Abstract
In February and March 2009, approximately 1,500 backyard pigs of variable age became sick, and approximately 700 of them died or were euthanized in the Lower Artibonite Valley and the Lower Plateau of the Republic of Haiti. The main clinical sign was posterior ataxia followed by paresis and/or paralysis on the second or third day of illness. No gross lesions were observed at postmortem examinations. The morbidity and mortality were approximately 60% and 40%, respectively. Diagnostic samples (whole blood, brain, tonsil, lymph nodes, spleen, and lung) were negative for Classical swine fever virus and African swine fever virus. Porcine teschovirus type 1 was detected by reverse transcription polymerase chain reactions in brain samples. Results of virus isolation, electron microscopy of virus particles, histopathological analysis on brain tissues, nucleic acid sequencing, and phylogenetic analysis of the viral isolate supported the diagnosis of teschovirus encephalomyelitis. The outbreak of the disease in Haiti is the first appearance of the severe form of teschovirus encephalomyelitis in the Americas. This disease poses a potential threat to the swine industries in other Caribbean countries, as well as to Central and North American countries.
Introduction
Teschovirus encephalomyelitis was first described as a particularly virulent, highly fatal encephalomyelitis of pigs and was previously known as Teschen disease (Center for Food Security and Public Health of Iowa State University: 2009, Teschovirus encephalomyelitis and porcine teschovirus infection. Available at: www.cfsph.iastate.edu/Factsheets/pdfs/enterovirus_encephalomyelitis.pdf). 20 The causal agent of teschovirus encephalomyelitis is Porcine teschovirus (PTV) serotype 1, which belongs to the genus Teschovirus, family Picornaviridae, order Picornavirales (Center for Food Security and Public Health of Iowa State University). There are at least 11 distinct serotypes of this species. Porcine teschovirus was previously classified as group I Porcine enterovirus (PEV) of the genus Enterovirus.10,11,21 Some virulent strains of PTV cause severe encephalomyelitis. Other strains of serotype 1 as well as other PTV serotypes can cause milder forms of the disease or inapparent infections in pigs. These milder forms of the disease have been given a variety of names, including Talfan disease, benign enzootic paresis, and poliomyelitis suum (Center for Food Security and Public Health of Iowa State University). 20
Outbreaks of teschovirus encephalomyelitis were reported in Europe from 1929 to the 1950s, with the virus spreading to other continents; however, the severe form of the disease has not been reported in Western Europe after 1980, and it is now considered rare (Food and Agriculture Organization of the United Nations [FAO]: 2009, Teschovirus encephalomyelitis in the Republic of Haiti. FAO Emergency Prevention System for Transboundary Animal and Plant Pests and Diseases. No. 4. Available at: ftp://ftp.fao.org/docrep/fao/012/ak137e/ak137e00.pdf; Center for Food Security and Public Health of Iowa State University: 2009, Teschovirus encephalomyelitis and porcine teschovirus infection). 20 Since 1996, teschovirus encephalomyelitis has been reported to the World Organization for Animal Health (OIE) by Belarus, Japan, Latvia, Madagascar, Moldavia, Romania, Russia, Uganda, and Ukraine (FAO; Center for Food Security and Public Health of Iowa State University). 20 Milder forms of the disease have been observed in Western Europe, Africa, 16 the United States,1,14 Canada, 16 Australia, 8 and China. 6 Porcine teschovirus can enter the body by ingestion and multiply in the gastrointestinal tract and associated lymphoid tissues, including the tonsils (Center for Food Security and Public Health of Iowa State University). The virus is excreted in feces and urine for several weeks, and the main route of transmission is fecal-oral, directly or indirectly, from contaminated sources of food or water (FAO).
In February and March 2009, an estimated 1,500 backyard pigs became sick, and approximately 700 of them died or were culled in the Lower Artibonite Valley and the Lower Plateau of the Republic of Haiti. Pigs of all ages were affected. Clinical signs included anorexia, locomotive disorders, nystagmus, opisthotonos, and paresis and/or paralysis of the hindquarters. Affected animals did not exhibit pyrexia, and no gross lesions were observed upon postmortem examinations. Morbidity and mortality were estimated at 60% and 40%, respectively.
Specimens (ethylenediamine tetra-acetic acid anticoagulated blood [EDTA blood], brain, tonsil, spleen, lung, and lymph nodes) collected from sick pigs were submitted to the U.S. Department of Agriculture National Veterinary Services Laboratories (NVSL) in Ames, Iowa, and the NVSL Foreign Animal Disease Diagnostic Laboratory (FADDL) at Plum Island, New York, for disease diagnosis. These samples were negative for Classical swine fever virus (CSFV) and African swine fever virus (ASFV). Porcine teschovirus serotype 1 was detected in brain tissue samples by reverse transcription polymerase chain reaction (RT-PCR). Results of virus isolation (VI) in tissue cultures, electron microscopy, histopathology of brain tissues, nucleic acid sequencing, and phylogenetic analysis were all consistent with a diagnosis of teschovirus encephalomyelitis in swine in the Republic of Haiti.
Materials and methods
Clinical specimens
All clinical specimens were collected from pigs presenting with clinical signs of neuropathy in the Lower Artibonite Valley and the Lower Plateau of the Republic of Haiti and submitted by the Chief Veterinary Officer of the Ministry of Agriculture, Port-au-Prince, Haiti. The samples included serum and EDTA blood from 6 pigs, as well as brain, tonsil, spleen, lung, and mandibular, renal, and mesenteric lymph nodes from 3 pigs.
Sample preparation and virus isolation
A 10% tissue homogenate was prepared for brain, tonsil, spleen, lung, and lymph nodes in Eagle minimal essential medium (EMEM) a supplemented with 4% fetal bovine serum (FBS) a using a mixer b at a setting of vibrational frequency of 18 cycles per second for a total of 2 min. All homogenates were clarified by centrifugation at 2,500 × g for 10 min and then filtered through 0.45-µm cellulose acetate filters. c For VI at FADDL, a volume of 0.5 ml of each filtrate was inoculated onto cultures of swine kidney (SK-6 and IBRS-2) and African green monkey kidney epithelial (Vero) cells grown in 25-cm2 cell culture flasks. The inoculum was adsorbed for 60 min at 37°C, and 5 ml EMEM with 4% FBS was then added to each flask. The flasks were incubated at 37°C and observed daily for cytopathic effect (CPE). When no CPE was observed after 3 days of incubation, the content of the flasks was frozen and thawed, and a volume of 0.5 ml lysate of each flask was inoculated into a new flask of each cell line described above. The cultures positive for CPE were aliquoted and frozen at −70°C for further analysis by electron microscopy and RT-PCR when needed. For isolation of CSFV, cultures of SK-6 cells negative for CPE were tested for CSFV antigens with the avidin–biotin complex (ABC) immunohistochemistry assay described below.
For VI at NVSL-Ames, porcine kidney (PK-15), swine testicle (ST), Vero V76, monkey kidney (Marc 145), and primary swine kidney (pSK) were used. A volume of 1– 2 ml of sample suspension was inoculated onto cells, incubated for 1–2 hr at 37°C, and rinsed 3 times with cell culture medium containing 2.5% FBS, and 8–10 ml of the cell culture medium was added to the flask. All cells were incubated at 37°C and observed daily for CPE. Cells were frozen when CPE was observed or after 5–7 days of incubation with no CPE. Cells were thawed and inoculated onto the same cells grown on coverslips in Leighton tubes d (1 ml/tube). Cells were incubated at 37°C for 1–2 hr, rinsed with cell culture medium containing 2.5% FBS, and maintained in the same medium. Cells were incubated at 37°C and observed daily for CPE. Cells on coverslips were fixed in 100% acetone at −20°C for 10 min and air dried when CPE was observed, or after 5–7 days of incubation with no CPE. A volume of 200 µl of porcine antiserum known positive for antibodies to PTV serotype 1 was added to each coverslip and incubated at 37°C for 30 min in a humidified chamber. The coverslips were rinsed in phosphate buffered saline (PBS) without Ca++ and Mg++ (pH 7.2) and soaked once in the PBS for 5 min. A volume of 200 µl of a 1:75 dilution of laboratory-prepared goat anti-pig immunoglobulin (Ig)G conjugated with fluorescein isothiocyanate (FITC) was added onto each coverslip and incubated at 37°C for 30 min in a humid chamber. The coverslips were rinsed in PBS (pH 7.2), soaked in the PBS for 5 min, rinsed in reverse-osmosis water, and allowed to air dry. After drying, the coverslips were mounted on microscopic slides with a mounting medium containing 50% glycerin and 50% PBS and examined under a fluorescence microscope. The PTV serotype 1–infected cells appeared bright green on a dark background.
Nucleic acid extraction
RNA was extracted from 140 µl of each tissue homogenate of brain, tonsil, spleen, and lymph node, as well as EDTA blood using a commercial kit. e RNA from each sample was eluted in 40 µl of RNase-free water f and stored at −70°C until RT-PCR was performed. DNA was extracted from 200 µl of each tissue homogenate of tonsil, spleen, and lymph nodes, as well as EDTA blood with a DNA extraction kit e following the manufacturer’s instructions. DNA from each sample was eluted in 100 µl of Buffer AE and stored at −70°C until PCR tests were performed.
Detection of Porcine teschovirus by RT-PCR
Two RT-PCR assays were conducted on RNA of brain samples as described previously. 22 The first assay was a nested RT-PCR for the detection of group I PEV (i.e., PTV serotypes 1–11). The second assay was specific for PTV serotype 1. In both assays, sample RNA was reverse transcribed using a commercial kit f according to the supplier’s instruction. PCR master mixes were prepared under the following conditions: 0.3 µM each primer, 0.2 mM each deoxyribonucleotide triphosphate, 1.5 mM MgCl2, 10 mM Tris-HCl (pH 8.3), 50 mM KCl, and 1.25 units of DNA polymerase. g PCR assays were performed on a thermal cycler. h PCR conditions were the same as described previously. 22 PCR products were analyzed by agarose gel electrophoresis with ethidium bromide staining. A sample was considered positive for PTV when it generated PCR products of the expected size of 158 bp in the nested RT-PCR, whereas a sample was deemed positive for PTV serotype 1 when it generated PCR products of the expected size of 683 bp in the PTV serotype 1–specific PCR.
Real-time RT-PCR for Classical swine fever virus
A TaqMan real-time RT-PCR was conducted on RNA of tonsil, spleen, lymph nodes, and EDTA blood samples for the detection of CSFV according to a previous study 15 with minor modifications. Primers and probe were designed to target the 5′ untranslated region (UTR) of the CSFV genome. Sequences of primers and probe were as follows: forward primer, 5′-TGCCCAAGACACACCTTAACC-3′; reverse primer, 5′-GGCCTCTGCAGCGCCCTAT-3′; and probe, 5′-TGATGGGAGTACGACCTG-3′. The probe was labeled with a 5′ reporter dye, 6-carboxyfluorescein (FAM), and a 3′ nonfluorescent quencher (NFQ). g PCR master mixes were prepared using a commercial RT-PCR kit. e The RT-PCR was carried out in a volume of 25 µl, including 2.5 µl RNA sample, 0.2 µM forward primer, 0.4 µM reverse primer, 0.2 µM probe, 12.5 µl of 2× master mix, e 50 nM 5-carboxy-X-rhodamine (ROX) reference dye, and 0.25 µl of RT mix. e It was performed on a real-time PCR system g using the following cycling conditions: 50°C for 30 min and 95°C for 15 min, followed by 45 cycles of 94°C for 15 sec and 56°C for 60 sec.
Real-time PCR for African swine fever virus
A TaqMan PCR was conducted on DNA of tonsil, spleen, lymph nodes, and EDTA blood samples for the detection of ASFV according to a previous study. 23 Oligonucleotide primers and probe targeted a gene encoding p72, a highly conserved structural protein of ASFV. Sequences of primers and probe were as follows: forward primer, 5′-CCTCGGCGAGCGCTTTATCAC; reverse primer, 5′-GGAAACTCATTCACCAAATCCTT; and probe, 5′-CGATGCAAGCTTTAT. The probe was labeled with a 5′ reporter dye, FAM, and a 3′ NFQ. g Reagents from a PCR kit g were used to prepare master mixture recipes according to the guidelines of the manufacturer for individual component concentrations. The PCR assay was performed on a thermal cycler i with 45 amplification cycles (95°C for 2 sec and 60°C for 30 sec).
Detection of antibodies to CSFV and ASFV
For antibodies to CSFV, a commercial test kit j was used for serum samples according to the manufacturer’s instructions. A sample was considered positive for the presence of antibodies to a pestivirus if its blocking percentage was 40% or greater. An immunoperoxidase–virus neutralization (IP-VN) test was performed to differentiate between antibodies to CSFV and Bovine viral diarrhea virus (BVDV) as described previously 18 with minor modifications. SK-6 cells were used in the IP-VN test. A 4-fold difference or more between the end-point titers for CSFV and BVDV was considered to be indicative for an infection by the virus yielding the higher titer. For antibodies to ASFV, an ELISA kit k was used following the manufacturer’s instructions.
Detection of CSFV and ASFV antigens
Tonsil, spleen, lymph nodes, and cultures of SK-6 cells were tested for the presence of CSFV antigens with the ABC immunohistochemistry assay using a commercial kit l and monoclonal antibody V3 m against the glycoprotein 55 of CSFV according to the manufacturers’ instructions. A direct fluorescent antibody test (DFAT) was used for the detection of ASFV antigens in tonsil, spleen, kidney, and lymph nodes as described by the following. Frozen tissue sections of lymph nodes and spleen were prepared by cut using a microtome. n The slides were fixed with 100% acetone at −20°C for 10 min and then air dried. A 1:10 dilution of ASFV DFAT conjugate was used to cover the section, and the slides were incubated at 37°C for 30 min in a humidified chamber. The slides were placed in a coplin jar filled with PBS (pH 7.2) containing 0.05% Tween-20 and incubated at room temperature for approximately 30 sec. The wash buffer was poured out and replaced with fresh wash buffer for the second wash. The slides were incubated with the wash buffer for 5 min on the countertop with slight agitation. This step was repeated 3 times. The slides were rinsed in running ultrapure laboratory grade water for 30 sec, then air dried and read with a ultraviolet microscope. A sample was interpreted positive for the presence of ASFV antigen when cells showed cytoplasmic fluorescent staining, and the known positive and known negative control tissues stained as expected.
Hemadsorption test for the detection of ASFV
Homogenates of brain, tonsil, spleen, lung, and lymph nodes were tested for ASFV using the hemadsorption (HAD) test. Inoculation of swine macrophages from peripheral blood was performed as described previously. 12 Briefly, peripheral blood macrophage cells were adhered to 96-well cell culture plates 1 day prior to inoculation. A volume of 100 µl of filtered 2-fold dilutions of tissue homogenate was prepared and added to each designated well. A minimum of 4 wells was used for each dilution of each sample. Inoculated plates were incubated at 37°C with 5% CO2 for 24 hr. Thereafter, a volume of 10 µl of 1% pig erythrocytes, which were collected from the same animal as the macrophages and suspended in PBS (pH 7.2), was added to each well, and the plates were observed for HAD and CPE daily under a microscope for 10 days. When no change was observed, the plates were frozen and thawed, and a volume of 50 µl of the supernatant in each well was sub-inoculated into fresh macrophage cultures for further incubation and observation as described above.
Electron microscopy
Aliqouted and frozen cultures of SK-6 cells positive for CPE in virus isolation were thawed at 37°C in a water bath. The cell lysate was spun at 10,000 × g for 20 min to remove debris, and the supernatant was harvested and spun in an ultracentrifuge o at 100,000 × g for 30 min. The pellet was resuspended in 20 µl of sterile water and adsorbed to polyvinyl formal carbon-coated grids that had been treated with 1% Alcian blue. The grids then were fixed with 4% formaldehyde p in PBS for 10 min at room temperature, washed in PBS (2×), and stained with 2% phosphotungstic acid. Grids were imaged with an electron microscope q operating at 80 kV. Viral images were captured with a digital camera, r and virus particles were measured with software. r
Histopathological analysis
Brain tissues that had been fixed in 10% neutral buffered formalin were paraffin-embedded, sectioned at 4 µm, mounted on glass slides, and stained with hematoxylin and eosin as described previously. 7
Sequence analysis for the entire genome of the Haitian isolate
The viral isolate was analyzed by pan-viral microarray as previously described. 2 Briefly, subsequent to scanning, a volume of 100 µl of sterile water at 95°C was added to the surface of the hybridized microarray and recovered. This was repeated 6 times, and resulting material was amplified by PCR using the ligated tag primers 5′-GTTTCCAAGTCACGATC-3′ and 5′-CATTACAAATGTGAACCTT-3′ as previously described. PCR products were agarose gel purified and cloned into TOP10 competent cells f and grown at 37°C. Ninety-six colonies were selected and added to a 96-well round-bottom plate with 25 µl of PCR mix, f PCR amplified, and submitted for sequence analysis. Resulting sequences were then used to design the primer pairs 5′-GTCACCGGGTATTGCACC AA-3′ and 5′-GGGCCAGACACAGTGATGG-3′, which were used for directed RT-PCR amplification of a single large genome fragment. In addition, primers 5′-CTGCAT TCCCATAAAGGAACTCC-3′ and 5′-CCATCACTGTGTC TGGCCC-3′ were designed and used for RT-PCR amplification of the 5′ and 3′ ends of the viral genome, respectively, using commercial reagents. f This process yielded 3 fragments spanning the complete genome of PTV serotype 1. RT-PCR products were subjected to Sanger dideoxy-sequencing using a cycle sequencing kit. f Sequencing reactions were purified using a purification system o and analyzed on a sequence analyzer. g Sequence contigs were read and assembled using a DNA sequence software. s
Phylogenetic analysis
Phylogenetic analysis was performed by alignment of the nucleotide sequence of the Haitian isolate with those of other teschoviruses using version 1.4 of ClustalW software. 19 Sequences were trimmed, edited, and exported to PHYLIP format using BioEdit software (version 7.0.5.3). 9 Phylogenetic analysis and assembly of the tree for PTV was performed using a software package t for analyzing the complete polyprotein coding region of teschoviruses.5,17 The bootstrap consensus trees were inferred from 1,000 replicates with calculation of the distance matrix based on nucleotide sequence. The output trees were then analyzed using the neighbor-joining method to produce a final consensus tree. Phylogenetic tree graphics were produced using a graphic tool. t Output tree was annotated to include percent consensus out of 1,000 trees with Adobe Illustrator. u
Results
Tonsil, spleen, and lymph node samples of all 3 pigs tested were negative using the following tests: real-time RT-PCR for CSFV, VI for CSFV, ABC immunochemistry assay for CSFV antigens, real-time PCR for ASFV, DFAT for ASFV antigens, and HAD test for ASFV. Brain and lung samples of these pigs were also negative using both the VI for CSFV and the HAD test for ASFV. The EDTA blood samples of all 6 pigs were negative in both the real-time RT-PCR for CSFV and the real-time PCR for ASFV. Serum samples of 4 (pigs 4 and 5) of 6 pigs were negative for antibodies to CSFV assayed by a commercial test kit j and the IP-VN test. The presence of antibodies to CSFV in the 2 sera could be due to the fact that the donor animals were vaccinated against CSFV because CSFV vaccination has been practiced in Haiti in past years. Five serum samples tested were negative for antibodies to ASFV tested by a commercial kit, k and 1 serum sample (from pig 1) was not tested in this assay because the quantity of the serum was not sufficient.
Brain samples of the 3 pigs were positive in the nested RT-PCR for group I PEV (PTV serotypes 1–11) and a separate RT-PCR for PTV serotype 1. Cytopathic effect was also observed in SK-6 and IBRS-2 cells inoculated with filtrates of brain tissue homogenates of these pigs. In addition, brain suspension–inoculated ST cells and pSK cells were positive in the indirect fluorescent antibody test using porcine antiserum known positive for antibodies to PTV serotype 1 as detected with anti-pig IgG conjugated to FITC. Under electron microscopy of cultures of CPE-positive SK-6 cells, spherical nonenveloped virus particles with a mean diameter of 25 ± 1 nm were observed (Fig. 1). The morphology of these virus particles is consistent with that of the family Picornaviridae.
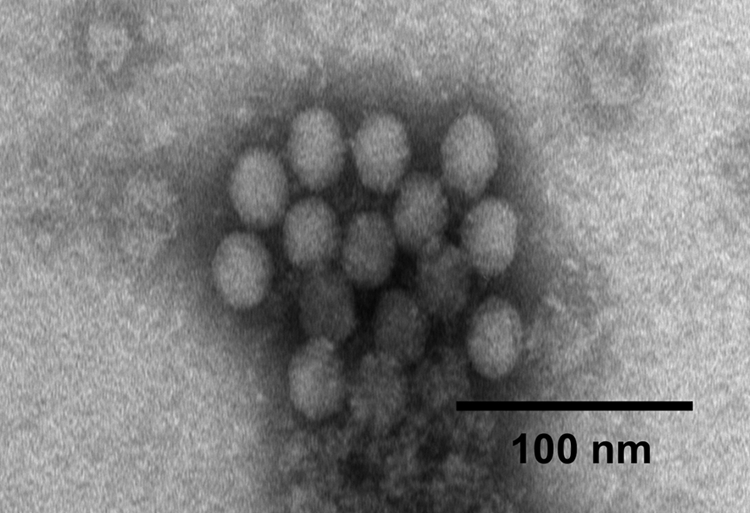
A clump of spherical nonenveloped virus particles with a mean diameter of 25 ± 1 nm isolated from cultures of SK-6 cells showing cytopathic effect. Bar = 100 nm.
In the histopathological analysis, the cerebrum, cerebellum, and brainstem of each of 3 brain samples were examined. The most significant microscopic finding was multifocal nonsuppurative encephalomyelitis. Lesions were characterized by multifocal perivascular cuffing of lymphocytes and plasma cells, as well as meningitis (Fig. 2A), multifocal gliosis (Fig. 2B), satellitosis (Fig. 2C), and neuronal necrosis (Fig. 2D). These changes were most severe in the cerebellum and throughout the brainstem. These results support the diagnosis of teschovirus encephalomyelitis.
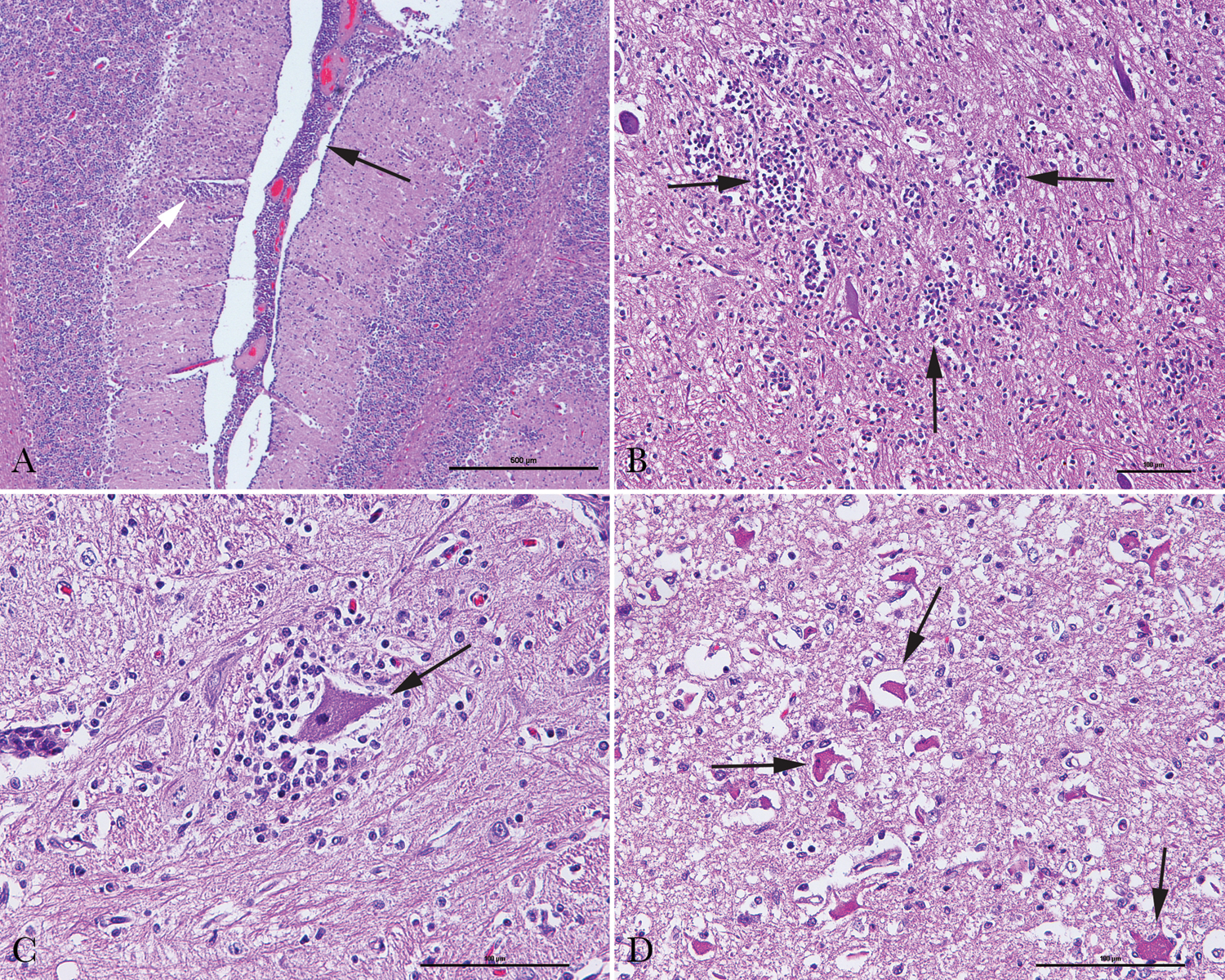
Histopathological analysis of brain tissues of pigs with neurological signs.
The entire genome of the virus isolate was sequenced and ClustalW aligned with nucleotide sequences of other PTV strains available in the GenBank database. The sequence identity between the Haitian isolate and 16 other PTV strains is 80%−86% and 69%−83% for the entire genome and the VP1 region, respectively (Table 1). Results of phylogenetic analysis on the polyprotein of PTV strains indicate that the Haitian isolate is most closely related to other PTV serotype 1 strains (Fig. 3). Among these PTV serotype 1 strains, strain Konratice (AF231768) was isolated in Czechoslovakia from pigs with porcine viral encephalomyelitis (Teschen disease). 3 In a previous study, 13 PTV serotype 1 strains Konratice and Bozen (AF231767) produced a lethal paralytic disease by intramuscular inoculation in pigs. Strain F65 (AJ011380) produced clinical signs resulting from neuroinvasiveness when inoculated into specific–pathogen-free pigs. 4 The Haitian isolate was designated as PTV-1/Haiti/2009. The nucleotide sequence of this isolate was submitted to GenBank with accession no. GQ914053.
Comparison of nucleotide sequences of Haitian isolate* of Porcine teschovirus (PTV) with other PTV strains.
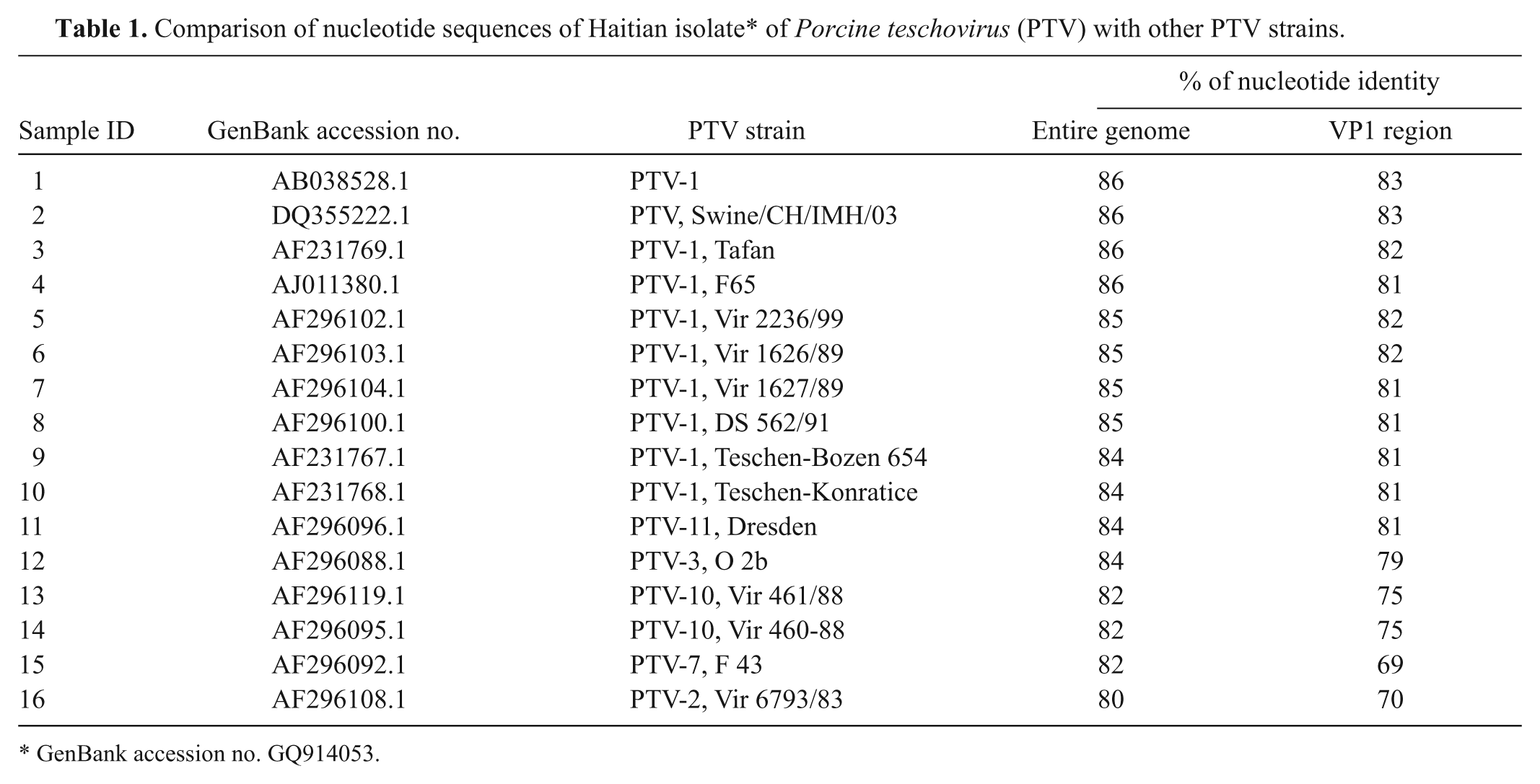
GenBank accession no. GQ914053.
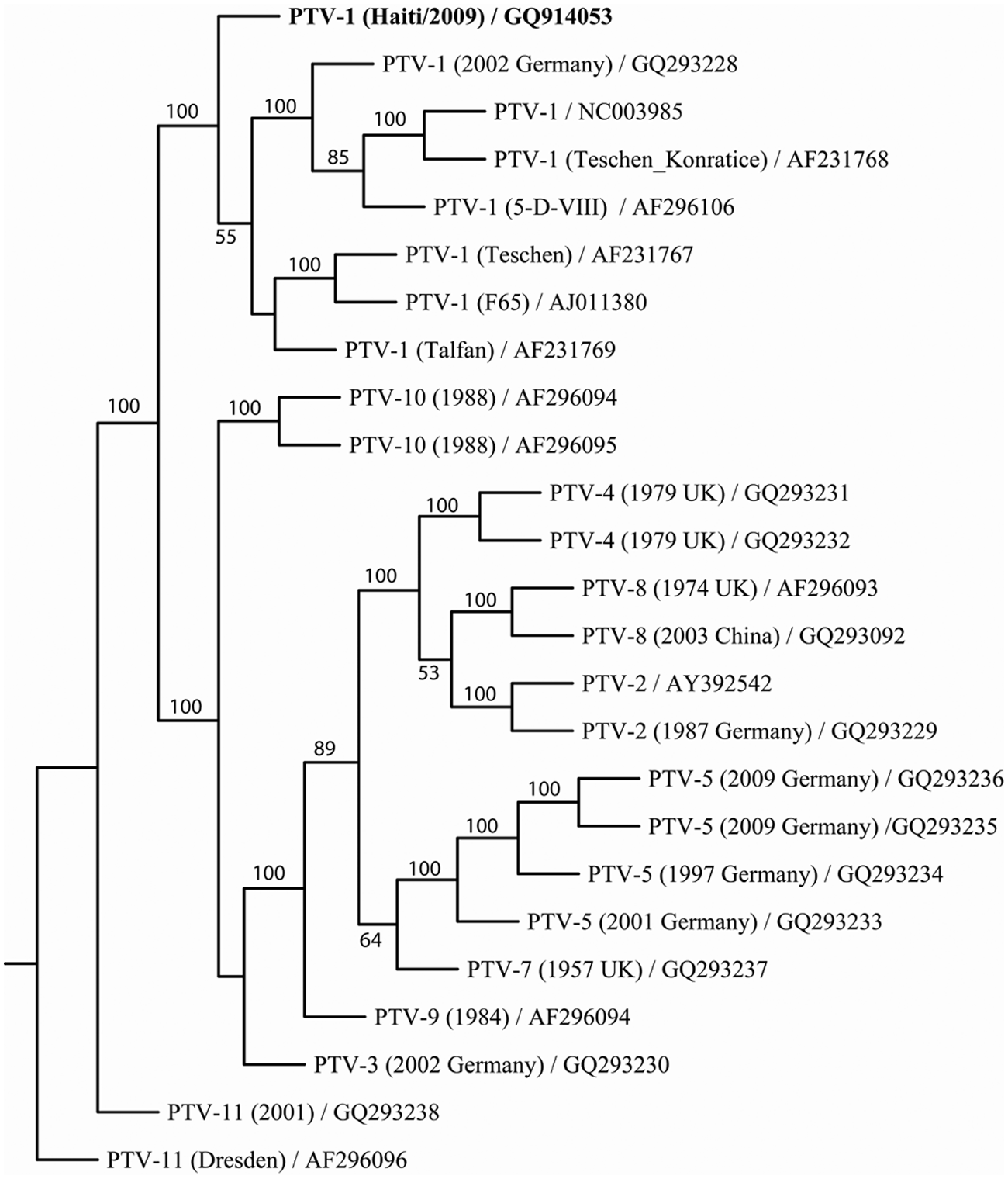
Phylogenetic analysis of the coding sequence for polyprotein of the Porcine teschovirus (PTV) strains. Nucleotide sequences of 25 PTV genomes were trimmed and aligned in ClustalW, and 1,000 bootstrap replicates were subjected to DNA distance and neighbor-joining methods. A consensus tree is shown with bootstrap values greater than 50% placed above tree branches. PTV serotype 1 from Haiti (in bold), while remaining distinct, clustered 100% of the time with other PTV serotype 1 genomes. GenBank accession numbers are indicated in the leaf descriptors on the consensus tree.
Discussion
Based on test results on samples collected from pigs showing neurological signs, the swine disease that occurred in the Lower Artibonite Valley and the Lower Plateau of the Republic of Haiti was diagnosed as teschovirus encephalomyelitis, and the viral isolate was identified as PTV serotype 1. The nonsuppurative encephalomyelitis characterized in the histopathological analysis on brain samples is also supportive of the disease diagnosis.
Nucleotide sequence analysis indicates that the sequence identity between the Haitian isolate and 16 other PTV strains is 80%−86% and 69%−83% for the entire genome and the VP1 region, respectively. The somewhat lower identity for the VP1 region than that of the entire genome is likely due to the relative higher variability of the VP1 region of viruses in the family Picornaviridae. The fact that the Haitian isolate is most closely related to other PTV serotype 1 strains in phylogenetic analysis on the polyprotein of PTV strains further confirms the identification of PTV serotype 1 for the Haitian isolate.
The outbreak of teschovirus encephalomyelitis in Haiti is the first known appearance of the severe form of the disease in the Americas. Factors responsible for the appearance of the disease in Haiti are not certain. In Haiti, no established regulations for the movement of commercial swine between different regions exist at the national level (FAO: 2010, Teschovirus encephalomyelitis in Haiti. FAO Emergency Prevention System for Transboundary Animal and Plant Pests and Diseases. No. 36. Available at: http://www.fao.org/docrep/013/i1958e/i1958e00.pdf). Individual farm operations, however, have the ability to purchase pigs locally and relocate them to their own facility for finishing. This uncontrolled movement of animals facilitates disease transmission. The disease has spread to many regions of the country, including areas near the border with the Dominican Republic. There is the potential for this disease to be introduced to other Caribbean countries, as well as to Central and North American countries by the movement of swine or contaminated products. The development of a vaccine against the disease may be warranted to mitigate risks of disease spread to countries in the Caribbean and other regions.
Footnotes
Acknowledgements
The work of sample submission and diagnosis for teschovirus encephalomyelitis in Haiti was a result of collaboration of several groups, including the Haitian Ministry of Agriculture, the Institute of International Cooperation in Agriculture, APHIS/IS Central America and Caribbean Area Office, and the NVSL. The authors thank Dr. Elizabeth Lautner, Dr. Paul Hauer, Dr. Beverly Schmitt, Tami Beach, Annette Olson, Linda Cox, Heather Petrowski, and many others in USDA/APHIS/NVSL for their support in various phases of this work. Ethan Macnow and Jonathan Artz of Plum Island Animal Disease Center provided assistance in preparing photos for pathological analysis of brain tissues.
a.
Lonza, Walkersville, MD.
b.
Mixer Mill MM 300, Retsch Inc., Newtown, PA.
c.
GE Water and Process Technologies, Trevose, PA.
d.
Bellco, Vineland, NJ.
e.
RNeasy Mini Kit, QIAamp DNA Mini Kit, QuantiTect Multiplex RT-PCR Kit, QuantiTect Master Mix, QuantiTect Multiplex RT Mix; Qiagen Inc., Valencia, CA.
f.
Superscript II Reverse Transcriptase, Platinum PCR SuperMix, 5′ RACE System for Rapid Amplification of cDNA Ends, BigDye Terminator v3.1 Cycle Sequencing Kit; Invitrogen Corp., Carlsbad, CA.
g.
AmpliTaq Gold DNA Polymerase, 7500 Fast Real-Time PCR System, GeneAmp EZ rTth PCR Kit, 3730XL DNA Analyzer; Applied Biosystems, Carlsbad, CA.
h.
DNA engine Tetrad 2 Thermal Cycler, Bio-Rad Laboratories, Hercules, CA.
i.
Cepheid Smart Cycler II, Cepheid Inc., Sunnyvale, CA.
j.
HerdChek CSFV Antibody Test Kit, IDEXX Laboratories Inc., Westbrook, ME.
k.
Ingezim PPA Compac ASF ELISA Kit, Ingenasa, Madrid, Spain.
l.
VECTASTAIN ABC-AP Kit, Vector Laboratories, Burlingame, CA.
m.
Cedi Diagnostics, Lelystad, the Netherlands.
n.
Microtome Cryostat, Thermo Fisher Scientific, Waltham, MA.
o.
Airfuge Air-Driven Ultracentrifuge, Agencourt CleanSEQ system; Beckman Coulter, Brea, CA.
p.
Electron Microscopy Sciences, Hatfield, PA.
q.
Hitachi Electron Microscope T-7600, Hitachi High-Tech, Tokyo, Japan.
r.
Advanced Microscopy Technologies Digital Camera, AMT Software; Advanced Microscopy Technologies, Danvers, MA.
s.
Sequencher, Gene Codes Corp., Ann Arbor, MI.
u.
Adobe Illustrator CS4 version 14.0.0, Adobe Systems, San Jose, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared that they received no financial support for their research and/or authorship of this article.
