Abstract
Traditionally, the bacteriological examination of mastitis milk samples is performed by culture followed by biochemical tests on the cultured bacteria to allow identification of the causative pathogen. Depending on the species involved, this classic identification is time-consuming compared to other techniques such as fluorescent in situ hybridization (FISH), a culture-independent method that utilizes oligonucleotides (labeled with a fluorophore) that are specific to a string of target DNA/RNA. In the current study, the applicability of FISH was evaluated for the detection of mastitis pathogens directly in milk samples. To remove interfering lipids and proteins from mastitis milk samples prior to FISH, a previously published enzymatic treatment with savinase was evaluated. FISH was performed using oligonucleotides specific for Staphylococcus aureus, Streptococcus agalactiae, Streptococcus uberis, Enterococcus faecalis, Enterococcus faecium, Escherichia coli, and Trueperella (Arcanobacterium) pyogenes. The enzymatic pretreatment and the sensitivity of FISH were evaluated using spiked whole milk samples and mastitis milk samples with bacterial loads of less than 103 up to 108 colony-forming units (CFU)/ml. Bacteria were reliably detected in milk samples with bacterial numbers of 106 CFU/ml or higher. However, bacteria present in numbers below 106 CFU/ml were not detectable in all cases. The ability of FISH to identify mastitis-causing pathogens directly in milk samples, and therefore earlier than classical culture methods, can supplement the classic diagnostic procedures for mastitis milk samples.
Introduction
Mastitis is one of the most important diseases in dairy cattle. Economic losses are estimated up to €200 (US$260) per affected cow and year. 12 Various species of streptococci, followed by coagulase-negative staphylococci, coliforms as well as Staphylococcus aureus are reported as predominant infectious agents in clinical cases of mastitis. 27 Rapid and reliable identification of the causative agent(s) can support effective management of the disease. 19
Generally, mastitis milk samples are examined with classical bacteriological culture-based methods, followed by biochemical tests for species identification. Depending on the species involved, identification requires 24–96 hr. 26 Less time-consuming variations on classic cultural methods have been developed to provide timely information. 17 Various culture-independent methods have been published for the rapid detection of mastitis pathogens in milk samples, as well as various polymerase chain reaction (PCR) approaches including real-time PCR methods.6,15,16
A culture-independent molecular method allowing the direct detection of pathogens in clinical samples is fluorescent in situ hybridization (FISH).24,25,29 Fluorescence-labeled oligonucleotide probes bind specific regions of the target RNA/DNA. Furthermore, several probes labeled with different fluorophores can be combined into a single test and therefore different species can be detected simultaneously. 20 Because of the high specificity, this method has been used for the identification of pathogens from culture and directly from various clinical samples4,11,13 and for the characterization of bacterial communities of animals.18,22 The applicability of FISH for the detection of bacteria such as pseudomonads in milk samples has been demonstrated using peptic nucleic acid probes in combination with flow cytometry.9,10 The microscopic examination of artificially spiked milk samples using FISH has been reported for Listeria and Salmonella. 21
The aim of the current study was to develop and evaluate a FISH protocol for the direct detection of pathogens in mastitis milk samples. FISH was performed using a set of oligonucleotide probes covering relevant mastitis pathogens. For most mastitis pathogens, useful probes have already been available for some years.3,14,24,28 Two published probes specific for the mastitis pathogens Streptococcus uberis and Trueperella (Arcanobacterium) pyogenes 30 were included. Additionally, an enzymatic pretreatment for mastitis milk samples 10 was adapted and evaluated in order to optimize the FISH procedure for the detection of pathogens in milk samples.
Materials and methods
Milk samples
Quarter milk samples used for the current study were obtained from routine diagnostic laboratories.a,b Samples were collected from dairy cows of herds located in the area surrounding Munich, Germany. Samples were randomly chosen and included mainly samples from animals with clinical mastitis. In some cases, subclinical mastitis was reported or no data were available concerning the character of mastitis. Ultra-high temperature (UHT)-treated milk (3.5% fat) purchased from a local grocery store was used for artificial bacterial spiking to evaluate the enzyme pretreatment of milk samples and to establish the detection limit for FISH.
Oligonucleotide probes
Previously published probes (and competitors) were used for the present study (Table 1). Four family- or genus-specific probes (Staphy, Strept, ENC 221, and EBAC 1790) as well as 7 species-specific probes (Sau, Straga, SUB 196, ENF 191, ENU 140, APYO 183, and ESC 468) were included in the study. Probe EUB 338, specific for the majority of eubacteria, 5 was used as a positive control for the FISH procedure and for the determination of the detection limit. Probes were synthesized and fluorescently labeled with 6-carboxyfluorescein (6-FAM) or cyanine 3 (Cy3).c,d Concentrations of stock solution were 50 ng/µl for probes labeled with 6-FAM and 30 ng/µl for probes labeled with Cy3. Synthesized competitors (unlabeled) were used in combination with their corresponding probe in order to avoid unspecific binding (Table 1).
Sequences of oligonucleotide probes used in the current study.
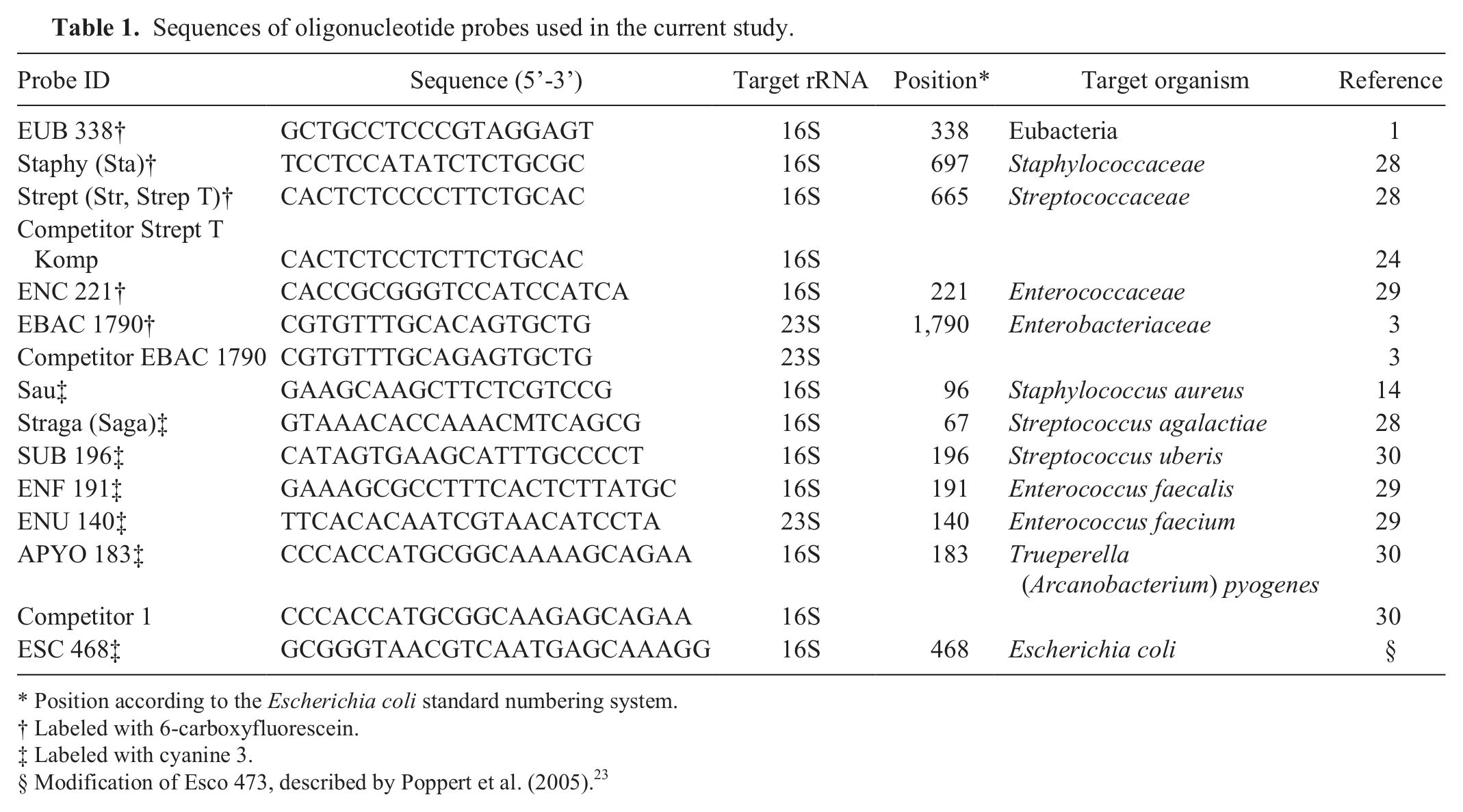
Position according to the Escherichia coli standard numbering system.
Labeled with 6-carboxyfluorescein.
Labeled with cyanine 3.
Modification of Esco 473, described by Poppert et al. (2005). 23
Conventional examination of milk samples
To compare results of FISH with usually reported semiquantitative cultural results, mastitis milk samples (10 µl) were streaked onto agar plates (blood agar, Gassner agar, e CNA [colistin and naladixic acid] agar with 5% sheep blood f ), and were diluted with 3 further dilution streaks. Bacterial growth was documented after 24 hr and after 48 hr of incubation. The quantity of colonies grown was determined semiquantitatively and expressed in 5 categories: ≤10 colony-forming units (CFU) as (+); colonies only in the first dilution streak as +; colonies in the second dilution streak as ++; colonies in the third dilution streak as +++; and close bacterial growth in every dilution streak as ++++.
To determine the total bacterial count quantitatively, milk samples were additionally diluted in steps of 1:10, and 100 µl of each was spread on blood agar plates e (supplemented with 7% [v/v] defibrinated sheep blood). After overnight incubation, colonies on agar plates were counted, and the total bacterial count of the original sample was calculated.
Identification of bacterial species
Preliminary identification of isolates was according to hemolysis characteristics, catalase test, cytochrome-oxidase test, and Gram staining. Definitive identification to species level was performed either with a latex agglutination test g or biochemical identification systems.h,i
Procedure for savinase treatment
Milk samples were homogenized on a vortex mixer. Subsequent pretreatment with savinase was performed using a modification of a previously described method to clear milk of proteins and lipids. 10 Briefly, 5 µl of savinase (protease from Bacillus cereus, activity ≥16 U/g) j was added to 500 µl of milk. The mixture was incubated at 30°C for 30 min. After centrifugation at 9,300 × g for 10 min, the lipid layer on top was gently removed, and the aqueous phase containing the digested proteins was carefully separated with a micropipette. The pellet was resuspended in 500 µl of 0.15 M NaCl k and centrifuged at 9,300 × g for 5 min (washing). If the supernatant was not clear, the washing step was repeated. Otherwise, the clear supernatant was carefully removed, and the pellet was dissolved in 100 µl of 0.15 M NaCl. This suspension was used for further investigations.
Evaluation of the savinase pretreatment and the detection limit of FISH
The enzymatic pretreatment of milk samples and the detection limit of FISH were evaluated with spiked milk samples. Commercial UHT-milk aliquots were inoculated with the following reference strains: Escherichia coli ATCC 25922, Enterococcus faecalis ATCC 29212, or Staph. aureus ATCC 29213. A series of controls (n = 75) with bacterial concentrations between 101 and 108 CFU/ml was prepared. Gram stains were prepared according to standard procedures both before and after savinase treatment using 8 µl of milk sample applied on a glass slide. The visible number of bacteria in a microscopic field was semiquantitatively documented in 6 categories: no bacteria (–), very few (+), few (+), moderate (++), many (+++), and in masses (++++). The influence of savinase pretreatment on milk samples was evaluated by comparing the before and after Gram stains with regard to the visually detectable number of organisms. To test whether the method was applicable to field samples, a set of 37 mastitis milk samples was examined and analyzed in the same way.
In order to determine the detection limit of FISH, the 75 spiked UHT-milk samples were examined with FISH (after savinase treatment, applying the FISH protocol as described in the following) using probe EUB 338. Samples containing bacteria in concentrations from 101 to 108 CFU/ml were evaluated by analyzing 50 fields of view at a magnification of 630× searching for detectable bacteria. The number of detectable organisms was semiquantitatively determined in 6 categories, as described previously for Gram staining. Additionally, the results of the 40 field samples containing species in pure culture were evaluated with respect to the detection limit.
Procedure and protocol of FISH for milk samples
Preparation of microscopic slides for FISH
An 8-μl aliquot of the resuspended pellet was applied to each well (transparent area) of a Teflon-coated 8-well adhesive glass slide k and air-dried. Subsequently, slides were fixed in absolute methanol e for 15 min and air-dried again. The slides were either examined immediately or stored at −20°C until analysis with FISH. Frozen microscopic slides were dehydrated in 50%, 80%, and 96% (v/v) ethanol e for 3 min during each step and air-dried before use.
Lysis of Gram-positive bacteria
Prior to hybridization, the cell wall of Gram-positive bacteria was permeabilized to allow the probes to penetrate the cells. Depending on probe specificity, lysis was performed per well either with 15 µl of lysozyme e (1 mg/ml) for the detection of streptococci, enterococci, and Gram-positive rods or with lysostaphin e (2 µg/ml) for staphylococci. Incubation time was 3 min (ENC 221, ENF 191, ENU 140), 5 min (APYO 183), 10 min (Strept, Straga, SUB 196), or 15 min (Staphy, Sau) in a humid chamber at 37°C. The reaction was stopped in absolute methanol for 3 min, and slides were air-dried.
Hybridization
For hybridization, a mixture of probes labeled with fluorophores, their competitors (unlabeled), and hybridization buffer (900 mM NaCl, 20 mM Tris-HCl l [pH 8], 0.01% [w/v] sodium dodecyl sulfate [SDS] buffer k ), containing formamide e at a concentration of 30% (v/v), was used. For each well, 10 µl of hybridization buffer was mixed with 1 µl of stock solution of each probe (50 ng/µl labeled with 6-FAM, 30 ng/µl labeled with Cy3) and competitor, if needed. The probe mixture was placed onto the respective wells of the slide. Slides were incubated at 46°C for 90 min in a humid chamber containing a tissue soaked with hybridization buffer. After hybridization, the slides were rinsed gently with distilled water and were washed in preheated washing buffer, containing 5 mM EDTA k (pH 8.0), 20 mM Tris-HCl (pH 8.0), 0.01% (w/v) SDS, 102 mM NaCl, and distilled water, at 48°C for 15 min. The rinse step was repeated. Slides were rapidly air-dried and embedded with 2 drops of an anti-bleaching agent m and examined with an epifluorescence microscope n at a magnification of 630×. Bacterial numbers detected by FISH were documented semiquantitatively in 6 categories, as described previously for Gram staining.
Evaluation of FISH for the direct detection of pathogens in mastitis milk samples
Quarter milk samples (n = 55) from cows with clinical and subclinical mastitis were examined. Cultural examination and FISH were performed concurrently on the day of arrival at the laboratory. Samples were homogenized, pretreated with savinase, and Gram stained.
For each sample, a total bacterial count, the semiquantification of bacterial growth, the identification of pathogens, and FISH were performed. Species-specific probes, usually labeled with Cy3, were applied in combination with a family- or genus-specific probe, usually labeled with 6-FAM.
Finally, the results obtained with FISH were qualitatively and quantitatively compared with the results generated by culture. The 2 approaches were designated as “results identical” if 1) a consistent match in bacterial genus or species (depending on the probe specificity) was observed and 2) a maximal deviation of 2 grades in the semiquantitative determination of bacterial numbers in culture and FISH was present.
Results
Pretreatment of milk samples with savinase
Prior to a comparison of FISH with cultural methods, a protocol to eliminate inhibitory milk ingredients had to be established. Preliminary FISH experiments using untreated spiked samples were unsatisfactory because fixed samples tended to peel off from the slides and FISH showed high levels of background staining (data not shown). Therefore, a previously published protocol for the removal of lipids and proteins from milk samples using enzymatic treatment with savinase and centrifugation steps was tested. In initial experiments, this protocol improved the results with UHT-milk samples, but the results with more viscous or floccose milk from acute mastitis cases were still not satisfactory (data not shown). Thus, a homogenization step using a vortex mixer before the enzymatic treatment, extended centrifugation times, and added washing steps were introduced. This modified procedure was evaluated by Gram staining using 75 UHT-milk samples spiked with different concentrations of different bacteria (Table 2). One aliquot of each sample with and without the modified pretreatment was investigated by Gram staining.
Evaluation of pretreatment with savinase and evaluation of the detection limit of the fluorescent in situ hybridization method in milk samples using the optimized procedure.*
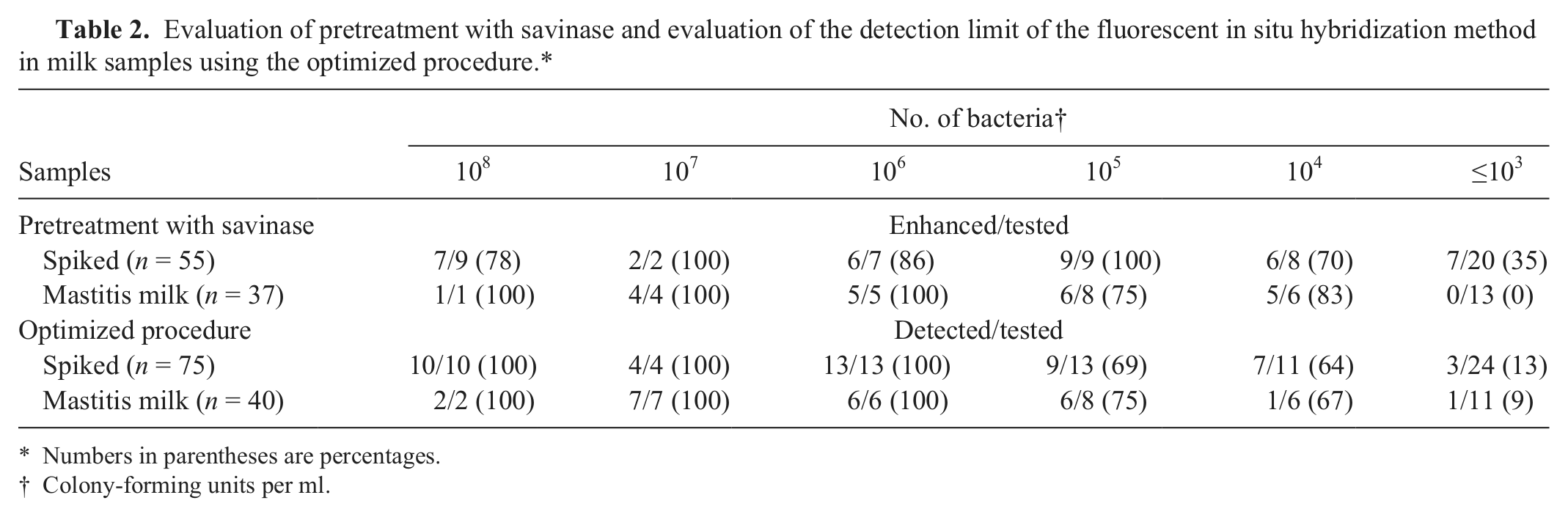
Numbers in parentheses are percentages.
Colony-forming units per ml.
In 20 of the 75 untreated spiked UHT-milk samples, the fixed material peeled off the slide, and the respective samples were excluded from evaluation. In 37 of the remaining 55 samples, the enzymatic pretreatment resulted in an increase of the amount of visible bacteria in a microscopic field of view (Table 2). Below a bacterial concentration of 103 CFU/ml, bacteria were rarely visible and an enhancement of the bacterial numbers could therefore only be documented in a few cases.
In the next step, the savinase treatment was evaluated using 37 milk samples from cows with clinical signs of mastitis (Table 2). The savinase pretreatment of the mastitis milk samples led to a significant increase in the number of bacteria that were visible in samples containing bacteria in concentrations of ≥104 CFU/ml. However, compared with spiked milk samples, no enhancement was noted in mastitis milk samples containing 103 CFU/ml or fewer. Gram stains of a milk sample with E. coli, representative for all samples, are shown in Figure 1. Bacteria are more difficult to identify without pretreatment (Fig. 1A) compared with the pretreated milk sample (Fig. 1B). Furthermore, a significant reduction of unspecific background staining was observed.
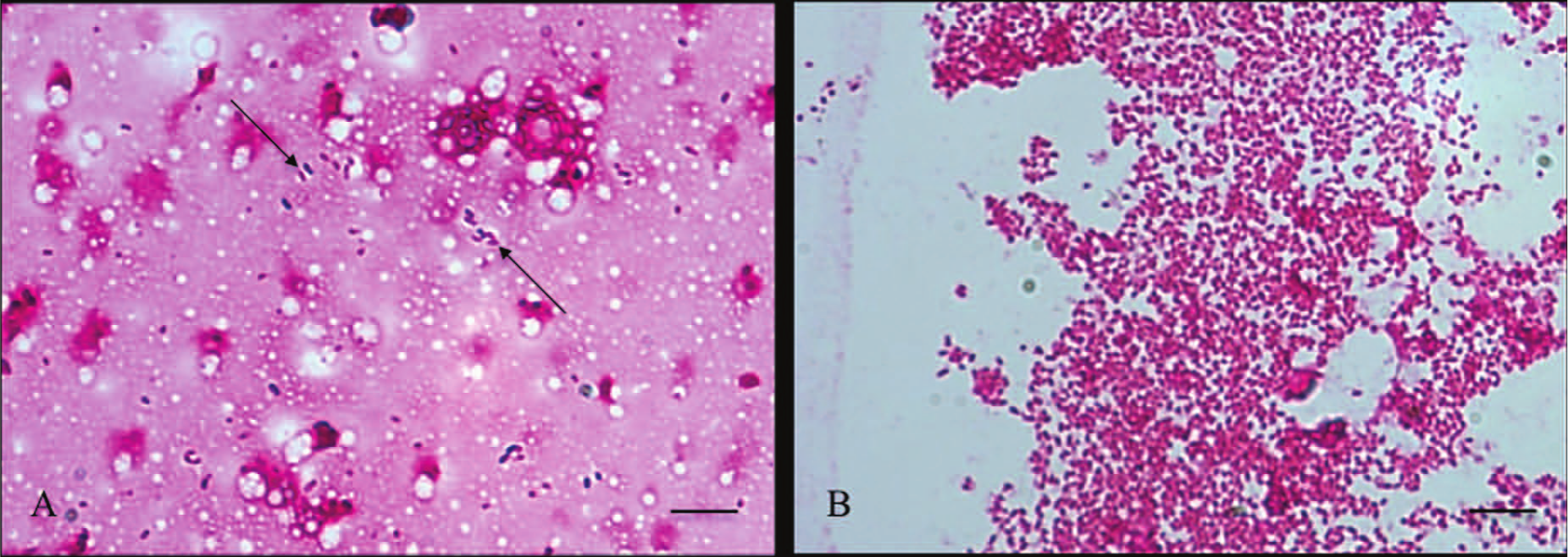
Effect of savinase pretreatment on a mastitis milk sample (containing Escherichia coli) visualized with Gram stain.
Detection limits of FISH
The 75 spiked UHT-milk samples were analyzed in order to determine the minimal number of bacteria that is required to be detected with the newly developed FISH protocol (Table 2). In samples with bacterial numbers of 106 CFU/ml or more, the detection rate of FISH was 100% (Table 2). In the range between 104 and 105 CFU/ml, successful detection was observed in 67% of the cases. However, if bacteria were present in concentrations of 103 CFU/ml or fewer, bacteria were only detected occasionally with FISH (13%).
Similar results were achieved by evaluating the detection limit in the 40 mastitis milk samples with only 1 species. Confirming the results with spiked UHT-milk samples, bacteria at a concentration of 106 CFU/ml or more were detected in all cases examined (Table 2).
Detection of bacteria in mastitis milk samples by FISH in comparison with culture
For the evaluation of the applicability of FISH in a diagnostic setting, 55 mastitis milk samples were examined by FISH on the day of arrival at the laboratory in parallel with conventional cultural examination. The samples for evaluation were divided into samples containing only 1 pathogen (hereafter referred to as pure cultures) and samples containing 2 or more bacterial species (hereafter referred to as mixed cultures). Forty of the 55 mastitis milk samples turned out to be pure cultures and 15 to be mixed cultures.
Pure cultures
The bacterial concentrations in the milk samples growing just 1 bacterial species ranged from less than 103 CFU/ml to 108 CFU/ml (Table 3). A detection rate of 100% was documented for FISH when bacterial concentrations exceeded 106 CFU/ml (Table 2). As seen in the previous experiments, the detection rate decreased in parallel with decreasing concentrations of bacteria (Table 3).
Results of fluorescent in situ hybridization and culture techniques on mastitis milk samples that contained pure cultures (n = 40).*
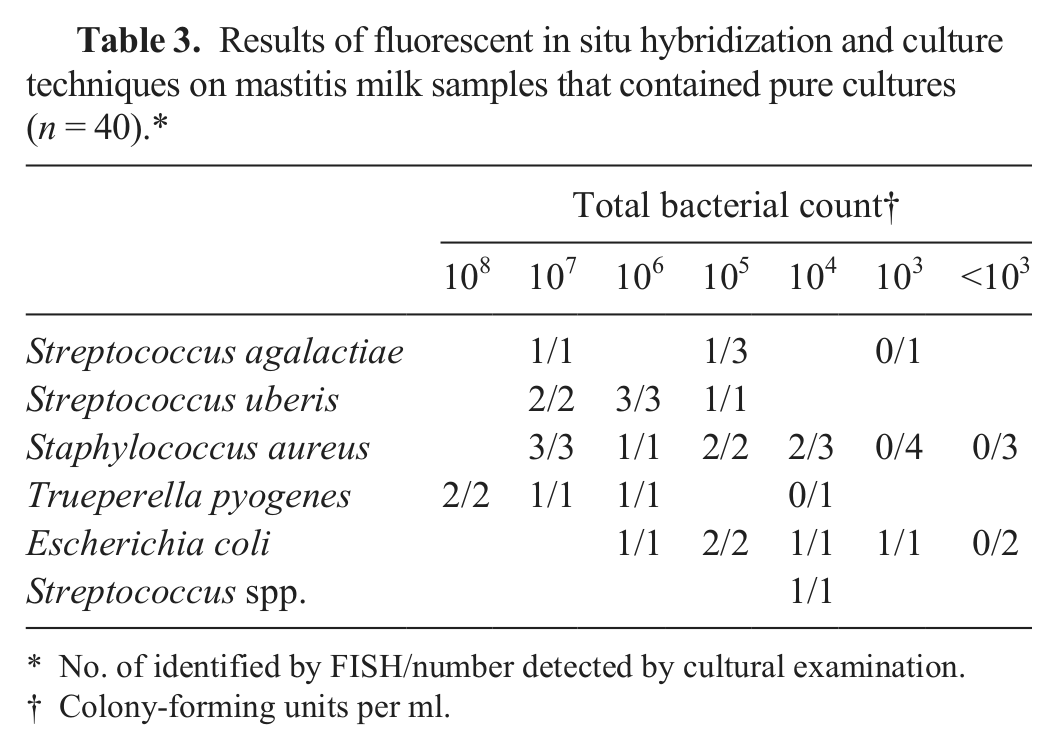
No. of identified by FISH/number detected by cultural examination.
Colony-forming units per ml.
Microorganisms detected were S. agalactiae (n = 5), S. uberis (n = 6), Staph. aureus (n = 16), T. pyogenes (n = 5), E. coli (n = 7), and Streptococcus spp. (n = 1). All bacteria that were detected by FISH were correctly identified to species level by FISH as well as by phenotypic methods. One Streptococcus species was identified only to genus level in both identification approaches.
Mixed cultures
In samples with a total bacterial count of 106 CFU/ml and greater, all FISH probes applied detected corresponding pathogens that were confirmed by cultural results (Table 4). Also, the concentration of bacteria determined with FISH was in the majority of samples tested, similar to the corresponding numbers obtained by culture. Hybridized pathogens could be identified clearly (Fig. 2).
Results of fluorescent in situ hybridization (FISH) and cultural examination of mastitis milk samples that contained mixed cultures (n = 15).
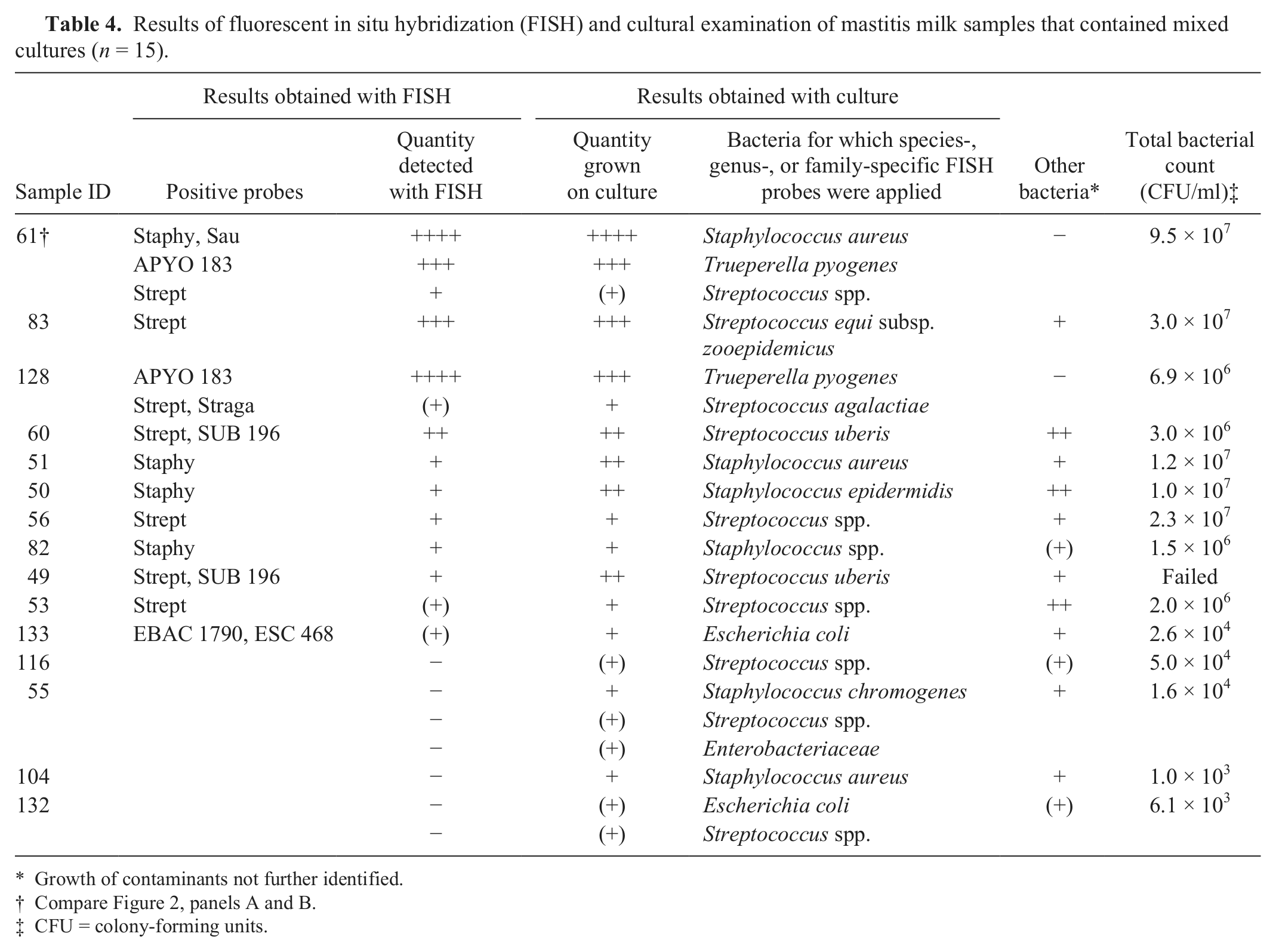
Growth of contaminants not further identified.
Compare Figure 2, panels A and B.
CFU = colony-forming units.
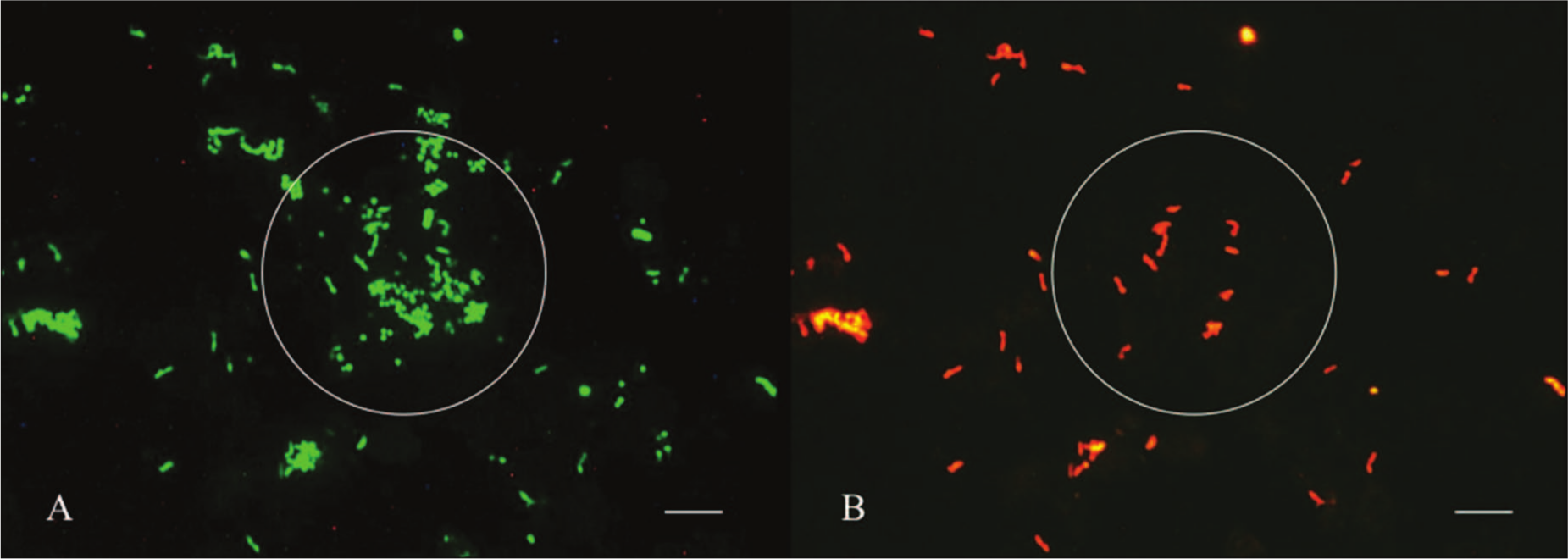
fluorescent in situ hybridization (FISH) analysis of the mastitis milk sample (ID 61, Table 4). Concurrent hybridization was performed with the Bacteria-specific probe EUB 338 (6-carboxyfluorescein [6-FAM]-labeled) and the species-specific probe APYO 183 for Trueperella pyogenes (cyanine 3 [Cy3]-labeled). Pictures of the fluorescent signal were taken from the same area.
However, detection by FISH failed in 4 samples with a total bacterial count of 103 to 104 CFU/ml. The numbers of bacteria on the agar plates confirmed the low numbers of organisms present [(+) or +]. In these samples, bacteria were consequently not detectable microscopically with the FISH method.
Discussion
The aim of the current study was to investigate whether FISH, a well-established diagnostic tool in human medicine,25,29 is suitable for the direct detection of mastitis pathogens in milk samples. For this purpose, a set of previously published probes was selected that target the most common mastitis pathogens, namely, Staph. aureus, S. agalactiae, S. uberis, Enterococcus faecalis, Ent. faecium, E. coli, and T. pyogenes. The most frequently observed pathogens in the 55 tested mastitis milk samples were Staph. aureus (n = 19) and various species of streptococci, especially S. uberis (n = 8) and S. agalactiae (n = 8). The probes specific for E. coli and T. pyogenes each detected corresponding pathogens in 6 samples. Bacteria detected in the present study therefore confirmed the suitability of the selected set of probes with the exception of enterococci. Probes targeting species of enterococci gave negative results in all mastitis samples examined. Neither Ent. faecalis nor Ent. faecium was found by FISH or culturally in this study.
In experiments with untreated mastitis milk samples, it was observed that material applied to slides tended to peel off during processing. Unfortunately, mastitis quarter milk samples do not often have a homogeneous consistency and, especially from acute cases, are often far more viscous or floccose than regular milk. Immune cells, immunoglobulins, and fibrin, commonly contained in significant amounts in mastitis milk samples, might be responsible for this effect. Furthermore, those ingredients can interfere with probes and induce unspecific binding. 8 Therefore, a previously published enzymatic clearing method9,10 for conventional milk samples, using savinase to remove proteins and lipids, was adapted for mastitis milk samples. The modifications included an extended centrifugation time and additional washing steps. The newly developed procedure improved adhesion of the samples on the slides, reduced nonspecific background staining, and also enriched the microorganisms in the pellet. For reasons of practicability, this procedure was used for all mastitis samples, including samples that were homogeneous in appearance.
Applying this optimized protocol, the detection limit of FISH was evaluated with a series of spiked UHT-milk samples containing bacteria in concentrations between 101 and 108 CFU/ml. A detection rate of 100% was observed for samples with bacterial concentrations of 106 CFU/ml or more. Detection of lower concentrations was also possible, but the reliability of FISH declined with decreasing numbers of bacteria in the milk sample. The detection and consequently the specification of fewer than 104 pathogens per ml was virtually impossible. A limit of 104 CFU/ml of milk, also reported in other studies, 10 seems rather high at first sight, but it represents the natural limit of a microscopic method as described elsewhere. 23 It was previously described that at a magnification of 1,000×, 2.5 microscopic fields have to be viewed in order to detect 1 single cell in 10 µl of a cell suspension with a concentration of 105 cells/ml. 2 In order to improve the detection rate of FISH, more milk might be used for examination, but handling of sufficiently large volumes is rather laborious and requires additional equipment.
All samples positive in culture but negative with FISH contained bacteria at or below the detection limit of FISH (Tables 3, 4). However, it should be kept in mind that clinically relevant mastitis pathogens, especially Staph. aureus, might be present in very low numbers.7,31 As an example, Staph. aureus occurred in the current study in 10 out of 40 samples in pure culture in numbers of 104 CFU/ml or fewer and was detected in only 2 of these samples. Therefore, FISH can be a reliable diagnostic tool for samples with high bacterial numbers but has to be combined with more sensitive test systems (e.g., a conventional agar-plate–based examination) for pathogens present in low numbers.
However, in addition to the qualitative detection of a particular pathogen, FISH allowed a semiquantitative estimation of bacteria. In the present study, the gradation of the bacterial amounts found with FISH in pure cultures was in accordance with results achieved by cultural examination. In addition to the determination of the presence of certain species in milk samples, the determination of the amount of bacteria present can give evidence of which bacteria may be involved in the disease, especially in cases of mixed samples.
Furthermore, FISH enables the selective detection of specific causative pathogens by applying only the corresponding oligonucleotide probe in a single test. Especially due to the use of species-specific probes, this method allows the detection of species that may be overlooked by cultural examination because of similar colony morphologies (e.g., of streptococci, enterococci, and lactococci or staphylococci and Micrococcus-like species).
In summary, FISH enables a reliable direct detection and identification of pathogens in mastitis milk samples. However, 2 major limitations of the method were observed. First, a pretreatment of mastitis milk samples with savinase was essential to achieve reliable results, but was rather laborious. Second, the detection limit of this method turned out relatively high, although within the expectable range for a microscopic method. For mastitis milk samples presenting low bacterial numbers, especially in subclinical cases, the detection limit was not satisfactory. Therefore, and despite the potential to identify pathogens very rapidly and specifically, this method should be regarded as a preliminary additional tool for the diagnosis of mastitis pathogens, and a cultural examination for confirmation will usually be required.
Footnotes
Acknowledgements
The authors thank A. Gangl and R. Huber-Schlenstedt, Tiergesundheitsdienst Bayern e.V., Poing, Germany, for making strains as well as milk samples available. Samples were also provided by P. V. Kremer, Reproduktionszentrum, Veterinary Faculty, Ludwig-Maximilians-University, Munich, Germany.
a.
Diagnostic laboratory of the Institute for Infectious Diseases and Zoonoses, Ludwig-Maximilians-University, Munich, Germany.
b.
Diagnostic laboratory of the Tiergesundheitsdienst Bayern e.V., Poing, Germany.
c.
Thermo Scientific, Ulm, Germany.
d.
MWG Biotech, Ebersberg, Germany.
e.
VWR, Darmstadt, Germany.
f.
BD Biosciences, Heidelberg, Germany.
g.
Slidex Staph Plus, bioMérieux, Nürtingen, Germany.
h.
API systems (ID 32 Staph, rapid ID 32 Strep, API Coryne, and rapid ID 32 E), bioMérieux, Nürtingen, Germany.
i.
Enterotube II ID, BD Diagnostics, Heidelberg, Germany
j.
Savinase 16.0L, Sigma-Aldrich, Taufkirchen, Germany.
k.
Roth, Karlsruhe, Germany.
l.
AppliChem, Darmstadt, Germany.
m.
Citifluor AF1, Science Services, Munich, Germany.
n.
Axioskop 2 plus, Zeiss, Göttingen, Germany.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was financially supported by the Akademie für Tiergesundheit e.V., Bonn, Germany. A. Gey received a grant for part of the study from the Bundesverband für Tiergesundheit e.V., Bonn, Germany.
