Abstract
The purpose of the current study was to investigate the prevalence and serovar distribution of Salmonella isolates in cobras and their environment at a snake park. A total of 166 fecal or intestinal samples were examined, comprising 39 samples from captive cobras (Naja kaouthia), 70 from recently wild-caught cobras, 19 from wild-caught cobras that had been kept on the farm for over 3 months, 18 from mice (Mus musculus), 12 from frogs (Hoplobatrachus rugulosus), and 8 from farm workers. Specific serological identification was performed, and pulsed-field gel electrophoresis (PFGE) was utilized for DNA analysis. Out of all snakes (n = 128), 20 of the 30 animals used for snake food and 3 of the 8 samples from personnel were positive for Salmonella spp. There were 228 Salmonella isolates, with a total of 29 serovars from subspecies I and IIIb, composed of 24 serovars from cobras and 5 from the other sources. Salmonella Amsterdam was predominant in captive-born and captive cobras, followed by S. Poona and S. Bareilly, respectively (P < 0.05). Salmonella I 4,[5],12:i:- was the sole serovar detected from the mice, while 3 serovars including Ramatgan, I 4,[5],12:e,h:-, and rough strain were detected only from frogs (P < 0.001). Salmonella Derby was only detected in workers. On the basis of the PFGE results, evidence of movement of isolates between human beings and snakes, and between snakes and frogs, was found for S. Poona and S. Wandsworth, respectively. The study suggests that Salmonella spp. act as true residents in the intestinal tract of cobras with high risk of environmental contamination through fecal shedding.
Introduction
Salmonella spp. are well-known bacterial pathogens that cause food poisoning in human beings and animals. 6 Fecal and blood contamination from reptiles has been believed to be a major source of salmonellosis in human beings, 28 especially those who consume meat or other products made from snakes, and also those who keep domesticate snakes as pets. 8 Cold-blooded animals are known to be carriers of Salmonella, especially carnivorous reptiles, which may intermittently shed into the environment. 3 Studies of Salmonella prevalence have found that approximately 40–100% of amphibians and reptiles carry Salmonella in their intestinal tracts, and may spread these in their feces.11,24 Septicemia, severe diarrhea, and fatal salmonellosis caused by contamination from reptiles have been reported in several countries.4,22 Thus, transmission of Salmonella from reptiles should be regarded as being of public health significance. 11
Materials and methods
Background of the snake park system
The snake park of the Queen Saovabha Memorial Institute (Bangkok, Thailand), the biggest snake farm in Thailand, is run for the production of antitoxin and as a reptile zoo for the public. Over 1,200 captive (born on the farm) snakes, including more than 15 common breeds, are kept in ponds or cages at a stocking rate of approximately 1–2 m2/snake. In addition, the farm holds snakes that have been caught from private homes in rural areas of Thailand and submitted to the farm for population replacement (wild-caught snakes). Wild-caught snakes are quarantined for 1 month before they are mixed with other captive snakes in a snake pond. After 3 months in captivity, such wild-caught snakes are designated as wild-caught captive. Caretakers routinely handle snakes during feeding, venom squeezing, and cage cleaning. Veterinarians also occasionally perform physical examinations and treatments. Only hand washing with ordinary soap is performed following snake handling. Frozen frogs (Hoplobatrachus rugulosus) and live mice (Mus musculus) derived from the same market for the past 10 years are used to feed the snakes. Generally, all snakes are fed once a week.
Sample collection
Between May 2007 and July 2009, at least 5 ml of fecal contents were randomly collected from the cloacae of 128 cobras (Naja kaouthia) by direct abdominal squeezing and digital manipulation. The cobras consisted of 39 captive cobras from 39 cages, 70 wild-caught cobras from 12 snake ponds (5–10 snakes per month/collection [i.e., 1 collection per month]), and 19 wild-caught captive cobras from 19 cages. All captive snakes were over 1 year of age and were of normal health status as judged by their behavior, appetite, respiratory rate, and skin appearance including lack of external parasite infestation. At least 10 g each of 18 feeder mice feces and 12 intestine tracts from frozen frogs were examined, as well as 8 fecal samples from 6 caretakers and 2 veterinarians on the farm. All samples were transported in sterile containers at 4°C and submitted to the laboratory of the Department of Veterinary Microbiology, Faculty of Veterinary Science, Chulalongkorn University (Bangkok, Thailand) within 24 hr.
Specimen processing and Salmonella serovar identification
Salmonella serovar isolation and identification was performed according to the methods recommended by the International Organization for Standardization (ISO) 6579, with minor modifications. 14 Fecal contents were pre-enriched in buffered peptone water, a overnight. The suspension was transferred into 3 enrichment broths: selenite F broth, a tetrathionate broth, a and Rappaport–Vassiliadis (RV) broth a at a 1:9 ratio and were incubated at 37°C for 24 hr, except for the RV broth, which was incubated at 42°C. 1 Following this incubation, the broths were incubated on modified semisolid RV medium a at 42°C for 24 hr, and xylose lysine deoxycholate agar a and brilliant green agar b supplemented with 1 mg/ml of novobiocin c at 37°C for 18–24 hr. Up to 3 suspect Salmonella colonies per sample were collected for serological identification. 12 For confirmation, all colonies were examined in a routine biochemical scheme using the triple sugar iron test a and motile indole lysine medium a as recommended. 19
Serological examination
For serovar identification, a single colony was used in a slide agglutination test using polyvalent antisera A-I, polyvalent OMA, polyvalent OMB, polyvalent OMC, polyvalent OMD, polyvalent OME, polyvalent OMF, and polyvalent OMG, d against somatic (O) Salmonella antigens. 15 After this screening, the isolates were serologically identified using all specific O- and H-antisera according to the Gard technique 17 and interpreted according to the principles of the Kauffmann–White scheme. 12
DNA profile analysis
Serovars common to both human beings and snakes or snake and feeds were analyzed by pulsed-field gel electrophoresis (PFGE) according to a previously published protocol. 2 Salmonella serovar Braenderup strain H9812 was used as the standard. The digested DNA samples were electrophoresed in a 1% molecular grade agarose gel in Tris–borate– ethylenediamine tetra-acetic acid buffer using a commercial system. e After staining with ethidium bromide, DNA fingerprints were visualized under ultraviolet illumination and documented electronically as a digital image. The sizes of DNA bands were detected and calculated by PhotoCaptMW program, f and band patterns were interpreted according to the approved criteria. 30 The band patterns of DNA fragments were compared by similarity matrix and clustered by unweighted pair-group method with arithmetic average for dendrogram construction using the online DendroUPGMA program (http://genomes.urv.cat/UPGMA). 18
Statistical analysis
The statistical analyses were conducted by using SAS. g The hosts of origin were classified into 6 groups (i.e., captive cobra, wild-caught captive cobra, wild-caught cobra, human being, frog, and mouse). For each serovar, 2 × 2 contingency tables were used between 2 host groups (e.g., captive cobra vs. wild-caught captive cobra, captive cobra vs. human being) and the isolation of the respective serovar (found/not found). Data were presented as the percentage of the occurrence of Salmonella serovars in each host group. The prevalence of each Salmonella serovar was compared among the various hosts of origin by using either chi-square analysis or Fisher exact test. Values of P < 0.05 were defined as statistically significant.
Results
The overall prevalence of Salmonella spp. in cobras, feeder frozen frogs, and mice were 100.0%, 91.6%, and 50.0%, respectively. Only 3 of the 8 human samples, 2 from caretakers and 1 from a veterinarian, were found positive (Table 1). Two subspecies (I and IIIb) and 29 serovars were identified (Table 2).
The prevalence of Salmonella in different sources at Queen Saovabha Snake Park, Thailand.
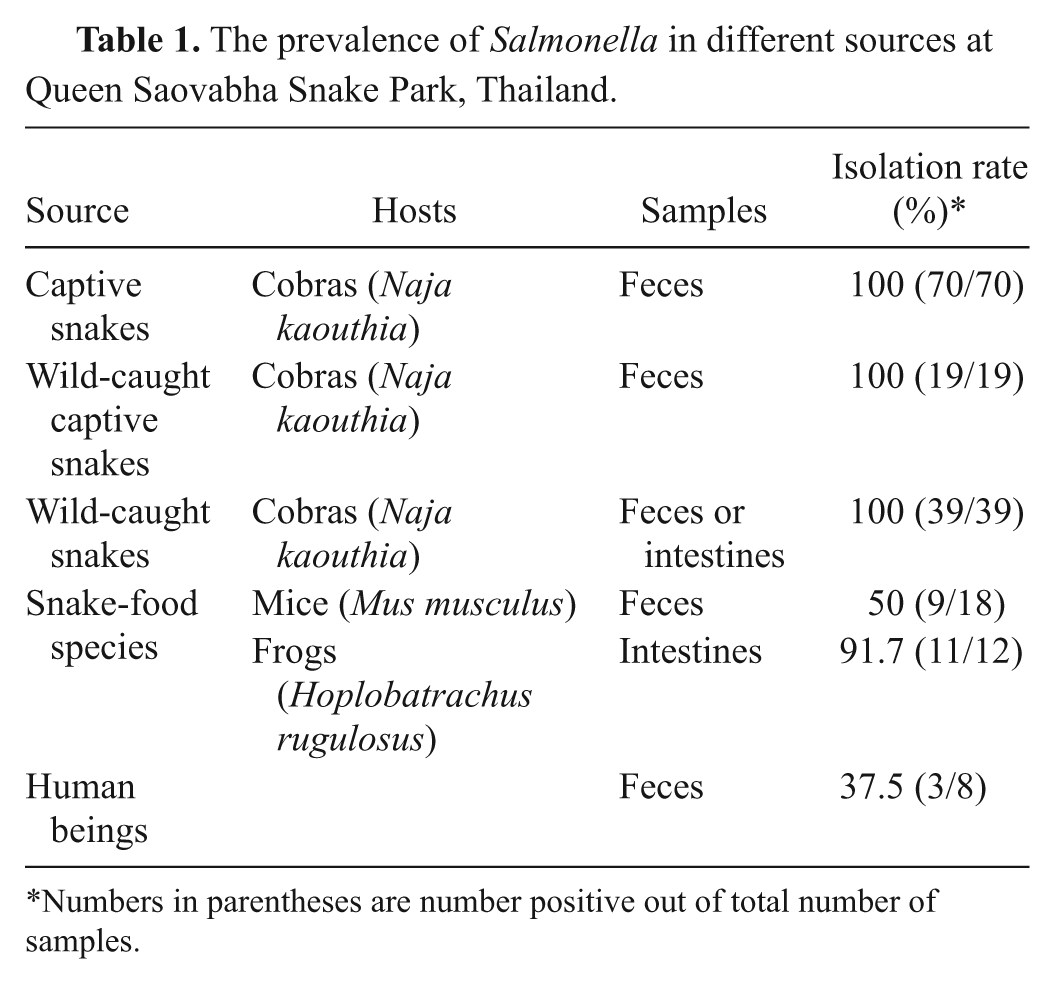
Numbers in parentheses are number positive out of total number of samples.
Comparison of serovar diversity of Salmonella spp. in captive cobras, wild-caught cobras, snake-food animals, and snake-associated workers at Queen Saovabha Snake Park, Thailand.*
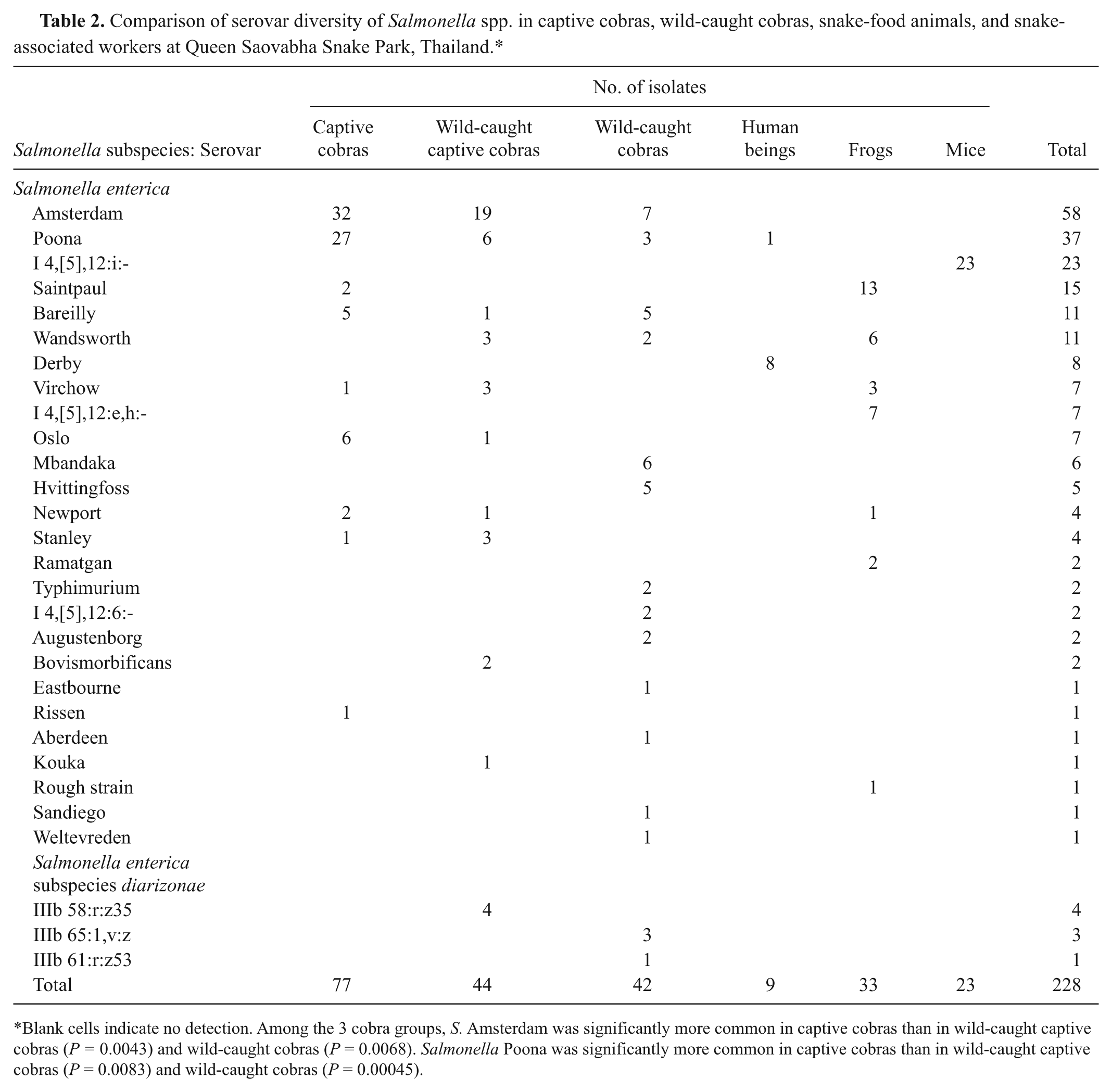
Blank cells indicate no detection. Among the 3 cobra groups, S. Amsterdam was significantly more common in captive cobras than in wild-caught captive cobras (P = 0.0043) and wild-caught cobras (P = 0.0068). Salmonella Poona was significantly more common in captive cobras than in wild-caught captive cobras (P = 0.0083) and wild-caught cobras (P = 0.00045).
The distribution of serovars in each host group is presented in Table 2. A total of 228 isolates were examined, and these were predominantly identified as S. Amsterdam (25.4%), followed by S. Poona (16.2%), S. I 4,[5],12:i:- (10%), S. Saintpaul (6.6%), S. Bareilly (4.8%), and S. Wandsworth (4.8%). All isolates derived from mouse feces were S. I 4,[5],12:i:-, which was significantly different from the situation in snakes, frogs, and human beings (P < 0.001). On the other hand, S. Virchow and S. Newport were detected in both frogs (9% and 3%, respectively) and snakes (2.4% and 1.8%, respectively, P = 0.6) and S. I 4,[5],12:e,h:-, S. Ramatgan, and rough strains were detected only in frogs (21.2%). The number of S. Saintpaul from frogs (39.3%) was higher than in cobras (1.6%; P < 0.0001). Salmonella Derby was detected in 3 human beings, 1 of whom also was colonized with S. Poona. Salmonella enterica subspecies diarizonae serovars IIIb 58:r:z35, IIIb 65:l,v:z, and IIIb 61:r:z53 were detected in wild-caught cobras. Among captive cobras, the prevalence of S. Amsterdam (42.1%) and S. Poona (27.2%) was higher than in wild-caught cobras (16.6% and 7.1%, respectively; P < 0.01) and the other hosts (Table 2).
Differences in the DNA fingerprints of serovars Newport, Poona, Wandsworth, and Virchow are shown in Table 3 and Figure 1, with Figure 1 also showing the patterns obtained with S. Derby. Based on the different of patterns, at least 2 representative patterns of each serovar from animals and all isolates from human beings are presented. The S. Poona derived from human beings and captive cobras were closely related whereas the serovars from wild-caught cobras all showed different patterns to those from human beings. The S. Derby isolates from human beings were essentially indistinguishable with only 1 isolate showing a minor difference. The S. Newport isolates from captive cobras were identical but different from those obtained from wild-caught cobras. On the other hand, the sole isolate from the frog was a nontypeable strain (no separated band). The S. Virchow isolates from frogs were possibly related to those obtained from captive cobras but were not related to the isolates from the wild-caught cobras. The S. Wandsworth isolates from wild-caught cobras were indistinguishable to those obtained from wild-caught captive cobras and frogs. Isolates of S. Saintpaul could not be typed by this PFGE technique.
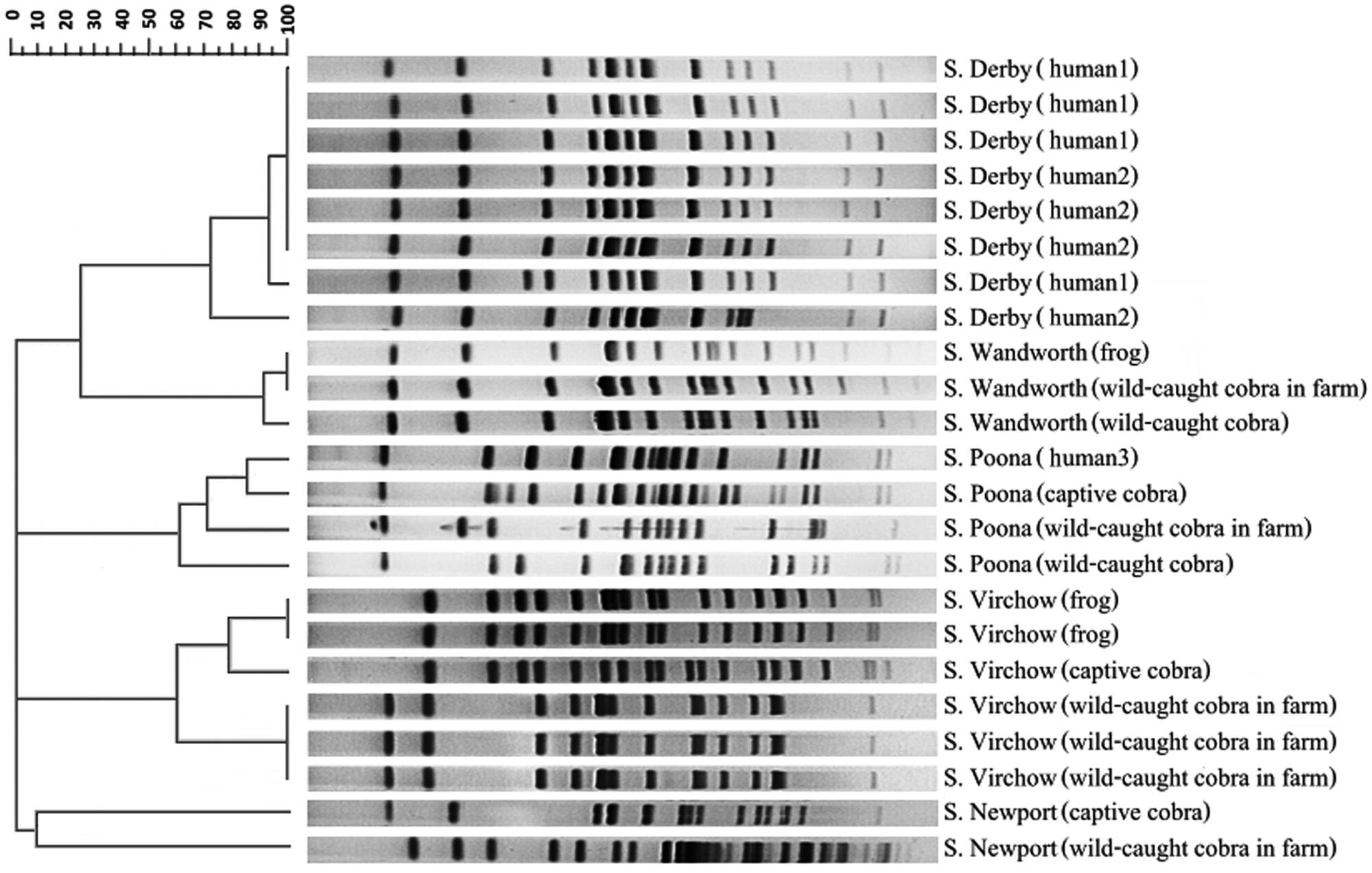
Patterns of XbaI-digested chromosomal DNA of selected isolates of Salmonella serovars Poona, Saintpaul, Wandsworth, and Newport on pulsed-field gel electrophoresis analysis by dendrogram alignment.
Comparison of pulsed-field gel electrophoresis (PFGE) band difference between 4 serovars isolated from human beings, cobras, and frogs at Queen Saovabha Snake Park, Thailand.*
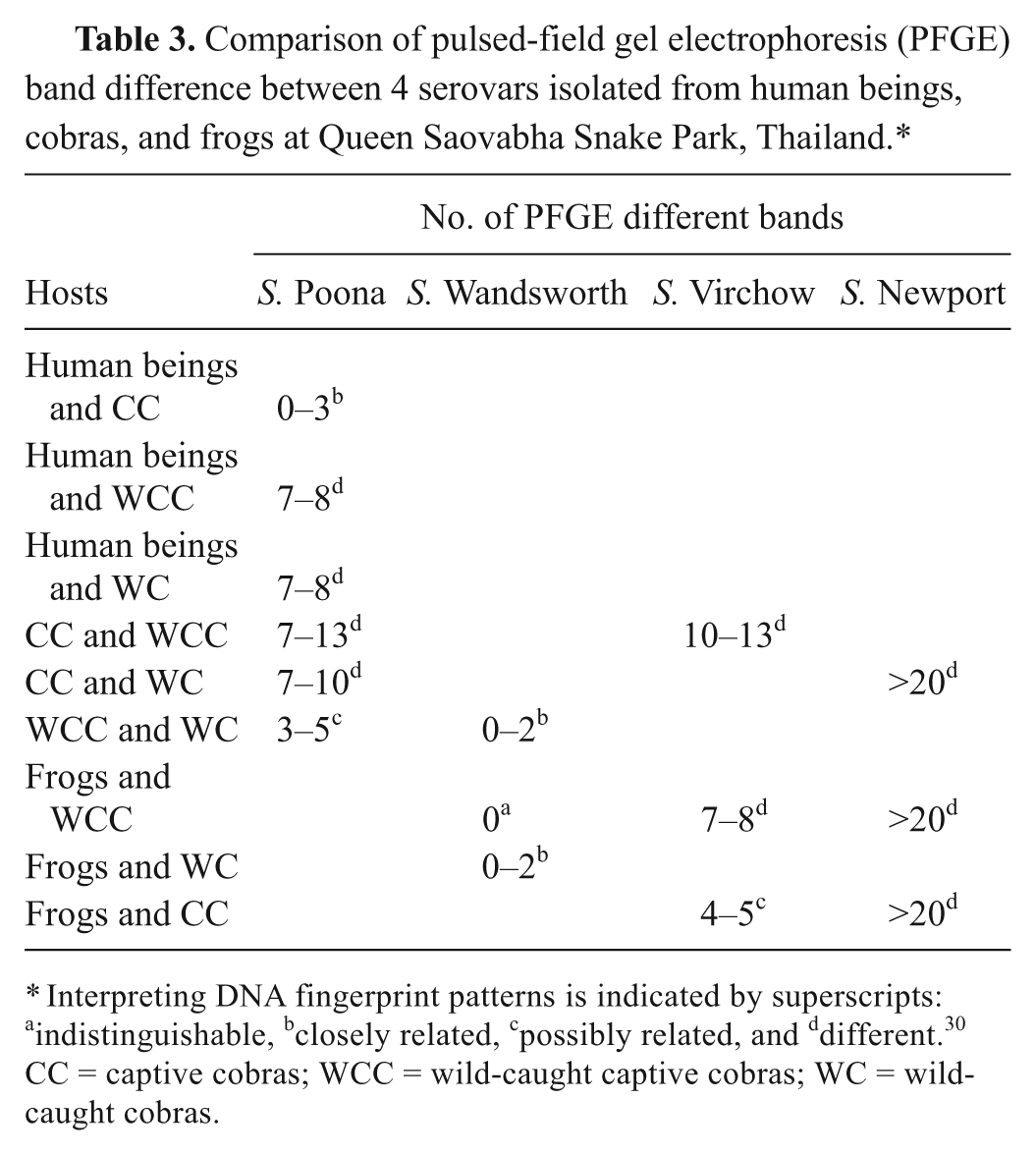

Interpreting DNA fingerprint patterns is indicated by superscripts: aindistinguishable, bclosely related, cpossibly related, and ddifferent.30 CC = captive cobras; WCC = wild-caught captive cobras; WC = wild-caught cobras.
Discussion
Previously, surveillance of Salmonella spp. from animal sources in Thailand has concentrated on food animals, with rates of 63% in slaughtered pigs 9 and 57% in chicken meat.26,27 Recently, reptile-associated salmonellosis has been reported in European countries, the United States, and Australia, where serovars Enteritidis, Typhimurium, Urbana, and Rubislaw, and Salmonella subspecies IIIa, subspecies IIIb (diarizonae), and subspecies IV (houtenae) were frequently obtained from patients with watery diarrhea and extra-gastrointestinal infection such as pleuritis, lymphadenitis, arthritis, meningitis, and cystitis, particularly in infants and immunocompromised hosts.16,21,23 Transmission of Salmonella spp. from pet reptiles to human beings could occur from direct contact and inappropriate hygiene by the owner after touching the pets. 29 Moreover, consumption of snake meat or products can be a direct source of human infection. 20 In Thailand, snake meat has been incorporated as a major ingredient in cooked and uncooked dishes for people in rural areas. Furthermore, raw cobra bile and blood mixed in liquor are popular potions that are believed to boost strength. Therefore, these practices pose a risk of zoonotic infection as all cobras tested in the current study contained Salmonella spp. While there has been no report of human salmonellosis associated with reptiles in Thailand, the current study provides evidence of the potential problem and highlights the need for good hygiene when handling snakes.
The role of Salmonella in snakes and frogs has not been elucidated, but Salmonella spp. may be resident or transient inhabitants of the intestine, liver, and gall bladder. 24 Serovars Amsterdam, Poona, and Bareilly were commonly found in captive cobras, as well as in wild-caught cobras. There was a much greater diversity of serovars found in wild-caught cobras, and these potentially could be introduced into the community at the park. The PFGE analysis provided evidence of movement of isolates between human beings and snakes, and snakes and frogs for S. Poona and S. Wandsworth, respectively. Transmission of Salmonella spp. into the farm did not appear to be associated with feeder mice, as these were all colonized by serovar I 4,[5],12:i:-, which was not found in the snakes. However, serovar S. I 4,[5],12:i:- has been identified in Taiwanese reptiles, 7 and a previous outbreak in snakes has been recorded associated with S. Typhimurium from feeder mice. 13 Regarding transmission to human beings, routine touching of snakes was not likely to be a common source of infection since the isolates from the staff were mainly S. Derby, and this serovar was not detected in the snakes. Thus, washing with ordinary soap after exposure to snakes may reduce the risks of transmission. 29 On the other hand, eating snake meat or exposure to their environment and products represents a high risk for salmonellosis. 10
The prevalence of Salmonella in cobras was very high compared to previous studies.7,25 Possibly the use of enrichment media including RV broth, tetrathionate broth, and selenite F broth enhanced the Salmonella recovery rate in the current study. Selenite F broth especially has been recommended for recovery of serovars harbored by reptiles. 1 Moreover, the use of at least 3 Salmonella-like colonies from each selective media increases the possibility of identifying colonized snakes and revealing the serovar diversity in the same sample.
Serovars Amsterdam, Poona, and Bareilly were the 3 most common Salmonella enterica serovars isolated from cobras in the present study, whereas serovars Heron, Bredeney, Treforest, I 4,[5],12:i:-, and Typhimurium were shown to be common in reptiles from Taiwan, 7 and serovars Bardo, Newport, and Panama were the predominant serovars in Japan. 25 The diversity of serovars apparently depends upon geographical area, and this information could be used to form a useful database for epidemiological investigations. In other countries, human salmonellosis due to S. Rubislaw, S. Paratyphi B, S. Urbana, and S. Typhimurium has been associated with reptile contamination.5,23
In conclusion, all snakes in a Thai snake farm were colonized with Salmonella, with the isolates consisting of 29 serovars, the majority belonging to subspecies enterica and a minority belonging to subspecies diarizonae. Salmonella bongori was not detected in the present study. The serovars detected in farm workers and feeder mice were distinct from those in cobras. Evidence of the transmission of isolates between human beings and snakes, and between snakes and frogs, was found for S. Poona and S. Wandsworth, respectively. Cobras represent a potentially important source of Salmonella contamination for human beings, especially for those people in situations of poor hygiene or working and/or living where snake meat or products are consumed.
Footnotes
Acknowledgements
The authors thank the staff from Snake Park of the Queen Saovabha Memorial Institute (Bangkok, Thailand) for assistance with obtaining isolates and the WHO National Salmonella and Shigella Center National Institute of Health, Department of Medical Sciences for serological analysis. The authors would also like to thank Dr. David Hampson of Murdoch University for assistance during preparation of the manuscript and Ms. Waree Niyomthum for assistance with the PFGE procedure.
a.
Difco™, Becton Dickinson France SAS, Le Pont-De-Claix, France.
b.
Mast Diagnostics Ltd, Bootle, United Kingdom.
c.
Sigma-Aldrich, St. Louis, MO.
d.
Serosystem, Clinag, Thailand.
e.
CHEF-DR® II, Bio-Rad Laboratories, Hercules, CA.
f.
PhotoCaptMW program, Vilber-Lourmat, Marne-la-Vallée, France.
g.
version 9.0, SAS Institute Inc., Cary, NC.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research project was supported by the Academic Affairs, Faculty of Veterinary Science, Chulalongkorn University during 2008–2010.
