Two Texas BTV-11 isolates from 2011 in particular were nearly identical to the domestic canine isolates. The relationship of these 4 isolates is depicted by their phylogenetic grouping in the trees produced for each of the genome segments (Figs. 1, 2). The Texas isolates 11-56803-9 and 11-56803-18 were identified from white-tailed deer (WTD) in the counties of Brazos and Nacogdoches, respectively, and shared 96–100% genetic identity and 99–100% predicted protein identity with the domestic canine isolates (Table 2, columns D, E).
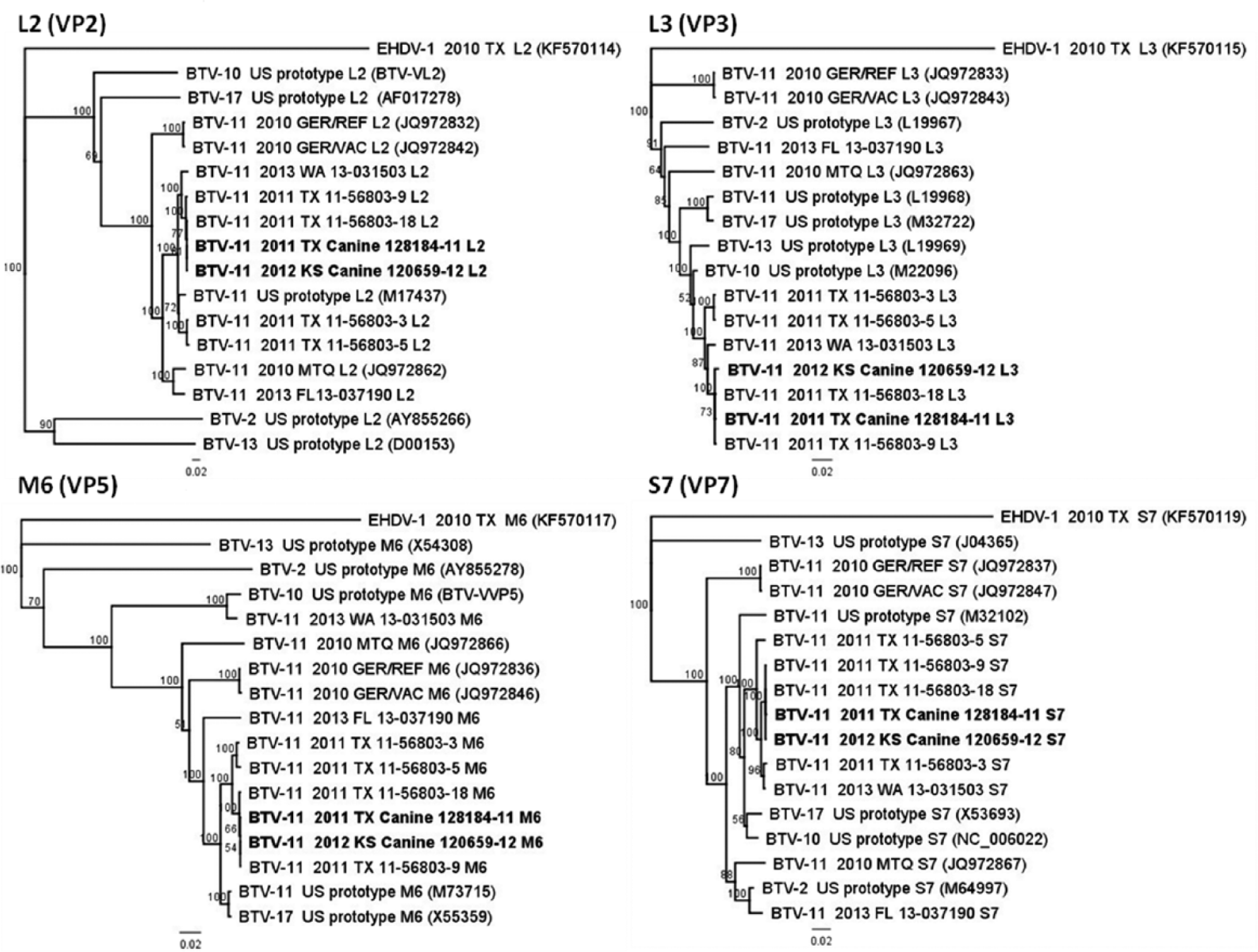
Phylogenetic trees generated for Bluetongue virus (BTV) outer capsid (VP2, VP5) and inner capsid (VP3, VP7) gene encoding segments. Neighbor-joining trees were built with other full-length gene sequences of BTV serotype 11 (BTV-11) and U.S. prototypic BTV strains, and 1,000 replicates were used to generate the bootstrap confidence values. The Epizootic hemorrhagic disease virus serotype 1 (EHDV-1) strain was included as the outgroup in this analysis. GenBank accession numbers are provided for previously published sequences. TX = Texas; KS = Kansas; WA = Washington; FL = Florida; GER = Germany; MTQ = Martinique.
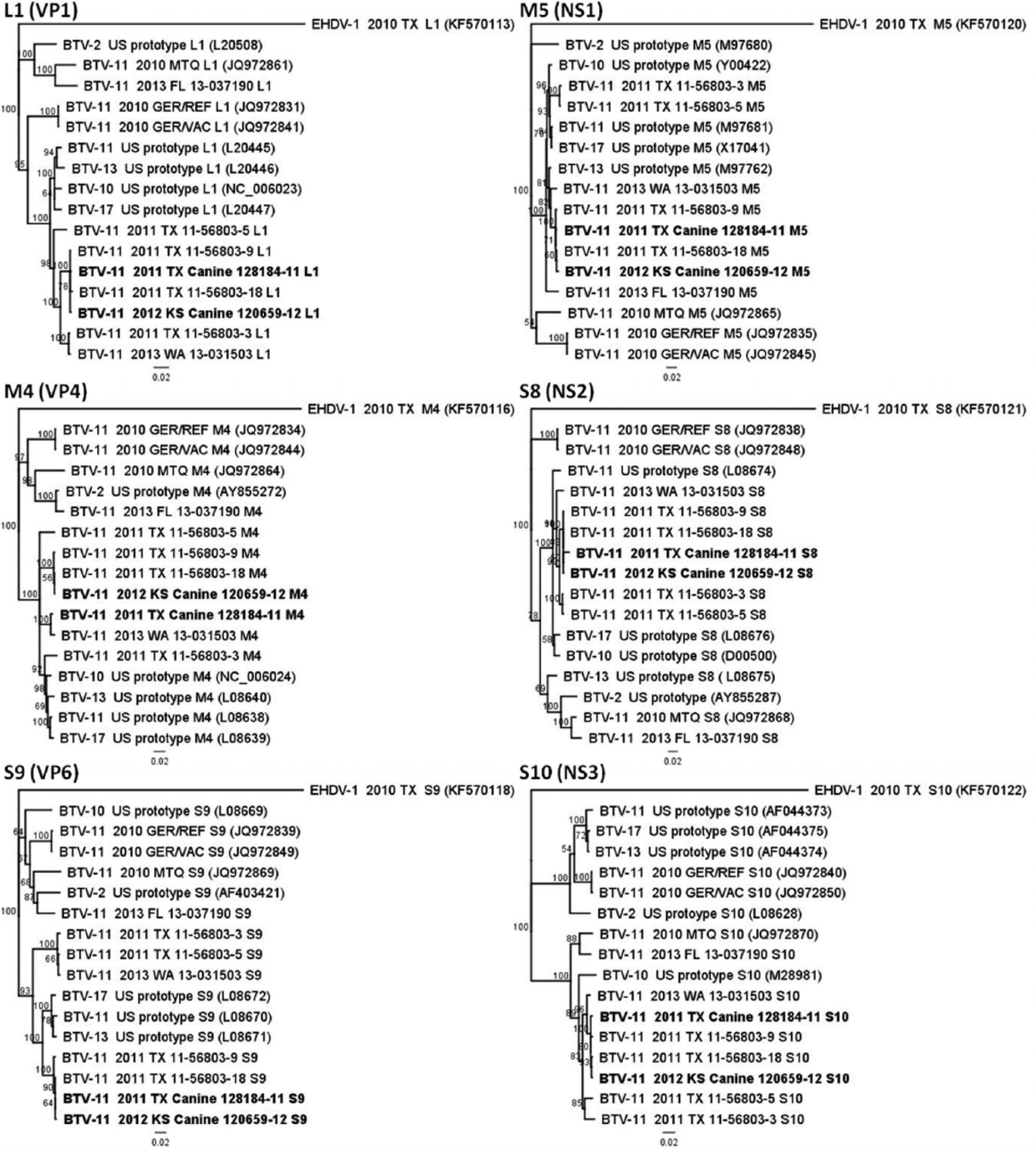

Phylogenetic trees generated for the inner structural (VP1, VP4, VP6) and nonstructural (NS1, NS2, NS3) gene encoding segments. Neighbor-joining trees were built with other full-length gene sequences of Bluetongue virus serotype 11 (BTV-11) and U.S. prototypic BTV strains, and 1,000 replicates were used to generate the bootstrap confidence values. The Epizootic hemorrhagic disease virus serotype 1 (EHDV-1) strain was included as the outgroup in this analysis. GenBank accession numbers are provided for previously published sequences. TX = Texas; KS = Kansas; WA = Washington; FL = Florida; GER = Germany; MTQ = Martinique.
The other two 2011 Texas isolates (11-56803-3 and 11-56803-5) were 91–98.9% genetically similar to the domestic canine isolates. With the exception of the M6 segment, the 2013 Washington BTV-11 isolate was 91–99.1% genetically similar to the domestic canine isolates. The M6 segment of the Washington isolate was most closely related to the BTV-10 US prototype strain (97.9%). All 3 of these isolates shared 91–99.7% predicted protein identity with the domestic canine isolates.
The BTV-11 Florida 2013 isolate was more related to the BTV-11 Martinique 2010 and the BTV-2 U.S. prototype strains, as illustrated by the trees for genome segments L1, M4, S7, S8, and S9 (Figs. 1, 2). This is not surprising, as it appears cocirculation and reassortment occurs among southeastern U.S. and Caribbean BTV strains.11,17 The German BTV-11 reference and vaccine strains were more distantly related and formed a distinct branch from the other BTV-11 and U.S. prototype strains (Figs. 1, 2).
Discussion
It has long been thought that BTV infection of domestic canines has been the result of vaccination with a contaminated vaccine or ingestion of contaminated raw meat. Yet, infection in dogs through vector transmission is possible, although it has not been directly proven and many questions remain unanswered. A serological surveillance study of domestic dogs conducted in Morocco in an area where BTV-1 had recently circulated in the ruminant population, found that out of a total of 187 samples tested, 21% were BTV antibody positive, of which 11% were identified as serotype 1. 14 The dogs surveyed were kenneled outside and fed only commercially produced canned food and did not have access to potentially infected meat or meat products, supporting the idea of infection through vector feeding. 14 However, detailed vaccination histories of the dogs surveyed were not mentioned. 14
In the report of the 2 U.S. domestic dog BTV cases, it was concluded that the infections were most likely due to direct transmission by the vector because contaminated raw meat and vaccination were ruled out as potential sources. 6 However, it has also been suggested that BTV-contaminated fetal bovine serum (FBS), used as a semen extender for artificial insemination, could be a suspected source of infection. 7 FBS was one of the sources proposed to be responsible for the BTV-11 contamination of the multicomponent canine vaccine. 8 This route of exposure is plausible as artificial insemination was performed in both of the domestic dog cases. 6 Unfortunately, no detailed information is available for whether FBS was used as an extender in these cases and thus this hypothesis can neither be confirmed nor disproved at present. There is also no sequence information available for the BTV-11 contaminant of the canine vaccine. However, because of the high sequence identity shared between the domestic canine isolates and the field strains circulating in the area during the same period of time, it is conceivable that natural infection by vector transmission could have occurred. In the Texas case report, all of the owner’s dogs were reported to have access to an outdoor area that was adjacent to a pasture with cattle. 6 As cattle are usually asymptomatic and there is no vaccination program for BTV-11 in the United States, the cattle could have been viremic without any knowledge of a subclinical infection. Deer would have also had access to the vicinity, thus there were 2 possible amplifying host sources that could have been fed on by the Culicoides vector, which could have then subsequently passed the virus onto the dogs. Nonetheless, Culicoides are often considered to be responsible for long-distance dispersal of BTV. Therefore, it is also plausible that in the absence of a primary host in the immediate vicinity, infected Culicoides could have alternatively fed on the dogs for a blood meal. An additional BTV-11 isolate from Iowa was reported in 2012 that shared 99% similarity with the canine isolates, although only partial sequences of a conserved region of the VP2 gene were evaluated. 6 Thus, the relationship of the Iowa BTV-11 strain to the canine and Texas isolates cannot be assumed, but it does support the hypothesis that this particular BTV-11 strain may have been circulating throughout the Midwest during this time.
Determining the epidemiology of BTV remains challenging given there is no official BTV surveillance program, and current available diagnostic tools are not effective for acquiring strain variations aside from serotype. Improved and more cost-effective diagnostic tools that can be used for the development of a better national surveillance strategy for BTV are needed. As mentioned above, disease severity is dependent on the virus strain as well as host, vector, and environmental factors. Subclinical infection, which is typical of BTV, can allow the virus to go unnoticed and thus be maintained and disseminated. Under ideal conditions, especially introduction to naïve flocks, which results in clinical infections, BTV can cause severe disease that can have significant economic impact due to effects on animal health and trade restrictions. Incursion of BTV into new territories10,16 is of genuine concern as it can have great consequences, as was the case with BTV-8 in cattle of Northern and Central Europe. 5 Although it has been known for a while that domestic canines are BTV susceptible, it remains unknown whether Culicoides actually feed on dogs, and whether viremia in dogs can result in BTV being passed on to the vector. The current study provides further support that vector transmission of a circulating BTV-11 field strain to domesticated dogs could have occurred, and also expands the limited inventory of whole genome–sequenced BTV-11 strains, which may aid in BTV epidemiology. Further studies are necessary to address host preference for canines among blood-feeding North American Culicoides midges, and differences in canine susceptibility to certain serotypes of BTV. Furthermore, pregnancy-associated immune suppression may explain why BTV-induced clinical disease is more often documented in pregnant dogs, but how sex, age, or pregnancy affect susceptibility and BTV viremia in dogs, and whether the virus can be passed forward from these animals to competent Culicoides vectors requires further investigation.
Footnotes
Acknowledgements
We thank Drs. Andrew Allison and Phillip Schumm for review of this article.
Authors’ note
Mention of trade names or commercial products in this report is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA. USDA is an equal opportunity provider and employer.
Authors’ contributions
NN Gaudreault and WC Wilson contributed to conception and design of the study. NN Gaudreault contributed to analysis and interpretation of data. DC Jasperson contributed to acquisition and analysis of data. EJ Dubovi, DJ Johnson, EN Ostlund, and WC Wilson contributed to acquisition of data. NN Gaudreault drafted the manuscript. All authors critically revised the manuscript, gave final approval, and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
American Type Culture Collection, Manassas, VA.
b.
QIAquick gel extraction, Qiagen Inc., Valencia, CA.
c.
AMV Reverse Transcriptase, Life Technologies, Grand Island, NY.
d.
TakaRa Ex Taq, Clontech Laboratories Inc., Mountain View, CA.
e.
Ion Torrent OneTouch, OneTouch ES, Personal Genome Machine; Life Technologies, Grand Island, NY.
f.
Ion Xpress Plus fragmentation library kit, Xpress barcode adapters, Ion library quantitation kit, OneTouch 200 template kit v2, Ion PGM sequencing 400 kit, Ion 314 chip; Life Technologies, Grand Island, NY.
g.
Geneious 6.0, Biomatters Ltd., Auckland, New Zealand.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the United States Department of Agriculture (USDA), Agricultural Research Service (project 5430-32000-006-00D).
