Abstract
Infectious bronchitis, a disease of chickens caused by Avian coronavirus infectious bronchitis virus (IBV), leads to severe economic losses for the poultry industry worldwide. Various attempts to control the virus based on vaccination strategies are performed. However, due to the emergence of novel genotypes, an effective control of the virus is hindered. In 1996, a novel viral genotype named IBV-QX was reported for the first time in Qingdao, Shandong province, China. The first appearance of an IBV-QX isolate in Europe was reported between 2003 and 2004 in The Netherlands. Subsequently, infections with this genotype were found in several other European countries such as France, Italy, Germany, United Kingdom, Slovenia, and Sweden. The present report describes the use of a new set of degenerate primers that amplify a 636-bp fragment within the S1 gene by reverse transcription polymerase chain reaction to detect the occurrence of IBV-QX infection in Switzerland.
Infectious bronchitis is an economically important viral disease of poultry occurring worldwide. 7 The causative agent is Avian coronavirus infectious bronchitis virus (IBV; order Nidovirales, family Coronaviridae, subfamily Coronavirinae, genus Gammacoronavirus). Clinical manifestations of the infection include respiratory distress, nephritis, and reduced egg production. 7 Because the disease represents a major economic problem within the poultry industry, various attempts to control the infection based on vaccination strategies are made.3-5,15 However, effective control of infectious bronchitis is hindered due to the considerable genetic variability of the S1 subunit of the viral spike glycoprotein, a major target of the acquired immune response.6,16,19 In a study of IBV in Switzerland in 2003, 19 of 45 tested flocks were positive in hemagglutination inhibition tests. The strains 4/91 and D274 were detected in 11 and 6 flocks, respectively, and mixed infections in 2 flocks (personal communication, RK Hoop, 2012). The emergence of a novel type of IBV, named QX, was described in Qingdao, Shandong province, China in 1996. 21 First publications concerning IBV-QX infections in poultry described proventriculitis as the main symptom. Later, severe renal pathology due to infection with IBV-QX was reported. 14 The first occurrence of the IBV-QX strain in Europe was described in The Netherlands between 2003 and 2004 followed by reports from several other countries including France, Italy, Germany, United Kingdom, Slovenia, and Sweden.1,2,10,11,13,20,22 The present report describes the detection of an IBV-QX infection in Switzerland.
In December 2011, a 2-year-old rooster of the Appenzeller Spitzhauben breed died and was submitted for necropsy. The animal, 1 of only 11 birds housed outdoor in a backyard flock, showed respiratory symptoms, apathy, and a reduced nutritional state. No evident pathological alterations were observed by postmortem examination except for an almost empty crop and intestine. Histopathological examination of kidney revealed a moderate interstitial nephritis. An infiltration of plasma cells as well as a slight edema was present in the mucosa of the trachea indicating a chronic tracheitis. Based on these findings, the presence of a nephropathogenic IBV strain was suspected. Materials from cecal tonsils and trachea as well as a cloacal swab were collected. RNA was extracted using a commercial kit a and reverse transcribed into complementary DNA (cDNA). b In order to screen for the presence of IBV RNA, a primer set described for the generic detection of coronaviruses 9 was used for amplification in combination with a commercial polymerase chain reaction (PCR) mix. c A 179-bp reverse transcription (RT)-PCR fragment was successfully amplified from RNA originating from the cecal tonsils and the cloacal swab.
To establish a novel two-step RT-PCR assay, the S1 coding sequences of 184 different IBV strains were aligned, and primers in conserved sequences were identified and chosen. The assay was performed using a cycle protocol consisting of an initial denaturation step for 3 min at 93°C, followed by 45 cycles of denaturation at 93°C for 1 min, annealing at 50°C for 1 min, and elongation at 72°C for 1 min, and a final elongation step at 72°C for 10 min. The PCR was carried out in 25-µl reactions containing (final concentrations) 12.5 µl of commercial PCR reaction mixture, c 1 µM of each primer (IBV_AV_For: 5′-TAGYGACYMTTTBGTVTGCACTATGTAG-3′ and IBV_AV_Rev: 5′-CTTGTGCDGTRCCATTAAYAAARTAR GC-3′) and 7.5 µl of reverse transcribed cDNA.
A 636-bp fragment, using the newly designed assay, was generated by the amplification of the cDNA produced from RNA originating from the cecal tonsils and the cloacal swab of the rooster. Moreover, RNA extracted from IBV strains Italy 02, 793B, 1466, D274, M41, and QX strain D388 were reverse transcribed into cDNA and could be successfully amplified with this new method, demonstrating the ability of the newly developed assay to detect non–IBV-QX as well as IBV-QX strains. The amplicon obtained from the clinical sample was sent for sequencing and analyzed for nucleotide identity using the BLAST program (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The sequence was deposited in GenBank (accession no. JX014252). The sequence of the amplicon was approximately 95% identical to the other IBV-QX strains and approximately 70–75% to the non–IBV-QX strains at the nucleotide level. This percentage of sequence identity is in accordance with results from another study reporting IBV-QX infection in a broiler flock in the United Kingdom. 11 At the amino acid level, the identity was approximately 90% to the IBV-QX strains and 65% to the non–IBV-QX strains. The possible allocation of the recently detected Swiss IBV isolate was explored by phylogenetic analysis. Based on the resulting tree (Fig. 1), the JX014252_CH isolate was found to cluster with the European QX strains. The most closely related strain was found to be the British isolate HM132098.
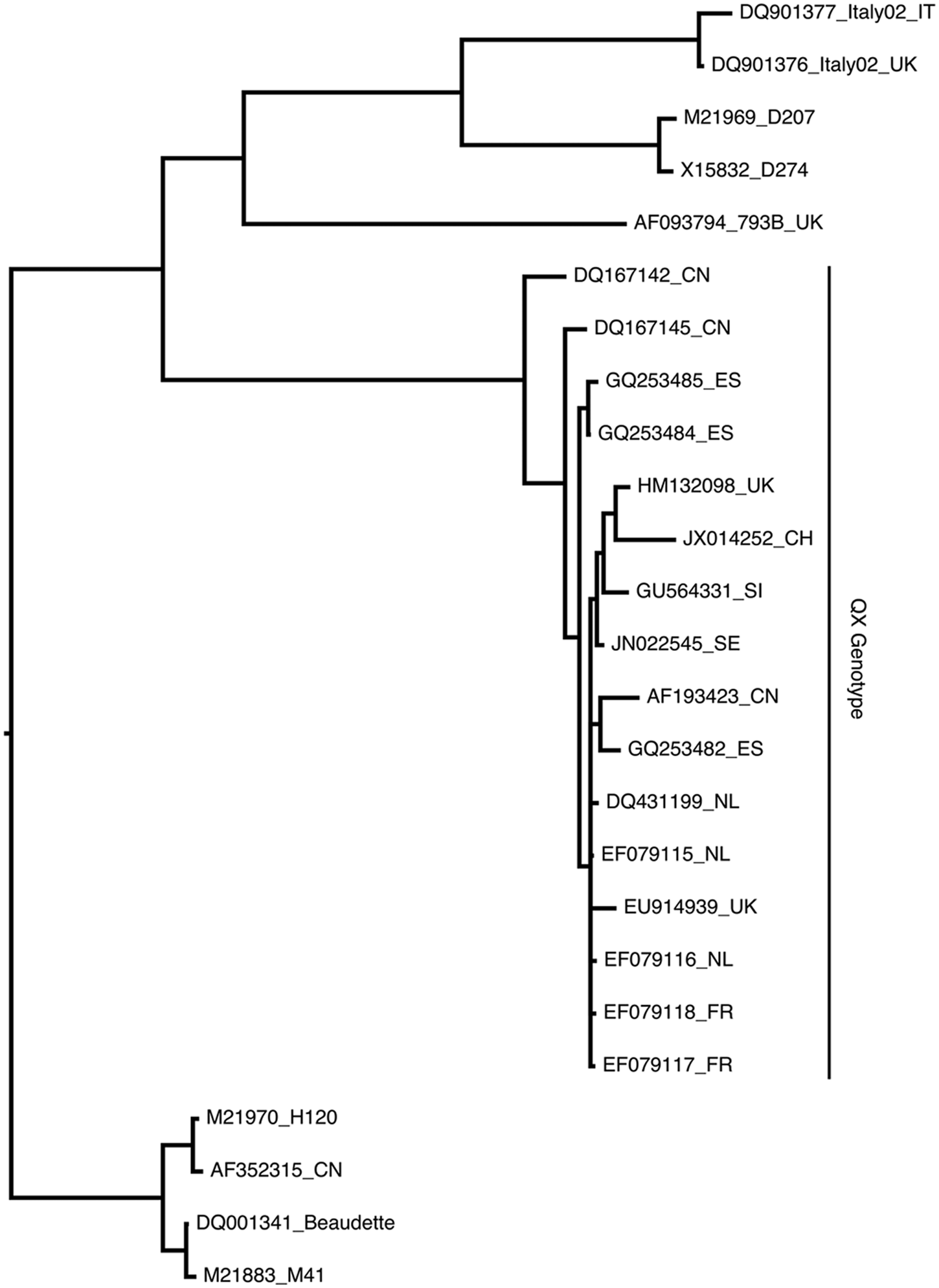
Phylogenic tree of 25 Avian coronavirus infectious bronchitis virus (IBV) strains. The partial coding sequences (corresponding to nucleotide 157–621 of DQ001341 [strain Beaudette]) for the spike proteins were aligned using MAFFT. 12 The optimal model of DNA evolution was evaluated for best fit of the data set using MODELTEST. 17 Bayesian phylogeny was inferred using MrBayes (HKYG, 10,000,000 generations). 18 Evaluation and final refinement of the tree were performed with BEAST. 8 The taxa of the tree are indicated by their GenBank accession numbers followed by the country of origin (CN: China, ES: Spain, FR: France, IT: Italy, NL: The Netherlands, SE: Sweden, SI: Slovenia, UK: United Kingdom) and/or the strain designations.
It is worth mentioning that the rooster was not vaccinated against IBV. Interestingly, several animals were newly integrated into the flock before the rooster developed clinical symptoms. Moreover, one of these birds was reported to show respiratory distress and reduced laying performance and died without any further diagnostic investigation. Therefore, the acquisition of these animals could most likely represent the reason for the introduction of the virus into the flock. The pathological findings in trachea and kidney present in the rooster sent for necropsy are in accordance with previous reports of IBV infections. Of note, cloacal and choanal swabs of 10 living animals from the same flock were taken and assayed by the novel RT-PCR assay. The amplification of the cDNA produced from RNA originating from 3 samples resulted in a positive signal. In addition, 1 PCR product was sent for sequencing, and the presence of IBV-QX infection in the flock was confirmed by the result obtained.
In conclusion, the detection and characterization of an IBV-QX infection in a Swiss poultry backyard flock became possible by applying a newly designed two-step RT-PCR assay in combination with sequencing. This novel approach might represent a valuable diagnostic tool to further evaluate IBV infection status not only in the backyard but also in commercial Swiss poultry flocks with respect to newly emerging IBV strains.
Footnotes
Acknowledgements
The authors thank Marius Birrer for collecting clinical samples. Richard Jones and Volker Thiel are also acknowledged for helpful discussions and suggestions.
a.
NucleoSpin RNA II kit, Macherey-Nagel GmbH & Co., Duren, Germany.
b.
Reverse transcription system, Promega Corp., Madison, WI.
c.
REDTaq ReadyMix, Sigma-Aldrich, St. Louis, MO.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
