Abstract
Mycobacterium avium subsp. hominissuis (Mah) infection was diagnosed in 5 captive bongo antelopes (Tragelaphus eurycerus) originating from a collection in a zoological garden. The animals suffered from emaciation. Postmortem examination revealed nodular lesions in the lungs of all 5 examined animals. Acid-fast bacilli were observed in the lungs of 4 animals. Culture and polymerase chain reaction identification based on IS901 negativity and IS1245 positivity confirmed Mah infection in the lungs of all 5 antelopes. In 3 animals, Mah was also isolated from other organs (liver, spleen, and kidney). Molecular analysis of these isolates using IS1245 restriction fragment length polymorphism and/or mycobacterial interspersed repetitive units–variable number tandem repeat revealed that the studied antelopes were infected by 1 identical genotype. Furthermore, in 2 antelopes, other genotypes were also detected. This shows the possibility of either genetic modifications occurring during infection or polyclonal infection. Culture examination of environmental samples from the enclosures holding the bongos revealed Mah in mulch bark, peat, and soil. Genotyping of these environmental isolates determined several genotypes with 1 dominant genotype that was identical to the dominant genotype detected in antelopes.
Mycobacterial infections with the development of tuberculous lesions in antelopes are most often associated with the Mycobacterium tuberculosis complex. 7 Therefore, the presence of tuberculous lesions in animals necessitates determination of the specific causal agent to assess potential epizootiological risks for other animals and human beings and to take appropriate preventive measures.
Mycobacterium avium subsp. hominissuis (Mah), a member of the Mycobacterium avium complex, is considered a potentially pathogenic mycobacterium that is able to cause infection in both animals and human beings.1,3,9,10 Infected animals usually display nonspecific symptoms (e.g., weight loss, apathy, or diarrhea). In some animals, infection might occur without any clinical signs. The most commonly affected tissues are the mesenteric lymph nodes.1,9,10 In the advanced stage of infection, systematic dissemination of mycobacteria is usually observed.1,9 The most common sources of Mah are various components of the environment, particularly water and soil (peat).4,6,8 The aim of the current study was to describe the mycobacterial infection of bongo antelopes (Tragelaphus eurycerus) originating from a zoological garden and to determine the possible source of the infection and threat to other animals.
Weight loss, cachexia, and diarrhea were observed in a group of bongo antelopes in a zoological garden over a 6-year period. The present report concerns 5 of these antelopes aged 8, 9, 16, 18, and 20 years, respectively. The antelopes were housed in a group of approximately 11 in an enclosure (enclosure 1) situated next to an enclosure of okapis (Okapia johnstoni; enclosure 2). Close to these enclosures, there was also another enclosure (enclosure 3) that was used for temporary housing of either antelopes or okapis. Two bongos died and a further 3 were euthanized because of their poor physical condition and old age (Table 1). Nodular lesions with necrotic centers and calcification were seen in the lungs of all 5 bongos during necropsy. Enlarged mesenteric lymph nodes and thyroid glands were noted in 2 animals.
Detection and characterization of Mycobacterium avium subsp. hominissuis in bongo antelopes (Tragelaphus eurycerus) from a zoological garden.*
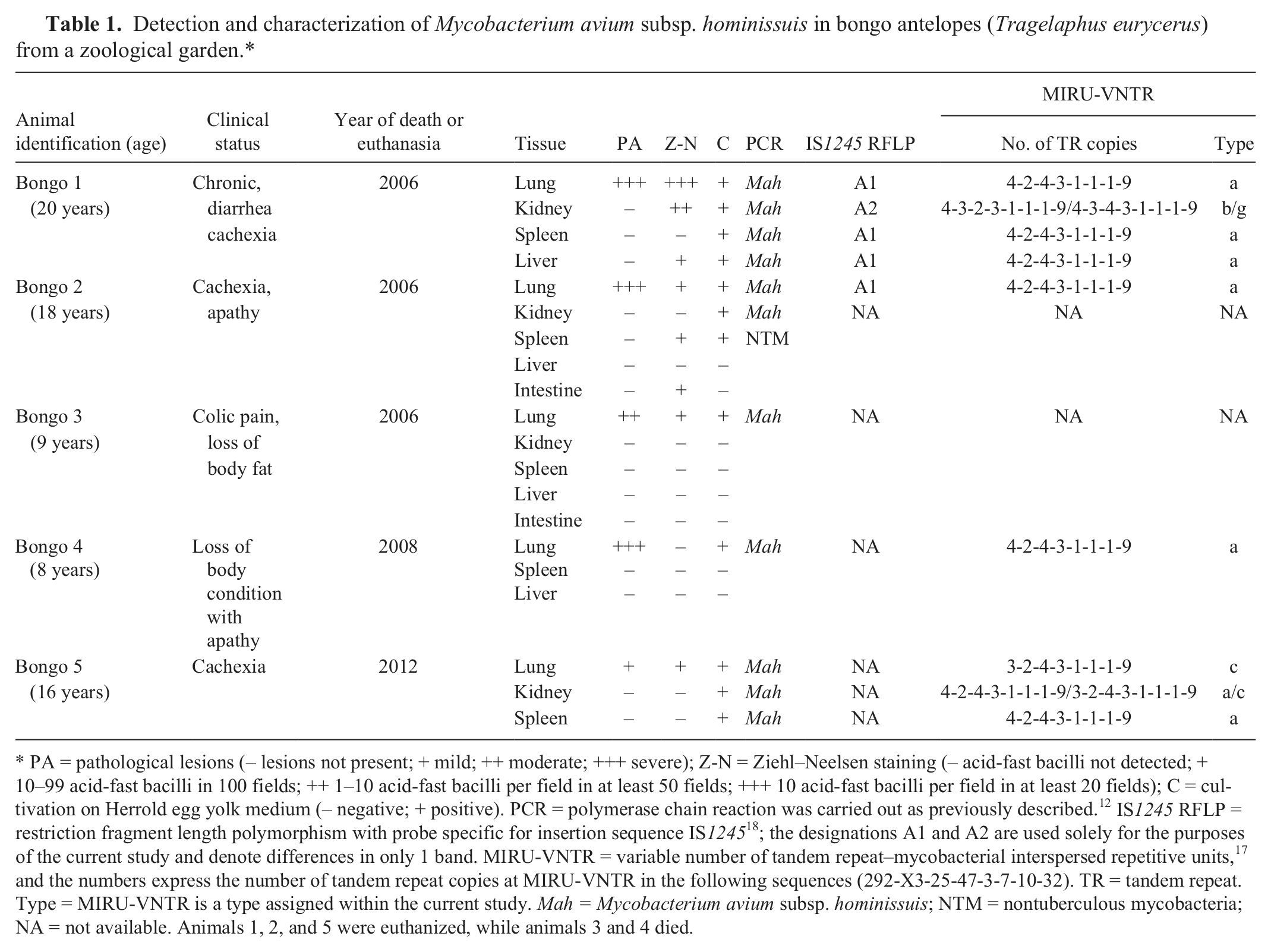
PA = pathological lesions (– lesions not present; + mild; ++ moderate; +++ severe); Z-N = Ziehl–Neelsen staining (– acid-fast bacilli not detected; + 10–99 acid-fast bacilli in 100 fields; ++ 1–10 acid-fast bacilli per field in at least 50 fields; +++ 10 acid-fast bacilli per field in at least 20 fields); C = cultivation on Herrold egg yolk medium (– negative; + positive). PCR = polymerase chain reaction was carried out as previously described. 12 IS1245 RFLP = restriction fragment length polymorphism with probe specific for insertion sequence IS1245 18 ; the designations A1 and A2 are used solely for the purposes of the current study and denote differences in only 1 band. MIRU-VNTR = variable number of tandem repeat–mycobacterial interspersed repetitive units, 17 and the numbers express the number of tandem repeat copies at MIRU-VNTR in the following sequences (292-X3-25-47-3-7-10-32). TR = tandem repeat. Type = MIRU-VNTR is a type assigned within the current study. Mah = Mycobacterium avium subsp. hominissuis; NTM = nontuberculous mycobacteria; NA = not available. Animals 1, 2, and 5 were euthanized, while animals 3 and 4 died.
To determine the etiological agent, samples of lung, spleen, liver, kidney, and intestine were collected from the 5 bongo antelopes for Ziehl–Neelsen staining and cultivation on Herrold egg yolk medium. Ziehl–Neelsen staining demonstrated the presence of acid-fast bacilli (AFB) in the lungs of 4 out of the 5 bongos examined. Furthermore, AFB were observed in the kidney, liver, spleen, and intestine in 2 bongos (Table 1). Culture on Herrold egg yolk medium 16 and subsequent polymerase chain reaction (PCR) identification 12 confirmed the presence of Mah (IS901–, IS1245+) in the lungs of all 5 animals. Mycobacterium avium subsp. hominissuis (IS901–, IS1245+) was also isolated from other parenchymatous organs (kidney, spleen, and liver) in 3 cases. One isolate from the spleen was determined to be a Mycobacterium sp. (Table 1).
To assess the source of infection and possible risks to other antelopes and other animals from the nearby enclosures (especially okapis), a total of 117 environmental samples (soil, peat, water, bark mulch, feed, straw with urine, dust, and house flies [Musca domestica]) and 22 fecal samples from bongos and okapis were examined using culture 15 and subsequent PCR identification of isolates. 12 Mycobacterium avium subsp. hominissuis was isolated from 15 out of 139 examined environmental samples (soil and peat, bark mulch, straw with urine, from 1 Musca domestica) including 2 fecal samples from okapis (Table 2).
Culture examination of the environment and stored bark mulch and genotyping of Mycobacterium avium subsp. hominissuis isolates from a zoological garden.*
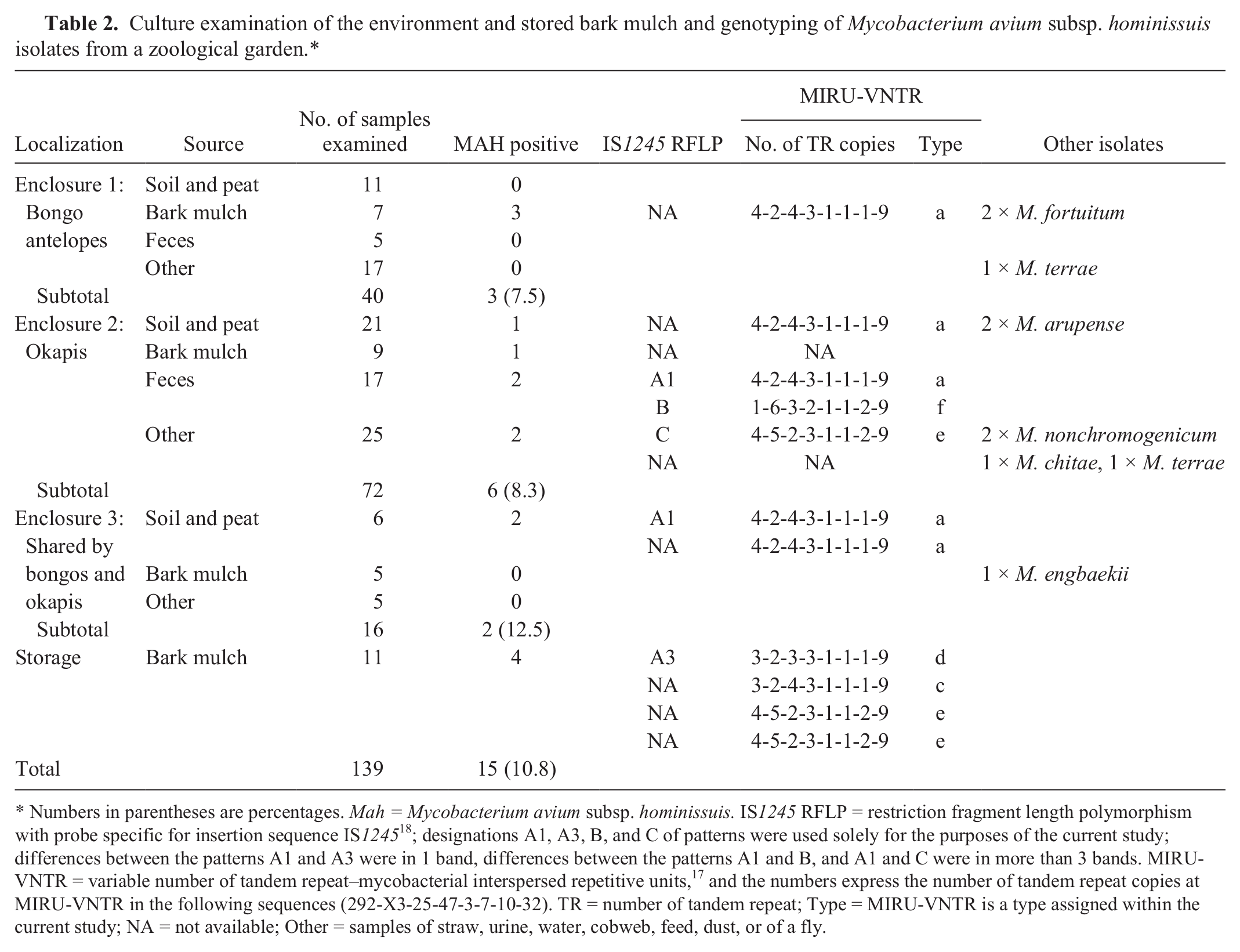
Numbers in parentheses are percentages. Mah = Mycobacterium avium subsp. hominissuis. IS1245 RFLP = restriction fragment length polymorphism with probe specific for insertion sequence IS1245 18 ; designations A1, A3, B, and C of patterns were used solely for the purposes of the current study; differences between the patterns A1 and A3 were in 1 band, differences between the patterns A1 and B, and A1 and C were in more than 3 bands. MIRU-VNTR = variable number of tandem repeat–mycobacterial interspersed repetitive units, 17 and the numbers express the number of tandem repeat copies at MIRU-VNTR in the following sequences (292-X3-25-47-3-7-10-32). TR = number of tandem repeat; Type = MIRU-VNTR is a type assigned within the current study; NA = not available; Other = samples of straw, urine, water, cobweb, feed, dust, or of a fly.
Mycobacterial isolates from environmental samples were further identified by sequencing of the 16S ribosomal RNA 5 or the hsp65 fragment. 11 Two isolates of Mycobacterium fortuitum and 1 isolate of Mycobacterium engbaekii were obtained from the bark mulch. The soil samples contained Mycobacterium arupense, while Mycobacterium nonchromogenicum was detected in cobweb and dust. Mycobacterium terrae and Mycobacterium chitae were isolated from straw with urine (Table 2).
Some Mah isolates from the bongos, okapi feces, and the environment were further genotyped using IS1245 restriction fragment length polymorphism (RFLP) and/or mycobacterial interspersed repetitive units–variable number of tandem repeat (MIRU-VNTR) typing as previously described.17,18 Both methods revealed different genotypes with 1 predominating genotype. The most common genotype was genotype A1, as assessed by IS1245 RFLP, and genotype a, as assessed by MIRU-VNTR analysis. As expected, all isolates with IS1245 RFLP pattern A1 produced an identical MIRU-VNTR pattern (pattern a). In contrast, isolates with IS1245 RFLP genotypes A2 and A3, differing by 1 fragment from the predominant profile A1, showed different MIRU-VNTR genotypes.
Although animals are exposed to Mah frequently, 4 clinical signs or pathological lesions are rarely observed.1,9,13 In the current study, Mah infection with associated pathological lesions in the lungs of 5 bongos housed in a zoological garden is reported. Before death or euthanasia, the antelopes were in poor body condition with a history of weight loss. This manifestation of Mah infection has been previously described in other animals.1,9 Both examined okapis sharing a temporary enclosure with the infected antelopes were without any clinical signs of infection, and Mah was only isolated from 2 fecal samples out of 17 collected from the okapis. Therefore, passive transport of Mah through these animals or asymptomatic infection is highly likely. The reason why the Mah infection developed only in the bongos has not been determined. An underlying disease may have played a role in the development of the Mah infection. 4 Also, the possibility that bongo and okapi antelopes have different genetic susceptibilities to Mah infection should be considered. 14
Molecular analysis of the Mah isolates revealed that all 4 antelopes were infected by 1 identical genotype (antelope 3 was not genotyped due to contamination of the culture). Furthermore, in 2 antelopes, other genotypes were detected in their organs (Table 1), suggesting that polyclonal infection or genotype reorganization took place during infection. Minor (1 or 2 bands) RFLP variations have been reported in M. avium isolates subjected to serial subcultures over 30 months. 2 Although the MIRU-VNTR results in the current study support the existence of different genotypes, the possibility exists that the minor RFLP variations between A1, A2, and A3 represent a genetic drift similar to that observed in vitro as noted above. 2
The most common route of Mah transmission to animals or human beings is by ingestion or inhalation of Mah directly from the environment that serves as a natural source of Mah infection.4,6,10 In the current study, Mah was isolated from 15 out of 139 examined environmental samples examined including 2 fecal samples from okapis (Table 2). Genotyping of these environmental isolates revealed several genotypes, including 1 predominant genotype that was identical to the major genotype detected in antelopes. In the past, peat was suggested to be a source of infection for pigs. 10 Both bark mulch and peat are materials with low pH (5.0–3.5), which suppresses the growth of the majority of bacteria but represents ideal conditions for mycobacteria.4,8 Although the possibility of the bark mulch being the source of infection has been considered, there is no convincing evidence to support such a causal relationship. The MIRU-VNTR analysis of Mah isolates found in storage bark mulch revealed genotypes that differed in the number of tandem repeats in 1 or more loci from the predominant genotype detected in the bongo antelopes and environment. In the IS1245 RFLP analysis, storage bark mulch isolates showed the A3 pattern that differed by 1 band from the predominant pattern A1 observed in isolates derived from the bongo antelopes and the environment.
A source of Mah may also be infected animals that shed the infectious agent through their feces into the environment. 9 In the current study, Mah was not isolated from any fecal samples from bongo antelopes, and the possibility of a fecal–oral route of infection in these animals seems unlikely. Although the negative results of fecal cultivation may be due to the low numbers of Mah in the feces of the bongo antelopes or intermittent Mah shedding in feces, 15 the isolation of Mah from the feces of the okapi antelopes demonstrates the robustness of the culture method used in the present study.
The spread of mycobacterial infection via the respiratory route is also possible. Although histology was not part of the current study, caseous pulmonary lesions and consistent isolation of Mah from the lungs of 5 bongo antelopes is consistent with the respiratory tract being the primary site of infection subsequent to inhalation of Mah.
In conclusion, infection of bongo antelopes with Mah is reported. A dominant genotype was associated with this infection and that genotype was also dominant in environmental samples associated with the antelopes.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Grant No. MZE0002716202 of the Ministry of Agriculture of the Czech Republic and the Ministry of Education, Youth and Sports of the Czech Republic (Grant “AdmireVet” no. CZ 1.05/2.1.00/01.0006-ED0006/01/01).
