Abstract
Nonhemolytic isolates of catalase-positive, Gram-positive cocci were observed that produced strong synergistic hemolysis with colonies of Staphylococcus pseudintermedius and Staphylococcus schleiferi subsp. coagulans in primary cultures of clinical specimens from dogs. The isolates appeared similar to nonhemolytic staphylococci but were not identified with conventional biochemical tests or commercial biochemical test kits. Partial 16S ribosomal RNA gene sequences from each of 9 tested isolates were nearly identical to that of Rothia nasimurium. The isolates produced positive Christie–Atkins–Munch–Peterson test reactions with Staphylococcus aureus (ATCC strain 25923) and Staphylococcus intermedius Hajek (ATCC strain 49052). The latter was identified in the current study as S. pseudintermedius. Rothia nasimurium appears to be a previously unreported commensal organism in dogs. Its potential significance as an opportunistic contributor to polymicrobial infections in dogs is not known.
Keywords
Synergistic hemolytic reactions are occasionally observed with Staphylococcus pseudintermedius in mixed bacterial cultures of clinical specimens from dogs. In many cases, the bacterium paired with Staphylococcus to result in synergistic hemolysis is thought to be clinically insignificant and is not identified to genus and species level. Christie–Atkins–Munch–Peterson (CAMP)–like synergistic hemolysis between β-hemolysin-producing isolates of Staphylococcus aureus or S. pseudintermedius and soluble factors produced by other weakly or nonhemolytic bacteria are well known. 15 The CAMP test has been useful for presumptive identification of pathogens such as Streptococcus agalactiae, Listeria monocytogenes, and Rhodococcus equi. 15 Similar synergistic hemolysis has been shown with common human skin commensals including Staphylococcus spp., Corynebacterium spp., Propionibacterium spp., Actinomyces neuii, and Turicella otitidis.8,12,15 Some group G Streptococcus isolates from dogs may also produce a CAMP-like reaction. 11 Two organisms, identified as coagulase-negative, Staphylococcus warneri–like and Micrococcus lylae, produced synergistic hemolysis with colonies of S. pseudintermedius in a primary culture from skin of a dog. 10 The present study describes the identification and characterization of Rothia nasimurium as an additional organism that produces strong synergistic hemolysis with Staphylococcus colonies in primary cultures of clinical specimens from dogs.
Bacterial isolates were from aerobic cultures performed on clinical specimens submitted to the University of Tennessee College of Veterinary Medicine, Clinical Bacteriology and Mycology Laboratory (Knoxville, Tennessee) from March 2011 to April 2013. Primary plates consisting of Columbia agar a with 5% sheep blood, b incubated at 35°C in an atmosphere of 7% CO2 for 24 hr, from cultures performed on samples from dogs, were inspected for pairs of colonies that produced strong synergistic hemolysis with suspected β-lysin–producing staphylococci (Fig. 1). Colonies showing such hemolysis from 9 unrelated clinical specimens were isolated in pure culture and saved for subsequent identification. Beta-lysin-producing staphylococci were identified by procedures that have been described for identification of common canine Staphylococcus species (e.g., S. pseudintermedius, S. schleiferi subsp. coagulans, and S. aureus).4,5 Briefly, these procedures included a short battery of conventional tests (hemolysis pattern, Gram stain reaction, and morphology; catalase, coagulase, acetoin production; and maltose, lactose, and trehalose fermentation), followed by polymerase chain reaction (PCR) amplification of a portion of the phosphate acetyltransferase gene (pta) and its subsequent direct DNA sequencing or restriction endonuclease fragment length polymorphism (RFLP) analysis using the restriction endonuclease, MboI. 2 Eight of the nine staphylococcal isolates were identified as S. pseudintermedius and 1 as S. schleiferi subsp. coagulans.
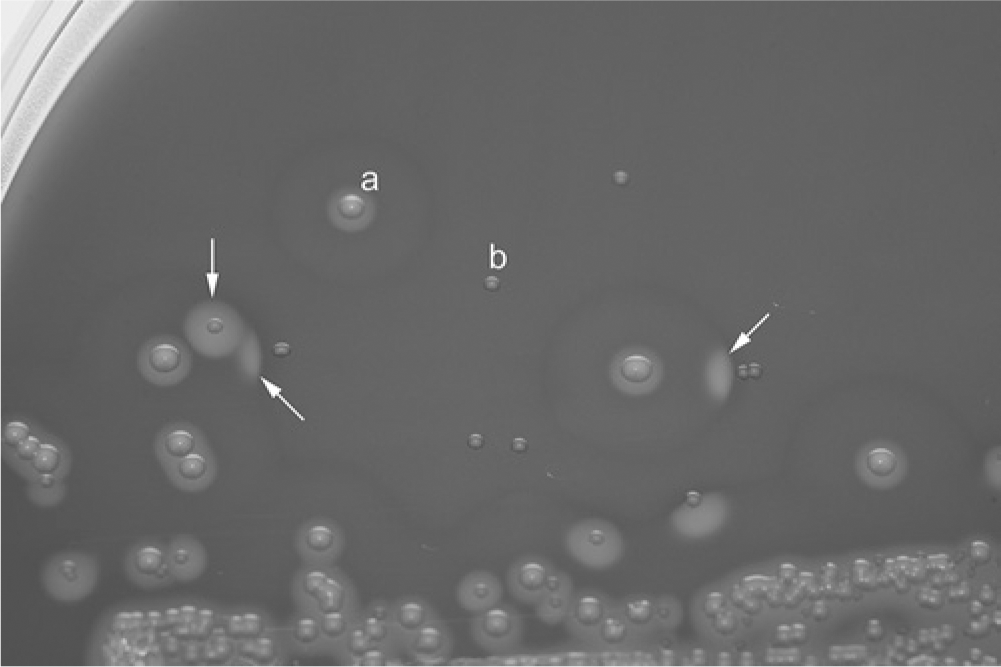
Primary blood agar plate showing synergistic hemolysis (arrows) between (
Isolates that produced synergistic hemolysis with Staphylococcus colonies on primary plates were initially examined for their colony characteristics, microscopic Gram stain appearance, catalase test reaction, and CAMP test reaction with S. aureus (American Type Culture Collection [ATCC] strain 25923) and S. intermedius Hajek (ATCC strain 49052). c The latter, originally isolated from a human blood culture, was identified in the current study by pta gene PCR/MboI RFLP 2 and thermonuclease gene (nuc) sequence analysis 13 as S. pseudintermedius. The isolates produced smooth, gray-white, nonhemolytic colonies, ≥1.0 mm in diameter, at 24 hr on blood agar. Microscopically, after 24-hr growth in brain-heart infusion broth a at 35°C, the isolates appeared as Gram-positive cocci, with diameters ≥1.0 µm, arranged in tetrads, clusters, and pairs (Fig. 2). Each of the R. nasimurium isolates produced a strong zone of CAMP-like enhanced hemolysis when cross-streaked with both S. aureus (ATCC strain 25923) and S. intermedius Hajek (ATCC strain 49052) on blood agar (Fig. 3). The isolates were catalase-positive following overnight growth on brain-heart infusion agar, a coagulase-negative at 24 hr, did not produce acetoin after 48 hr, and were modified oxidase test d negative. Maltose fermentation reactions were negative on bromocresol purple maltose agar a at 24 hr, but for 8 of the 9 isolates, maltose, lactose, and trehalose fermentation reactions were positive at 48 hr in phenol red broths a containing these sugars. The isolates produced biocodes 3761773122 and 2760773122 with a commercial biochemical test kit, e neither of which resulted in acceptable identifications. In an automated biochemical identification system, f the isolates produced biocode 061032301241031, which gave very good identification (95% probability) of Globicatella sanguinis and biocode 061132305261031, which gave an acceptable identification (87% probability) of Kocuria kristinae. The difference in these 2 biocodes was in variable reactions for lactose fermentation and growth in 6.5% NaCl. Neither commercial database contained biocodes for R. nasimurium. Positive test results included the following: 4-methylumbelliferyl-β-D-glucosidase, L-phenylalanine arylamidase, 4-methylumbelliferyl-α-D-glucosidase, L-pyroglutamic acid arylamidase, L-tryptophan arylamidase, L-arginine arylamidase, methyl-α and β-glucosidase, proline and leucine arylamidase, alanine and phenylalanine and proline arylamidase, leucine arylamidase, alanine arylamidase, β-galactosidase, L-pyrrolydonyl arylamidase, D-trehalose, sucrose, D-mannitol, maltotriose, glycerol, fructose, maltoside, esculin, D-ribose, optochin resistance, D-mannose, and D-maltose. Negative test results were observed for the following: phosphatase, β-D-glucuronidase, p-nitrophenyl-β-D-glucosidase, α-D-galactosidase, L-aspartate arylamidase, β-galactopyranosidase, α-mannosidase, urease, pholphatidyl inositol phospholipase C, arginine dihydrolase, L-lactate alkalinization, novobiocin resistance, polymyxin B resistance, O129 R resistance, bacitracin resistance, D-raffinose, cyclodextrin, arabinose, D-amygdalin, N-acetyl-β-D-glucosamine, β-D-cellobiose, D-xylose, D-sorbitol, salicin, D-galactose, and pullulan. Variable test results were observed for L-valine arylamidase, L-isoleucine arylamidase, growth in 6.5% NaCl, and lactose.
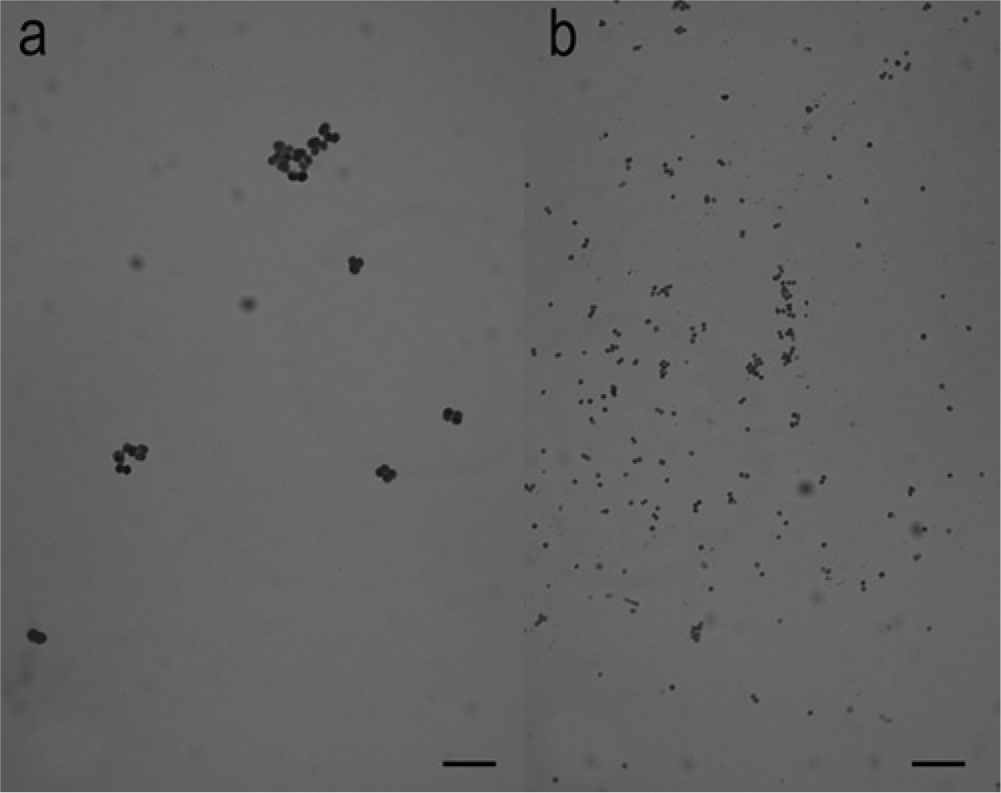
Gram stain microscopic appearance of Rothia nasimurium (
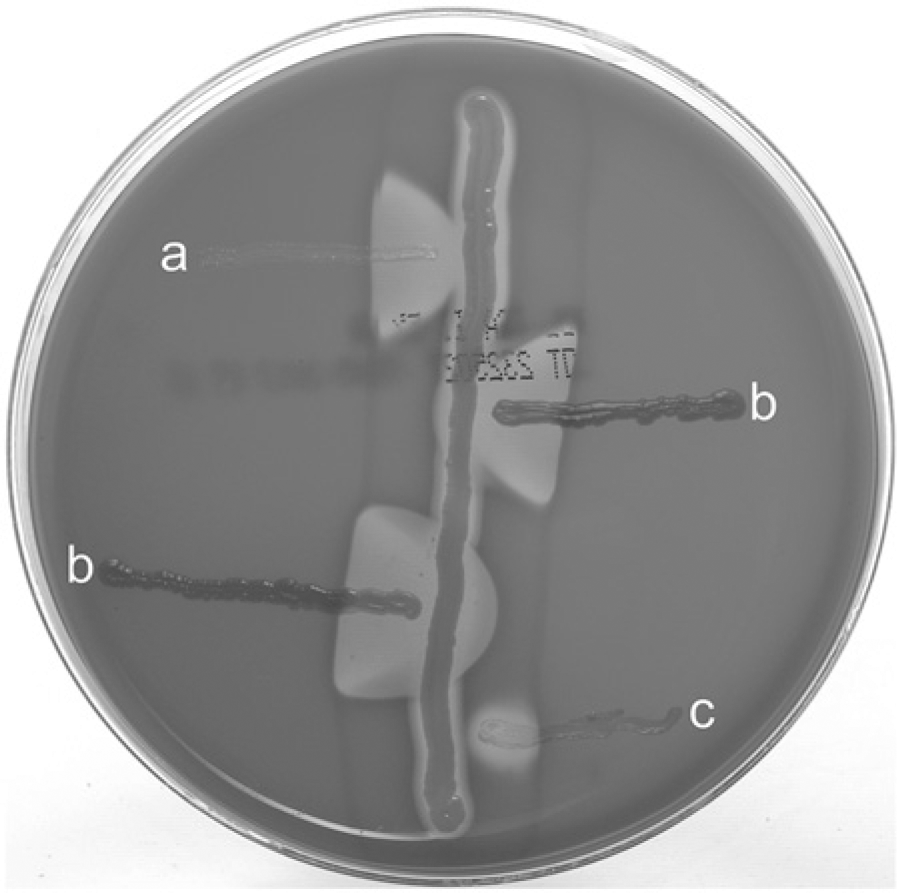
CAMP test among (center streak) Staphylococcus aureus ATCC strain 25923, (
DNA was extracted from the isolates by physical disruption with 0.1-mm glass beads, g and a portion of the 16S ribosomal RNA (rRNA) gene was amplified by PCR with eubacterial primers Broad 1 and Broad 2. 14 The PCR mix was treated h to remove single-stranded DNA and submitted to the University of Tennessee Molecular Biology Resource Facility for bidirectional sequencing with the PCR primers. The 16S rRNA gene sequences of the isolates were edited and aligned using commercial software i and compared to sequences in the GenBank database using the BLASTn algorithm. 1 The isolates that paired with Staphylococcus to produce strong synergistic hemolysis were identified as most closely resembling the type strain of R. nasimurium by partial 16S rRNA gene sequencing (fragment sizes ranging from 1366 bp to 1421 bp, with 100% query coverages, 0 expect values, and 99% identities with GenBank accession no. AJ131121). 6 DNA sequence similarities among the canine Rothia isolates were from 99.4% to 100%. Although sequences from the canine Rothia isolates were close enough in similarity to be included in the R. nasimurium species, they were more closely related to each other than to that of the R. nasimurium type species and were more distantly related to that of an outlier species, Rothia mucilaginosa (Fig. 4).
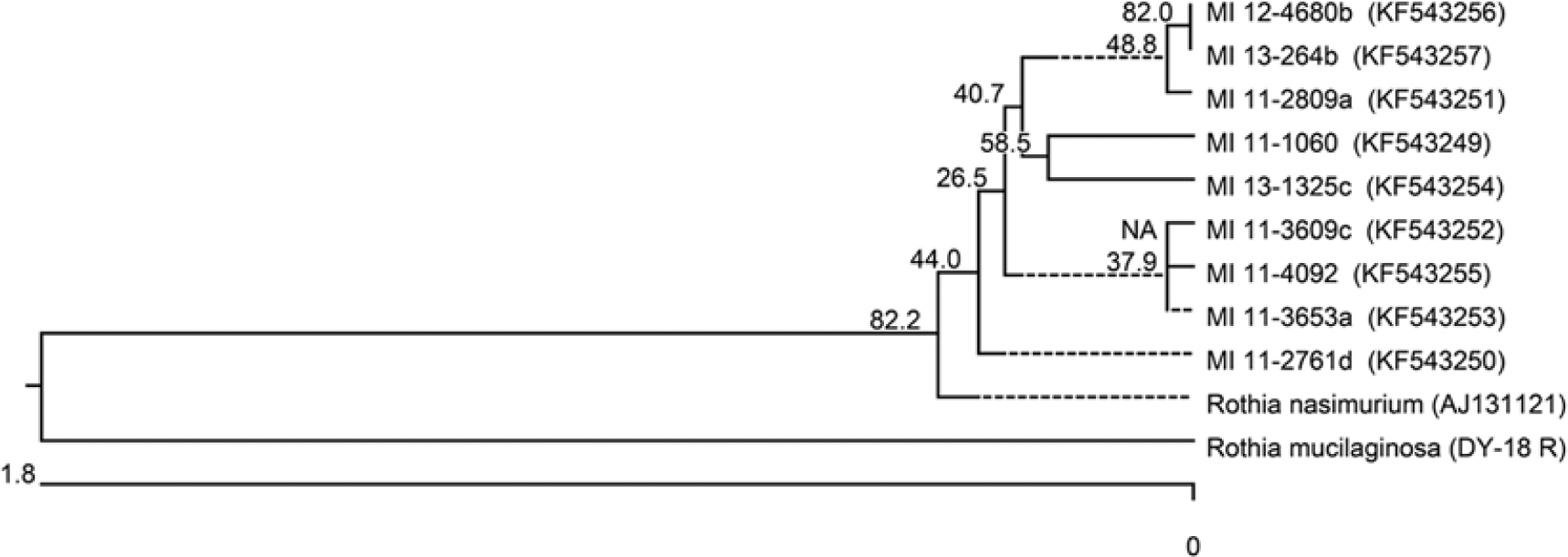
16S ribosomal RNA gene sequence–based phylogenetic tree of canine Rothia isolates and related GenBank entries from the type strain of Rothia nasimurium and strain Rothia mucilaginosa. Sequence accession numbers are given in parentheses. The tree was generated by the neighbor-joining method, and the bootstrap values, shown at the branches, were calculated from 1,000 replicates. Evolutionary analyses were conducted in Molecular Evolutionary Genetics Analysis (MEGA) software version 5.0. j
Rothia nasimurium, or a very closely related species, appears to be a commensal organism found in dogs. The isolates identified in the present study were from superficial body sites. Seven samples were from skin lesions, 1 was from the external ear canal, and 1 was from seminal fluid. Published descriptions of R. nasimurium are based on a single isolate from a mouse. 6 Phenotypic characteristics and 16S rRNA gene sequences observed in the current study were consistent with those described for the type strain of R. nasimurium (Culture Collection University of Goteborg strain [CCUG] 35957); however, little is known about phenotypic strain variation within the species, its genome sequence, or its host distribution among different animals. In a previous report, an isolate obtained from either canine dental plaque or saliva was assigned “a most likely estimated genus identification” as Rothia based on 16S rRNA sequence analysis, and closest identity (92.7%) of the 473-bp fragment was with sequence from R. nasimurium. 7 There are also unpublished descriptions of 3 isolates from the tonsils of healthy dogs listed as R. nasimurium in the online catalog of a biological repository. k Each of these had similar cell fatty acid profiles and 1 isolate had a 750-bp fragment rRNA gene sequence with 99% identity to that of the type strain of R. nasimurium. Minor variations in biochemical profile as observed in the current study and the inability to identify R. nasimurium with commercial biochemical test systems place a great deal of weight on the results of 16S partial rRNA gene sequencing to confirm the identity of clinical isolates. Additional phenotypic and genotypic markers are needed to facilitate identification and determine the prevalence and body site distributions of R. nasimurium in dogs.
It is not known if the R. nasimurium isolates reported herein were normal flora contaminants or opportunistic pathogens involved in polymicrobial infections. All specimens were from clinically infected body sites; however, in 5 of the 9 cultures, 3 or more bacterial or fungal species (most of which could also be considered as commensals, opportunists, or contaminants) were isolated. Bacterial species from several different genera, including Staphylococcus, Micrococcus, Streptococcus, Listeria, Rhodococcus, Corynebacterium, Arcanobacterium, Actinomyces, Turicella, Trueperella, and Propionibacterium, produce CAMP-like synergistic hemolysis on sheep blood agar with β-lysin–producing Staphylococcus species.3,8-12,15 The current study demonstrates that R. nasimurium can be added to such a list. Rothia species isolated from dogs have received little attention in the veterinary literature. Their role as commensals or as opportunistic pathogens in mixed infections with Staphylococcus spp. merits further investigation. The strength of hemolysis enhancement produced by the canine R. nasimurium isolates was equal to or greater than that observed with a control strain of Streptococcus agalactiae. In a mouse model, a CAMP-like factor produced by Propionibacterium acnes enhanced the cytolytic effects of S. aureus on macrophage and exacerbated S. aureus–induced skin lesions. 12 From the present study, it was concluded that R. nasimurium is a common inhabitant of dogs and that synergistic hemolysis with S. pseudintermedius is a characteristic feature of canine R. nasimurium isolates. For the veterinary diagnostician, the presence of R. nasimurium colonies in a confluence of growth with a β-lysin–producing Staphylococcus on blood agar could mistakenly prompt futile attempts to isolate and identify a presumed β-hemolytic colony. Conversely, if present in the absence of a β-lysin-producing Staphylococcus, R. nasimurium colonies could be presumptively identified as those of coagulase-negative Staphylococcus-like organisms, and the prevalence of R. nasimurium in clinical samples from dogs would remain unknown. Screening of nonhemolytic Staphylococcus-like colonies using the CAMP test may provide an initial step for preliminary identification of R. nasimurium isolates from dogs.
Footnotes
Acknowledgements
The authors thank Brian Johnson, Rebekah Jones, Matthew Riley, Anik Vasington, and Phillip Snow for their technical assistance.
a.
BBL, BD Diagnostic Systems, Sparks, MD.
b.
Hemostat Laboratories, Dixon, CA.
c.
American Type Culture Collection, Manassas, VA.
d.
Remel Microdase disk, Thermo Fisher Scientific, Waltham, MA.
e.
BBL Crystal identification systems gram-positive ID kit, BD Diagnostic Systems, Sparks, MD.
f.
Vitek GPS card, Vitek 2 automicrobic system; bioMérieux Inc., Durham, NC.
g.
BioSpec Products Inc., Bartlesville, OK.
h.
ExoSAP-IT, USB Corp., Cleveland, OH.
i.
Lasergene version 7.2, DNASTAR Inc., Madison, WI.
k.
Strains CCUG 46188, CCUG 46190, and CCUG 46343, Culture Collection University of Goteborg, Goteborg, Sweden.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported in part by a grant from the Center of Excellence in Livestock Diseases and Human Health, University of Tennessee.
