Abstract
Approximately 8,000 isolates of
Keywords
Introduction
The U.S. Food and Drug Administration (FDA) has published guidelines for dairy cattle veterinarians on the judicious use of antimicrobial drugs (U.S. FDA/CVM: 2002, Judicious use of antimicrobials for dairy cattle veterinarians. Available at: http://www.fda.gov/downloads/AnimalVeterinary/SafetyHealth/AntimicrobialResistance/JudiciousUseofAntimicrobials/UCM095571.pdf) as has the Canadian Veterinary Medical Association. 5 Responsible stewardship involves activities that aim to minimize the emergence and spread of antimicrobial resistance and preserve the efficacy of current veterinary drugs. Surveillance for antimicrobial susceptibility among pathogens is an important component of stewardship.3,18 Data regarding loss of in vitro susceptibility in pathogens can be used by veterinarians, livestock producers, pharmaceutical companies, and others working to support the health and welfare of animals. It has been noted that “Access to timely and accurate antimicrobial resistance data may also help to prevent inappropriate responses to anecdotal reports of resistance, which can unfairly bias the perceived efficacy of an agent, complicate antibiotic strategies and confuse laboratory testing efforts.” 11
There are few publications of national surveys of the antimicrobial susceptibilities of pathogens isolated from dairy cattle on farms in the United States or Canada.2,3 To address this lack of data, the animal health business unit of Pfizer Inc. (currently operating as Zoetis™) established a Mastitis Pathogen Antimicrobial Susceptibility Surveillance Program in 2002 to monitor the susceptibility of the major mastitis pathogens to antimicrobial drugs used for treatment of mastitis in North America. The objective of the program was to monitor bovine mastitis pathogens that had been isolated from sick animals and report on minimal inhibitory concentration (MIC) distributions over time. The microorganisms tested in the program,
Materials and methods
Isolates
Between 2002 and 2010, 25 veterinary laboratories submitted isolates of
Isolate origin and number of isolates per year by region for a 9-year (2002–2010) survey of antimicrobial susceptibility of bacteria that cause bovine mastitis in the United States and Canada.*
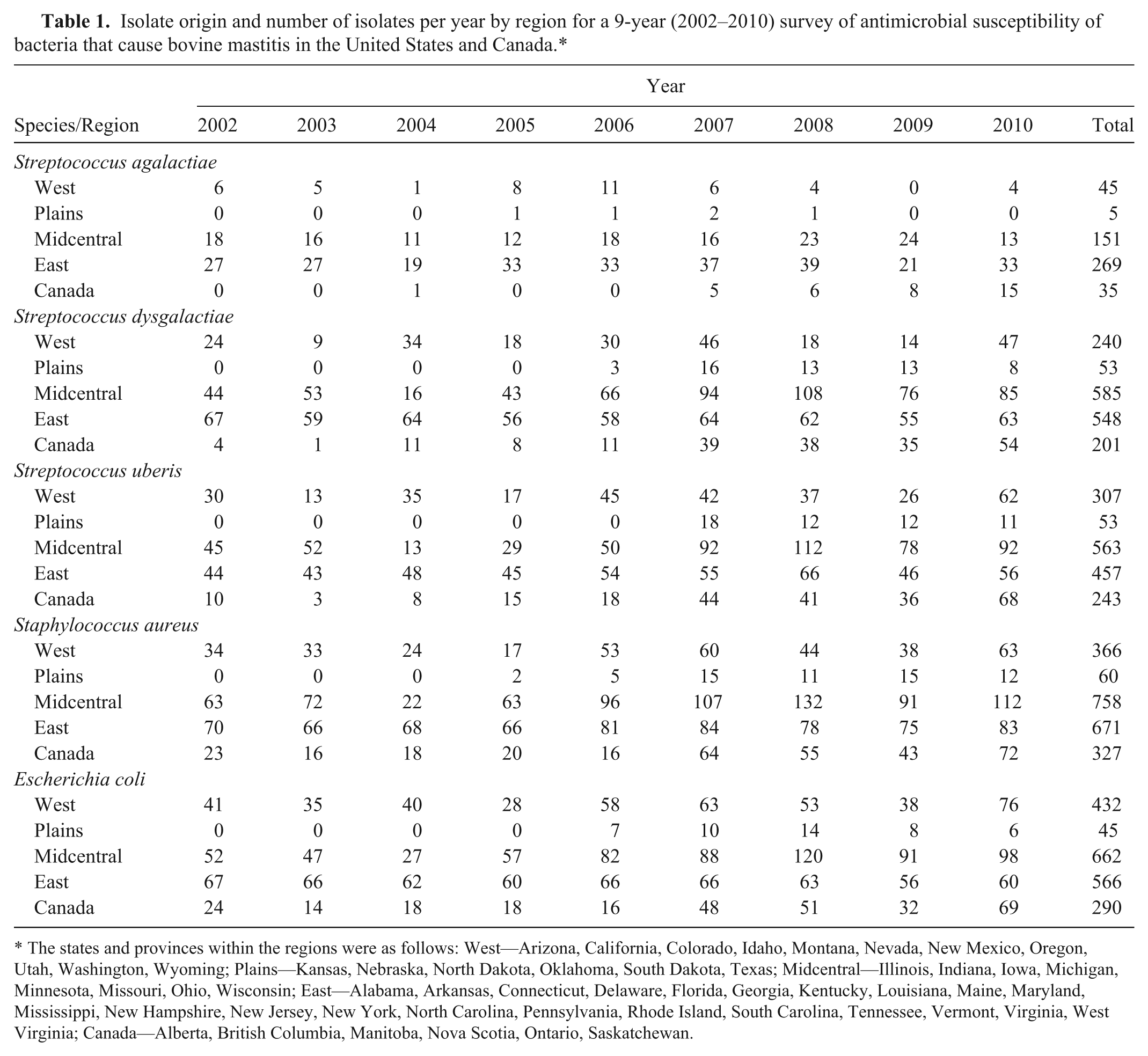
The states and provinces within the regions were as follows: West—Arizona, California, Colorado, Idaho, Montana, Nevada, New Mexico, Oregon, Utah, Washington, Wyoming; Plains—Kansas, Nebraska, North Dakota, Oklahoma, South Dakota, Texas; Midcentral—Illinois, Indiana, Iowa, Michigan, Minnesota, Missouri, Ohio, Wisconsin; East—Alabama, Arkansas, Connecticut, Delaware, Florida, Georgia, Kentucky, Louisiana, Maine, Maryland, Mississippi, New Hampshire, New Jersey, New York, North Carolina, Pennsylvania, Rhode Island, South Carolina, Tennessee, Vermont, Virginia, West Virginia; Canada—Alberta, British Columbia, Manitoba, Nova Scotia, Ontario, Saskatchewan.
The isolates had been recovered from either clinical or subclinical cases of bovine mastitis. The isolates were selected by the laboratories, which had been requested not to use susceptibility as a criterion for submission. Each laboratory was limited to the number of isolates (typically less than 25 isolates of each species) that could be submitted each year, so that there was not an overrepresentation from any geographic region. Laboratories were discouraged from submitting multiple isolates of the same bacterial species from an individual herd to prevent testing of epidemiologically related strains. The majority of the isolates came from animals within the same state or province of the submitting laboratory; however, isolates were also submitted from additional states or provinces. Isolates were identified by the submitting laboratory before shipment to Zoetis. Confirmation or further characterization of isolates, when necessary, was conducted using standard biochemical tests and commercially available identification systems.a,b
Minimal inhibitory concentration determinations
Minimal inhibitory concentrations for all isolates were determined using a broth microdilution system, c which conforms to the standards of the Clinical and Laboratory Standards Institute (CLSI) for testing veterinary pathogens. 6 Antimicrobial agents were selected either as representatives of their antimicrobial class for the purposes of susceptibility testing or based on interest and availability in commercial, approved intramammary preparations. The custom 96-well microtiter panels included doubling dilutions of ampicillin, ceftiofur, cephalothin, erythromycin, oxacillin, the combination drug penicillin–novobiocin, and pirlimycin. Concentration ranges for each antimicrobial drug were chosen to encompass appropriate CLSI quality control ranges and clinical breakpoints wherever possible. 6
All MIC testing was done at 1 of 2 laboratories (Zoetis, Kalamazoo, Michigan or Microbial Research Inc., Fort Collins, Colorado). Direct colony suspensions were used when testing all organisms, and suspensions were prepared to yield a final concentration of bacteria of approximately 5 × 105 colony-forming units/ml. The growth medium for all streptococcal isolates was cation-adjusted Mueller–Hinton broth supplemented with 2.5–5% lysed horse blood. All other isolates were tested with unsupplemented Mueller–Hinton broth. Both laboratories adhered strictly to CLSI standardized methods, including performance of quality control tests. 6
Results
Between 2002 and 2010, 505 isolates of
The distribution frequencies of ampicillin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
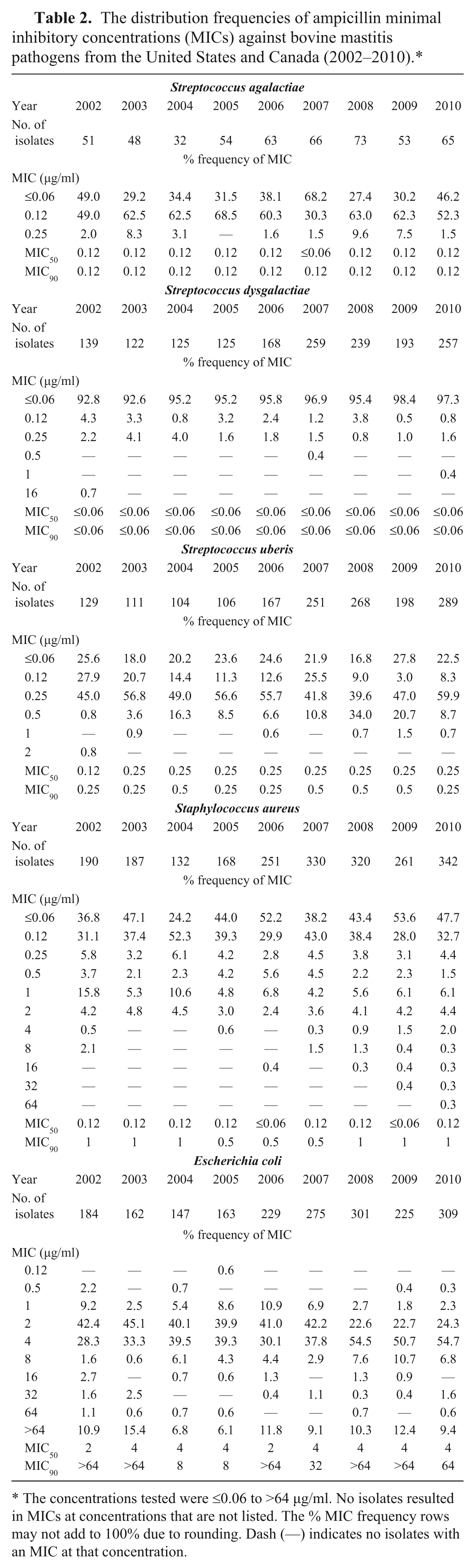
The concentrations tested were ≤0.06 to >64 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration.
The distribution frequencies of ceftiofur minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*

The concentrations tested were ≤0.06 to >64 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration. The CLSI-VAST–approved upper limit for susceptibility was 2 µg/ml, and the lower limit for resistance was 8 µg/ml for ceftiofur against
The distribution frequencies of cephalothin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
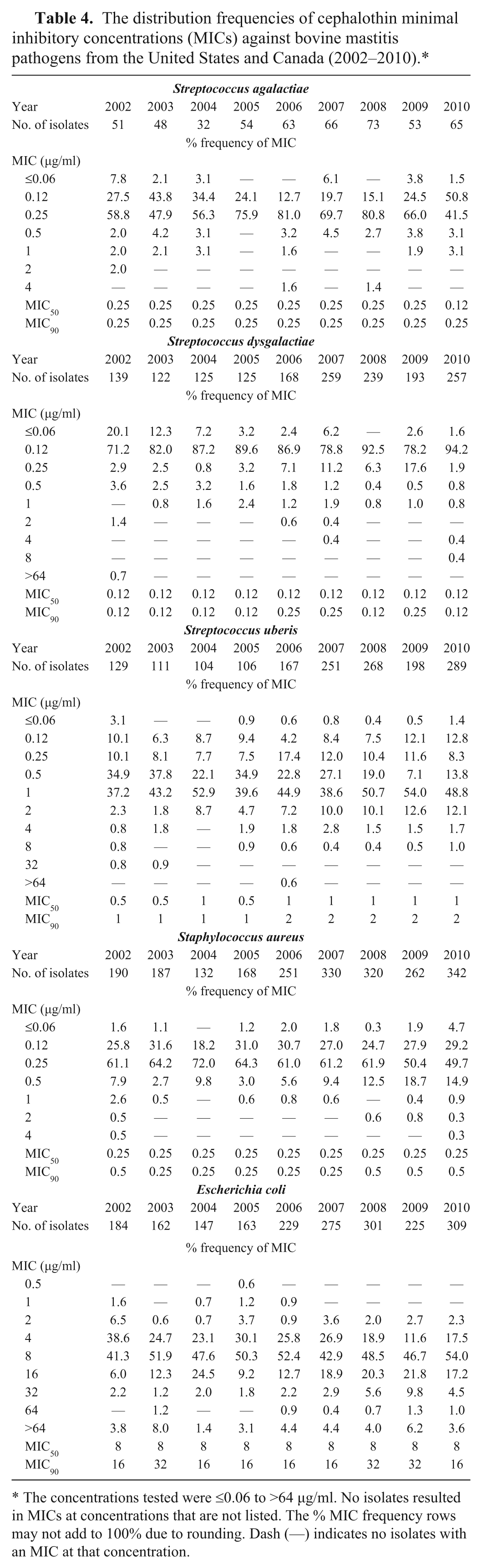
The concentrations tested were ≤0.06 to >64 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration.
The distribution frequencies of erythromycin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
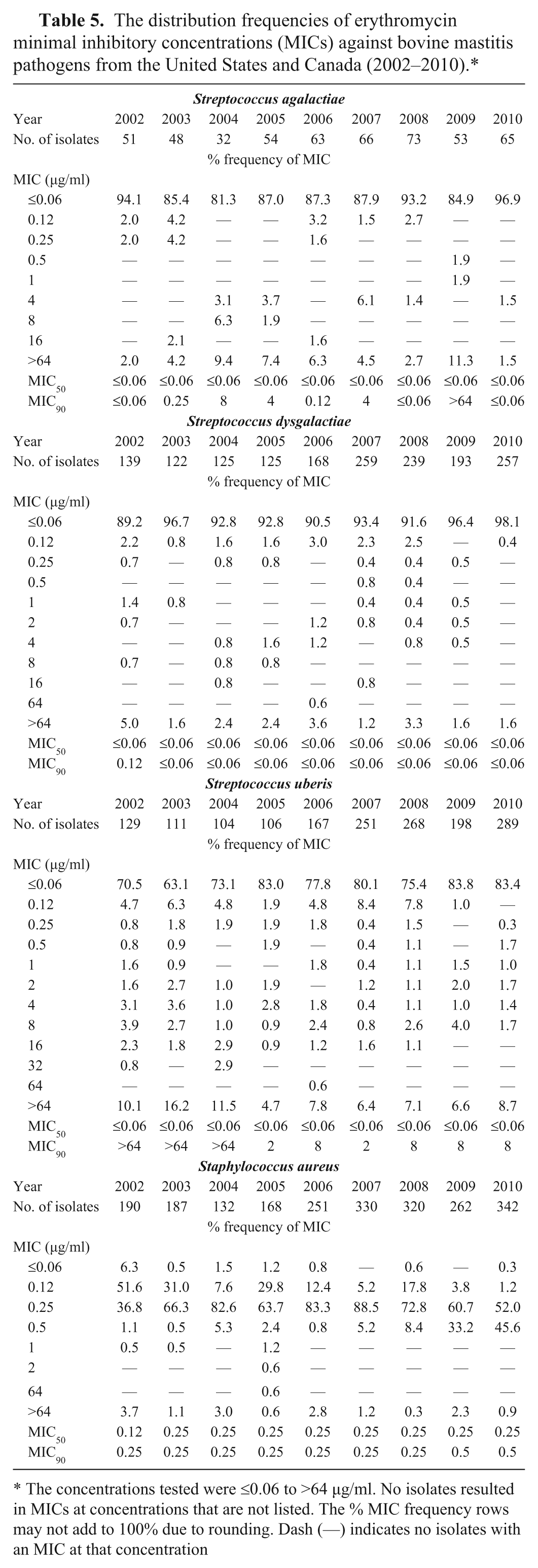
The concentrations tested were ≤0.06 to >64 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration
The distribution frequencies of oxacillin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
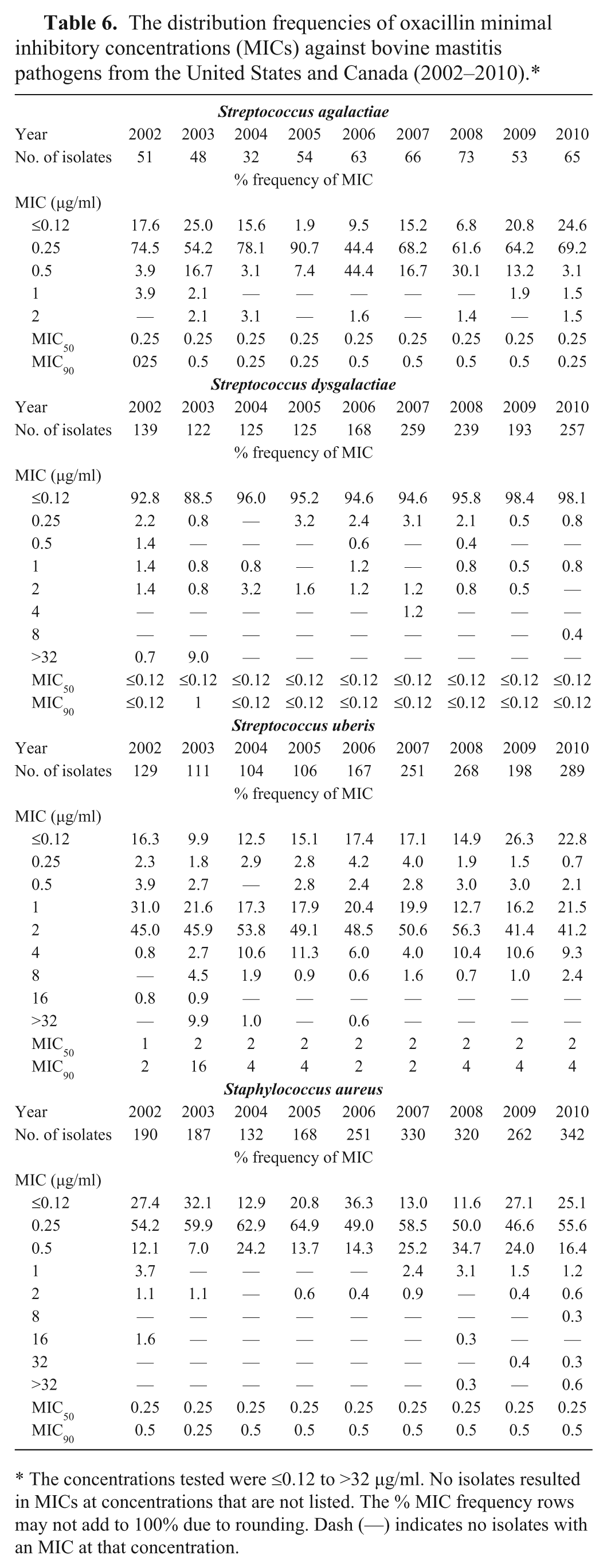
The concentrations tested were ≤0.12 to >32 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration.
The distribution frequencies of penicillin–novobiocin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
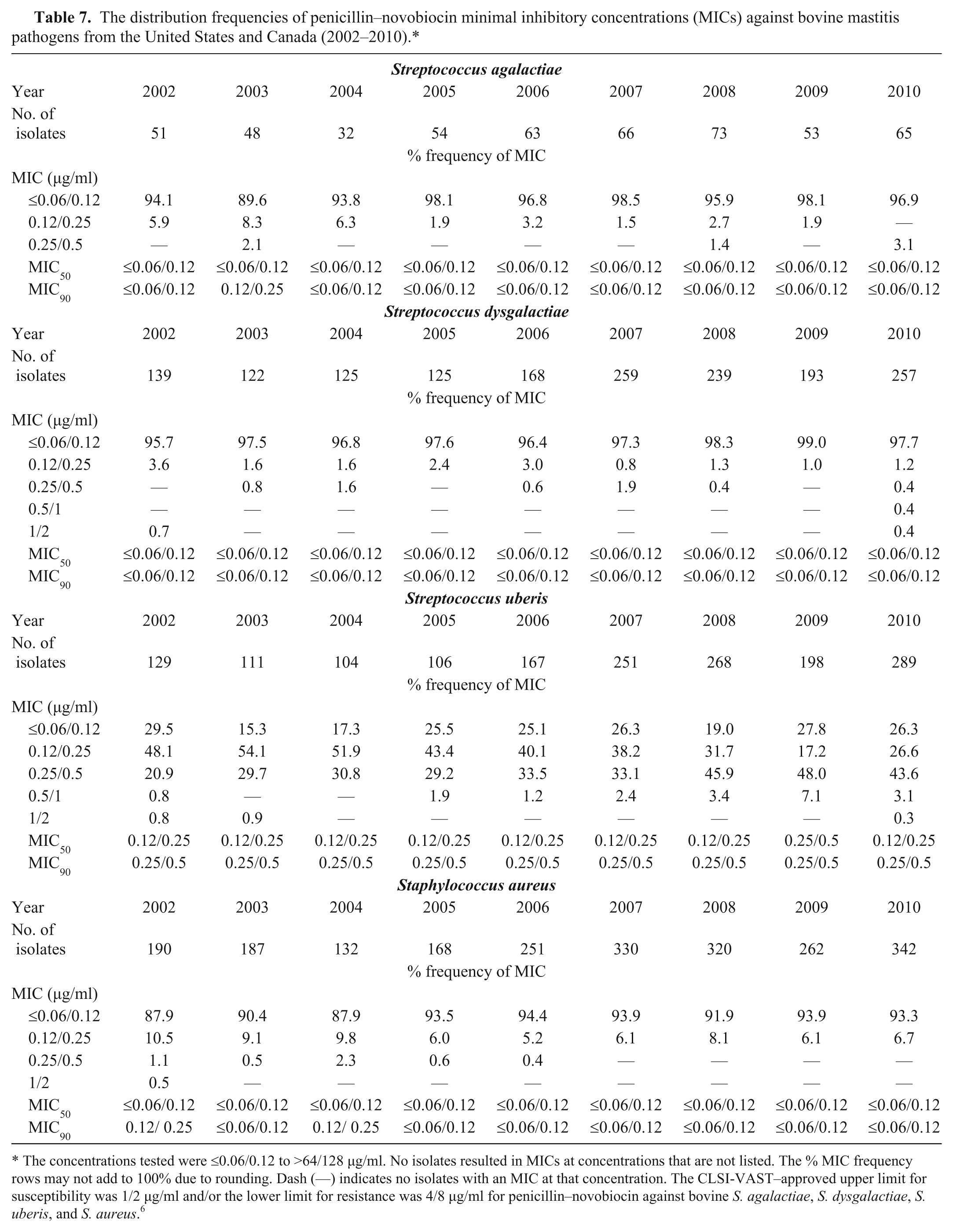
The concentrations tested were ≤0.06/0.12 to >64/128 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration. The CLSI-VAST–approved upper limit for susceptibility was 1/2 µg/ml and/or the lower limit for resistance was 4/8 µg/ml for penicillin–novobiocin against bovine
The distribution frequencies of pirlimycin minimal inhibitory concentrations (MICs) against bovine mastitis pathogens from the United States and Canada (2002–2010).*
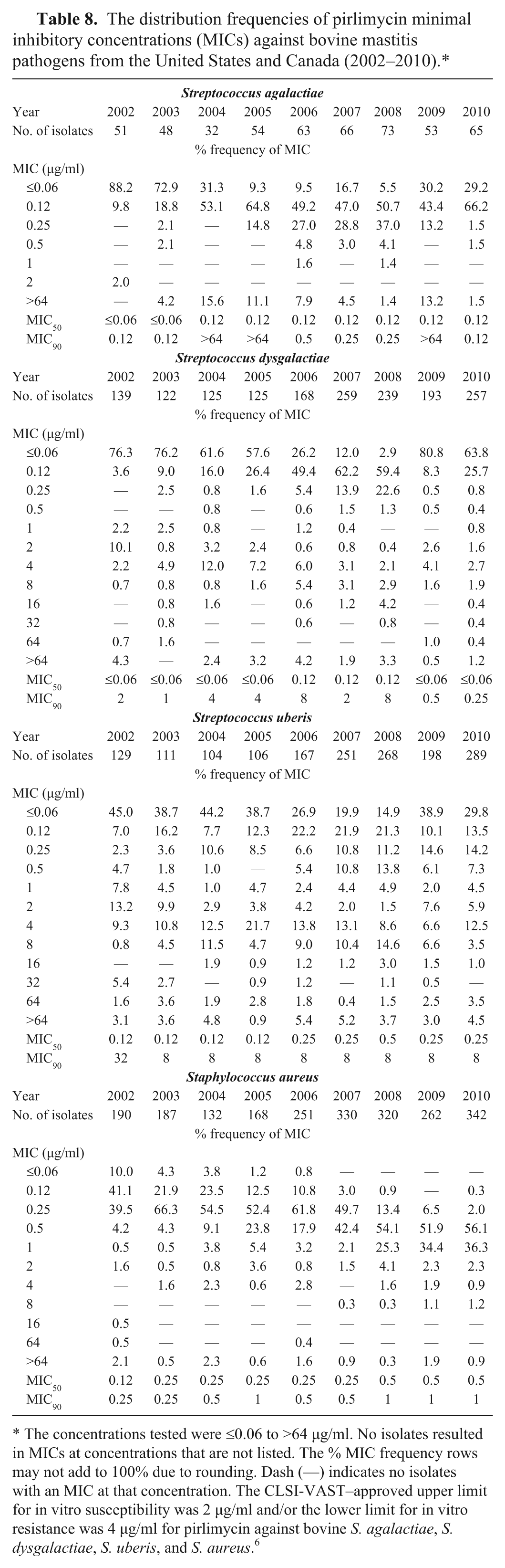
The concentrations tested were ≤0.06 to >64 µg/ml. No isolates resulted in MICs at concentrations that are not listed. The % MIC frequency rows may not add to 100% due to rounding. Dash (—) indicates no isolates with an MIC at that concentration. The CLSI-VAST–approved upper limit for in vitro susceptibility was 2 µg/ml and/or the lower limit for in vitro resistance was 4 µg/ml for pirlimycin against bovine
Ampicillin
The ampicillin MICs against the staphylococci and streptococci isolates were low, and MIC50 and MIC90 values for these pathogens were similar, within a single doubling dilution, over the 9 years of the survey (Table 2). The MIC values of ampicillin against
Ceftiofur
Streptococci and
Cephalothin
There were no apparent differences, year to year, in the cephalothin MIC50 and MIC90 values greater than a single doubling dilution among the 3 streptococcal species and
Erythromycin
Table 5 provides data on the frequency distribution of erythromycin MICs and the number of isolates tested each year from 2002 to 2010. While there was variability in the distribution of MICs, overall, the data did not provide evidence that there was any obvious increase or decrease in erythromycin susceptibility among the bovine streptococci over the 9 years of the survey. There was no change beyond a single doubling dilution in the erythromycin MIC50 or MIC90 values for
Oxacillin
Over the 9 years, oxacillin MIC50 and MIC90 values against the streptococci remained unchanged, except in 2003 when increases in the MIC90 of
Penicillin–novobiocin
All of the
Pirlimycin
As with erythromycin, there was variability in the distribution of pirlimycin MICs among the streptococci (Table 8). There appeared to be no clear temporal trend in the proportion of streptococcal isolates that were susceptible (MIC ≤2 µg/ml) to pirlimycin. More than 90% of
Discussion
There is a need for surveillance information about temporal changes in susceptibility of bovine pathogens to those antimicrobial drugs that are commonly used to treat infections, including mastitis.2,3 Reviews of surveillance studies that looked at antimicrobial susceptibility among mastitis pathogens in the United States
2
and elsewhere
15
noted that these surveys were not recent and that more current data are needed. These reviews2,15 also stated that a longer time frame to study changes in susceptibility is desirable, as are data from a wider geographical area. There are few such programs that monitor susceptibility over a whole country and for multiple, consecutive years.
3
Germany has initiated a surveillance program in which bovine mastitis pathogens (including
Overall, there did not appear to be any substantial changes in the distribution of MICs to the antimicrobial drugs tested in this 9-year study of mastitis pathogens. Previously reported high levels of in vitro susceptibility of mastitis pathogens to penicillins8,9,14,16 appear, from the data in the current survey, to have been sustained over time. While
There has been considerable discussion as to the most useful presentation of data obtained from susceptibility programs.7,20 In vitro data are often summarized using such values as MIC50, MIC90, median, mode, or range of MIC values. However, these statistics may not reflect the sometimes subtle shifts in MICs over time that may provide a more sensitive indicator of changes in susceptibility. Other summary values for describing MIC susceptibility data utilize clinical breakpoints or epidemiological cutoff values, but specific veterinary interpretive criteria have not been established for all drugs approved for use in treatment of mastitis. While the CLSI Subcommittee on Veterinary Antimicrobial Susceptibility Testing (CLSI-VAST) has approved veterinary-specific interpretive criteria for ceftiofur, penicillin–novobiocin, and pirlimycin against the Gram-positive mastitis pathogens tested in the surveillance program reported herein, only ceftiofur has interpretive criteria against
An additional challenge is the potential for interpretive criteria, be they clinical breakpoints or epidemiologic cutoff values, to be changed by standards committees, such as CLSI, and the resulting complication of interpreting historical data in light of these revised criteria. Such difficulties can be minimized by publication of the full MIC distribution for each specific “bug-drug” combination. 20 While such publication does result in large data tables, this is the approach that has been taken in the current article as it enables a number of summary values to be examined. It is also the approach that has been taken with a number of national surveillance programs that examine the antimicrobial susceptibility of zoonotic and commensal enteric bacteria, such as in Canada (CIPARS: 2010, Final report 2007. Available at: http://www.phac-aspc.gc.ca/cipars-picra/pdf/cipars-picra-2007-eng.pdf; CIPARS: 2011, Final report 2008. Available at: http://www.phac-aspc.gc.ca/cipars-picra/2008/pdf/cipars-picra-2008-eng.pdf), the United States (NARMS: 2010, 2007 executive report. Available at: http://www.fda.gov/AnimalVeterinary/SafetyHealth/AntimicrobialResistance/NationalAntimicrobialResistanceMonitoringSystem/ucm209340.htm), and Denmark (DANMAP: 2011, 2010–Use of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from food animals, foods and humans in Denmark. Available at: http://www.danmap.org/Downloads/~/media/Projektsites/Danmap/DANMAPreports/Danmap_2010.ashx).
Antimicrobial resistance surveillance programs are subject to a number of limitations, including sampling bias.3,10,18 In a study of published antimicrobial resistance surveillance studies, 17 the authors concluded that sampling bias and the failure to address multiple occurrences are common, and that case definition and laboratory practices and procedures may also influence the validity of the results. It is important that the isolates tested are representative of the population of interest. For the current mastitis surveillance program, isolates were collected from across the United States and Canada, especially from those states and provinces that contain large proportions of the national dairy herd populations (Table 1). The U.S. veterinary diagnostic mastitis laboratories that submitted isolates for this surveillance program are located in states in which more than 90% of the U.S. dairy cattle are found. The Canadian isolates came from laboratories in provinces with a substantial number of dairy operations. However, many, possibly most, bovine mastitis pathogens are not isolated and cultured, raising the possibility that the isolates from these state and provincial laboratories may not represent the whole population of pathogens. In addition, information was not available on any recent antimicrobial drug therapy that the animals may have received and so the isolates may have included bacteria from animals that had not been treated or had been treated and failed treatment.
In conclusion, the present report provides an extensive survey, in terms of geography and time, of the susceptibilities of North American bovine mastitis pathogens to beta-lactam, macrolide, and lincosamide drugs. Monitoring in vitro susceptibilities of clinical isolates can provide an early warning for the emergence of resistance. 21 The data indicate that there were no substantial numerical changes in the in vitro susceptibility of bovine mastitis pathogens from dairy cattle in Canada and the United States to these drugs during the years 2002–2010. The data support the conclusion of previous reports 15 that scientific evidence derived from local and regional studies in the United States and elsewhere does not support widespread, emerging resistance to common antimicrobial drugs among mastitis pathogens, even to drugs that have been used in dairies for many years.
Footnotes
Acknowledgements
The authors would like to thank the following veterinary diagnostic laboratories for their generous assistance by providing the bacterial isolates for this study: Arizona Dairy Herd Information Association, Cornell University (Quality Milk Production Services), High Plains Veterinary Outlet, Indiana Animal Diagnostic Laboratory (Purdue University), Iowa State University, Louisiana State University (Hill Farm Research Station), Michigan State University, Ohio Department of Agriculture, Pennsylvania State University, Texas A&M (College Station), University of Tennessee, Udder Health Systems Inc., University of California, Davis (VMTRC-Tulare), University of Guelph, University of Illinois, University of Minnesota (Laboratory for Udder Health), University of Montréal, Prairie Diagnostic Services Inc., University of Wisconsin (Madison), University of Wisconsin (Barron), Vermont Agency of Agriculture, Food & Markets Laboratory, and Washington State University. The authors would also like to thank Microbial Research Inc., as well as colleagues who provided thoughtful comments and review of this article.
a.
API Microbial Identification kits, bioMérieux Inc., Durham, NC.
b.
Biolog Microbial Identification System, Hayward, CA.
c.
Sensititre Division, Trek Diagnostic Systems Inc., Thermo Scientific, Cleveland, OH.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Cynthia J. Lindeman, Lacie Johansen, and Lisa M. Mullins are employed by Zoetis Inc. (formerly a business unit of Pfizer Inc.). Ellen Portis is a former employee of Zoetis Inc. Gillian A. Stoltman (Waterwood Consulting LLC), a former employee of the animal health business unit of Pfizer Inc., is currently providing consultation services for Zoetis Inc.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and or publications of this article: This study was funded by the animal health business of Pfizer Inc. (currently operating as Zoetis Inc.).
