Abstract
Moraxella bovoculi was isolated and identified in ocular fluid samples collected from 9 racehorses with infectious keratoconjunctivitis in China in 2013. All 9 M. bovoculi isolates were hemolytic, Gram-negative diplococci that were phenylalanine deaminase positive. The sequence of the 16S ribosomal DNA (rDNA) gene of the isolates matched the 16S rDNA sequence of M. bovoculi. Amplification of the 16S–23S intergenic spacer region followed by AfaI digestion produced a 600–base pair product, a result characteristic of M. bovoculi isolates. The phylogenetic analysis based on the 16S rDNA sequence confirmed the strain isolated in the current study had genetic homology with M. bovoculi.
Keywords
Infectious keratoconjunctivitis (IKC), a highly contagious ocular disease of cattle, results in various degrees of corneal scarring, and can cause permanent blindness following corneal rupture and is a common, economically important, production-limiting disease. 4 Moraxella bovis has been identified as the primary etiological agent of IKC.9,12 However, the recently characterized Moraxella bovoculi has been isolated from corneal ulcers of cattle in the United States 4 and Uruguay14,15 and from reindeer with keratoconjunctivitis in Norway. 16 It was initially reported that M. bovoculi could be distinguished from M. bovis and Moraxella ovis on the basis of the presence of phenylalanine deaminase activity. 4 However, the presence of phenylalanine deaminase activity is now known not to be a consistent feature of M. bovoculi. 2
At the present time, there is only limited information on M. bovoculi, with most of the information coming from the United States.3,7 Although M. bovoculi infection induction studies have failed to document the ability of M. bovoculi to induce IKC ulcers in calves, 8 it is still not known whether M. bovoculi plays a secondary role in the pathogenesis of IKC. Moreover, M. bovoculi can be isolated from eyes of completely normal calves.1,13 The current study reports on the characterization of M. bovoculi isolates obtained from the eyes of racehorses with IKC symptoms in China and the phylogeny of the strain.
During the summer of 2013 on an English Thoroughbred racehorse breeding farm in China, 10 horses from 3- to 9-month-old exhibited corneal ulceration and edema, photophobia, and lacrimation. The 10 swabs of ocular secretions from these IKC-affected horses were plated onto 5% sheep blood agar, and incubated at 37°C. After 48 hr, colonies that exhibited β-hemolysis and were white to off-white, circular, and convex with an entire margin and a moist-looking surface were observed for 9 of the 10 swabs. No other pathogenic bacteria were isolated. The colonies were shown to consist of Gram-negative diplococci. The isolates were subcultured for further biochemical and molecular testing. The 9 isolates were identified asM. bovoculi by a commercial identification system. a All 9 isolates were phenylalanine deaminase positive when tested as previously described. 4
A microdilution system was used for performing antimicrobial sensitivity tests based on the methodology from the Clinical Laboratory Standards Institute (CLSI). 5 The isolates were streaked onto trypticase soy plates containing 5% sheep blood cells and incubated at 37°C for 24 hr. Two to three isolated colonies were used to inoculate 2 ml of brain–heart infusion broth b and incubated at 37°C without carbon dioxide for 6 hr. This broth was then used to inoculate the commercial microdilution system. a The minimum inhibitory concentration was determined for 15 antimicrobial agents. The 9 isolates were all sensitive to oxytetracycline (≤0.12 µg/ml), ceftiofur (≤0.12 µg/ml), florfenicol (≤0.25 µg/ml), penicillin (≤0.25 µg/ml), and resistant to cloxacillin and trimethoprim–sulfamethoxazole (>256 µg/ml) following the guidelines of the CLSI. 5
Molecular identification by sequence analysis of the 16S ribosomal DNA (rDNA) and the internal transcribed spacer (ITS) region between the 16S and 23S rRNA genes was used in this study. Genomic DNA was isolated using a genomic DNA extraction kit. c A polymerase chain reaction (PCR) assay was performed using 2.5 mM deoxyribonucleotide triphosphate, 10 μM of each primer, 2 μg of DNA, and 1 U of Taq DNA polymerase. d The cycling reactions were 32 cycles of 1 min at 94°C, 90 sec at 53°C, and 2 min at 72°C. The 16S rRNA gene was amplified using universal primers for bacteria 27f (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492r (5′-GGTTACCTTGTTACGACTT-3′). The products were sequenced, and the initial sequence identification was conducted by BLAST. According to nucleotide–nucleotide GenBank search (http://blast.ncbi.nlm.nih.gov), the sequence was 99% identical to the sequences of 16S rDNA of M. bovoculi. Moreover, the results of the sequencing of the 16S rDNA indicated that the 9 isolates were identical and showed 99% homology with the M. bovoculi type strain 237 (GenBank accession no. NR043583). A 600–base pair product was obtained following amplification of the ITS region coupled with restriction enzyme AfaI e digestion analysis, a result that is typical of M. bovoculi as previously described.2,14
Sequence assembly was carried out using commercial software. f The sequence of the 16S rRNA gene of the isolated strain was deposited in GenBank (KF637298). Multiple sequence alignments based on the 16S rRNA gene sequence of the strain in the current study and other reference strains in the database were conducted with ClustalW (http://www.clustal.org/). Phylogenetic analysis was performed based on the 16S rRNA gene sequence by the neighbor-joining (NJ) method. The NJ tree was constructed with MEGA5.2 software, and the robustness of the phylogenetic analysis was determined by bootstrap analysis with 1,000 replications (http://www.megasoftware.net). The phylogenetic tree showed that M. bovis and M. bovoculi isolates grouped in different branches of the dendrogram, with both being clearly differentiated from the M. ovis reference strain (Fig. 1).
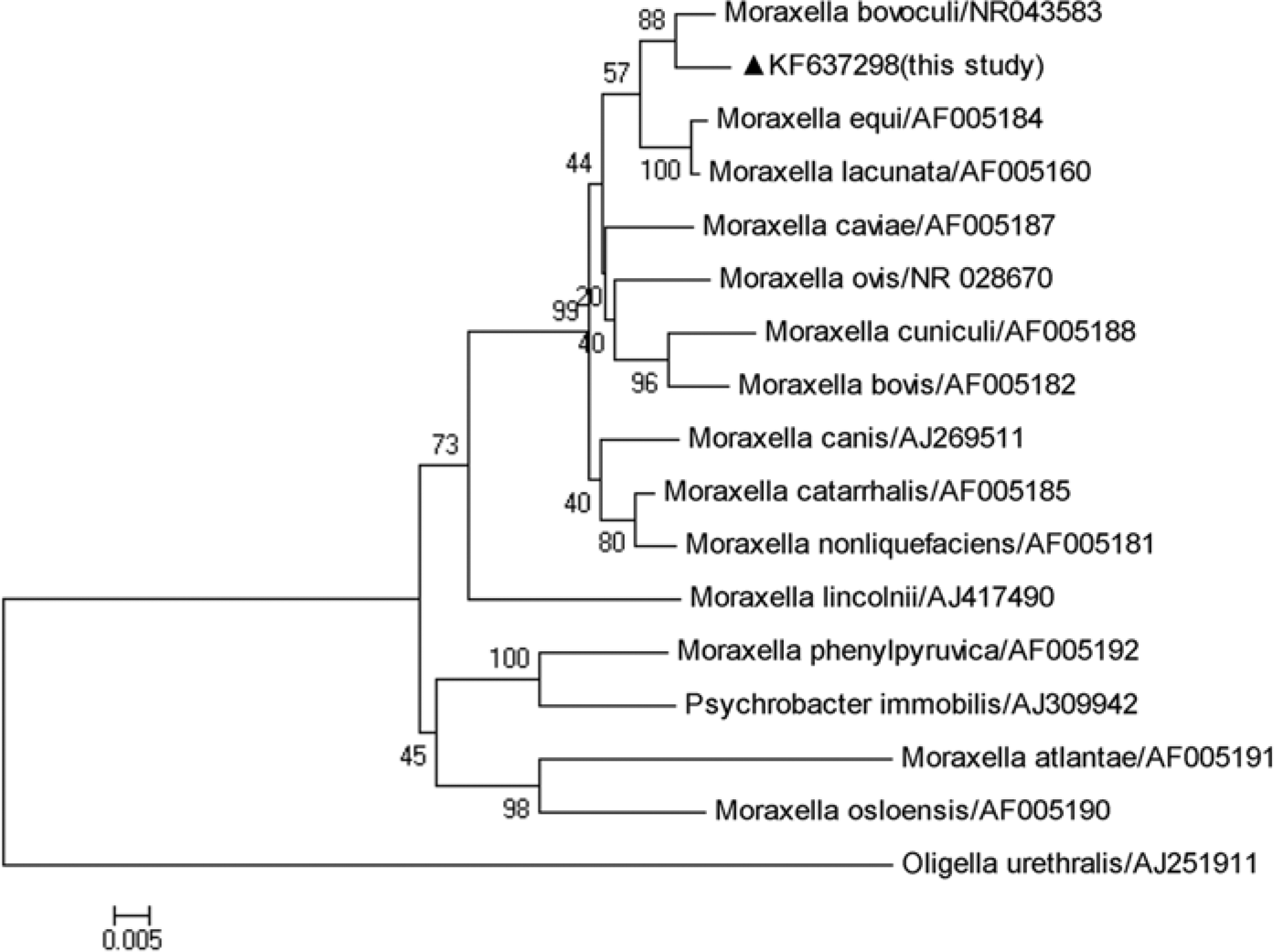
The phylogenetic tree of the genus Moraxella created by cluster analysis (using MEGA version 5.2) of 16S ribosomal RNA gene sequences. Oligella urethralis served as an outgroup. Bootstrap values obtained from 1,000 replicates are shown at the major nodes. The isolate identified in the current study is indicated by a solid triangle.
Conjunctivitis is considered to be one of the more common ocular conditions seen in horses, and previous reports have described the isolation of Moraxella sp. that serologically and biochemically resemble M. bovis from the eyes of horses affected with conjunctivitis.10,11 The results of the present study showed that the isolate from racehorses constituted a branch with M. bovoculi. Moreover, the isolate could be distinguished from the nearest phylogenetic relatives, M. equi, M. bovis, and other related strains of the genus Moraxella. The isolation of M. bovoculi, as reported in the current study, indicates that this species may be an important cause of IKC in Chinese horses.
An autogenous vaccine made from M. bovoculi isolates from active infectious bovine keratoconjunctivitis (IBK) cases has been shown to be ineffective as a management tool for preventing or reducing the incidence of IBK. 6 In the present study, application of oxytetracycline ophthalmic ointment resulted in recovery of horses affected with conjunctivitis.
In summary, the current study reported the isolation ofM. bovoculi associated with IKC in horses in China, and provided some information on antibiotic treatment of equine IKC from which M. bovoculi was isolated. Further research is required to establish the impact that these findings will have on IKC control, particularly vaccination.
Footnotes
a.
Phoenix 100 identification/susceptibility system, BD, Franklin Lakes, NJ.
b.
Difco Laboratories Inc., Detroit, MI.
c.
DNeasy blood & tissue kit, Qiagen GmbH, Hilden, Germany.
d.
rTaq DNA polymerase, Invitrogen Corp., Carlsbad, CA.
e.
QuickCut, Takara Biotechnology (Dalian) Co. Ltd., China.
f.
SeqMan program, DNASTAR Inc., Madison, WI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by grants to the Wildlife-borne Diseases Surveillance Project (31072126, 31101806), National Science and Technology Ministry (2013BAD12B00), and the Knowledge Innovation Program of the Chinese Academy of Sciences (grant KSCX2-EW-J-2).
