Abstract
Three tests for typing clinical isolates of Actinobacillus pleuropneumoniae biovar 2 were compared: 1) standard coagglutination with type-specific antisera against serovars 1–12 of biovar 1 of A. pleuropneumoniae; 2) a previously described polymerase chain reaction system for detecting the apx genes encoding the ApxI, ApxII, and ApxIII toxins in A. pleuropneumoniae; and 3) a restriction fragment length polymorphism analysis of the transferrin-binding protein B gene. The panel of strains tested included 112 field isolates of biovar 2 recovered from pigs between 1979 and 2007 in Italy and Spain, and reference strains for all described serovars of both biovars. The values of Simpson index of diversity obtained for the 3 methods were 0.68, 0.20, and 0.60, respectively. Coagglutination assays identified the field isolates as belonging to serovars 2 (9 strains), 4 (13 strains), 7 (61 strains), 9 (17 strains), and 11 (1 strain). Eleven strains were not typeable, and cross-reactivity was observed between serovars 2 and 4, 4 and 7, and 9 and 11. Isolates of A. pleuropneumoniae biovar 2 displayed 2 apx patterns: ApxII+ (94 strains) and ApxI+/ApxII+ (18 strains). The restriction fragment length polymorphism analysis assigned the strains tested to 3 different patterns. This method distinguished between biovar 2 reference strains and field strains that could not be identified by other methods, thus constituting a useful complementary test for the typing of A. pleuropneumoniae biovar 2.
Actinobacillus pleuropneumoniae causes porcine pleuropneumonia, a contagious disease that may lead to sudden death or poor performance in infected growing pigs, and thus contributes significantly to economic losses in industrialized pig production. Biovar 1 (nicotinamide adenine dinucleotide [NAD]-dependent) and biovar 2 (NAD-independent) strains of A. pleuropneumoniae can be distinguished according to the requirement for β-NAD for growth. 20 Each of the biovars can be subdivided into a number of serovars. The current serotyping scheme applies to both biovar 1 and 2 and recognizes 15 serovars, with serovars 13 and 14 exclusive to biovar 2. Several antigenic assays for biovar 1 serotyping have been described. 18,22 However, the usefulness of these established assays for typing biovar 2 isolates remains to be determined. Nevertheless, the same panel of antisera raised against biovar 1 reference strains are commonly used to type field isolates of the biovar 2 cluster. In this procedure, some biovar 2 isolates react strongly with biovar 1 serovars 2, 4, 7, and 9. 16 However, nontypeable biovar 2 strains and strains with antigenic determinants common to more than one serovar in biovar 1 have been described, 16 rendering this methodology unsuitable for the typing of biovar 2 isolates in general.
DNA-based molecular assays for detecting and typing A. pleuropneumoniae have been described, and some have been proposed as additional tools to support antigen-based typing techniques. 4,9,11,13,14,21,23 The DNA-based methods are based on the amplification and analysis of the nucleotide sequences of 1 or more genes, and allow either individual serovar identification or the establishment of groups of genetically related serovars. However, molecular typing methods have not been extensively used for classifying A. pleuropneumoniae biovar 2 isolates.
Actinobacillus pleuropneumoniae isolates display different toxin gene (apx) patterns. The apx patterns are inherent to a given serovar and play an important role in virulence. 1 Hence, typing A. pleuropneumoniae on the basis of apx toxin profile enhances and facilitates differentiation between A. pleuropneumoniae isolates according to virulence properties. 10 In the current study, the polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) method based on the variability of the transferrin-binding protein B (tbpB) gene encoding the transferrin-binding protein B, was adapted for the typing of A. pleuropneumoniae biovar 2 isolates. 7 The discriminatory power of this method was compared with those of serotyping and PCR toxin typing for a sample of 112 biovar 2 field isolates from Spain and Italy.
Typing assays were validated by testing the following panel of bacteria: 1) 112 biovar 2 field isolates recovered from pigs between 1979 and 2007 in Spain and Italy; 2) the A. pleuropneumoniae reference strains for serovars 1–12 and serovar 15 (all biovar 1); and 3) the A. pleuropneumoniae reference strains for serovars 13 (N-273) and 14 (3906; both biovar 2). Additionally, a reference biovar 2 strain (N-282, known to be serovar 2), 19 and a Spanish field isolate (45687), which has been typed as biovar 2 serovar 14 by an external laboratory (University of Montreal, Canada), were also used. All strains were plated on chocolate agar and incubated overnight at 37°C under aerobic conditions. Phenotypic identification of A. pleuropneumoniae to species level was based upon previous recommendations. 5 Identification was confirmed by single multiplex PCR with a commercially available PCR mixture. a The PCR test for the species-specific detection of A. pleuropneumoniae biovar 1 and 2 sequences has been shown to be suitable and to perform well. 3,8
Antigen-based typing was based on coagglutination (CoA) tests with a panel of type-specific antisera for serovars 1–12 b of biovar 1, as previously described. 17 All isolates giving a single, serovar-specific reaction within 30 sec were assigned to the serovar concerned. Isolates cross reacting with 2 or more serovars were considered nontypeable (NT).
Whole-cell DNA was isolated from exponential growth phase cultures of the A. pleuropneumoniae strains, with a spin-column procedure based on the use of a silica membrane–based isolation kit. c The apx gene patterns were obtained with a previously described PCR apx typing system. 9 A previously described tbpB gene PCR–based RFLP method was adapted. 7 This method exploits the variability of the tbpB gene, which encodes a 60-kDa protein component of the A. pleuropneumoniae transferrin receptor. The TbpB protein differs considerably between A. pleuropneumoniae serovars and induces a serovar-specific protective immune response. 6,24 The tbpB genes were amplified using specific primers, and the PCR amplicons were digested with AvaII as previously described. 7 The PCR-RFLP patterns were compared visually, and fragment patterns were arbitrarily named with an uppercase letter (A–Z).
To quantify the discriminatory ability of the 3 typing systems in the current study, the D value for each system was calculated as previously described. 12 In this approach, the closer a D value is to 1, the more discrimination is being achieved.
The CoA assay assigned a single serovar to 101 (90.2%) of the 112 A. pleuropneumoniae biovar 2 isolates tested. Serovars 2 (9/101), 4 (13/101), 7 (61/101), 9 (17/101), and 11 (1/101) were identified, and the D value was 0.68. The remaining 11 strains (9.8% of the sample) were NT as they either cross reacted with serovars 2–4, 6–9, or 11, in various combinations, or displayed no reaction at all. The most common cross-reactions were those between serovars 2 and 4, serovars 4 and 7, and serovars 9 and 11. The A. pleuropneumoniae biovar 2 reference strain N-273 reacted strongly with biovar 1 serovar 7 antiserum, whereas strain N-282 reacted with biovar 1 serovar 11 antiserum, and strain 3906 was not typeable. There was some influence of geography on the distribution of the CoA serovars with serovar 7 being dominant in Spain and serovar 9 dominant in Italy (Table 1).
Typing results for 112 biovar 2 Actinobacillus pleuropneumoniae field strains obtained by coagglutination (CoA), transferrin-binding protein B (tbpB) gene polymerase chain reaction (PCR)-based restriction fragment length polymorphism (RFLP), and A. pleuropneumoniae RTX (Repeats in the structural ToXin; Apx) toxin PCR.*
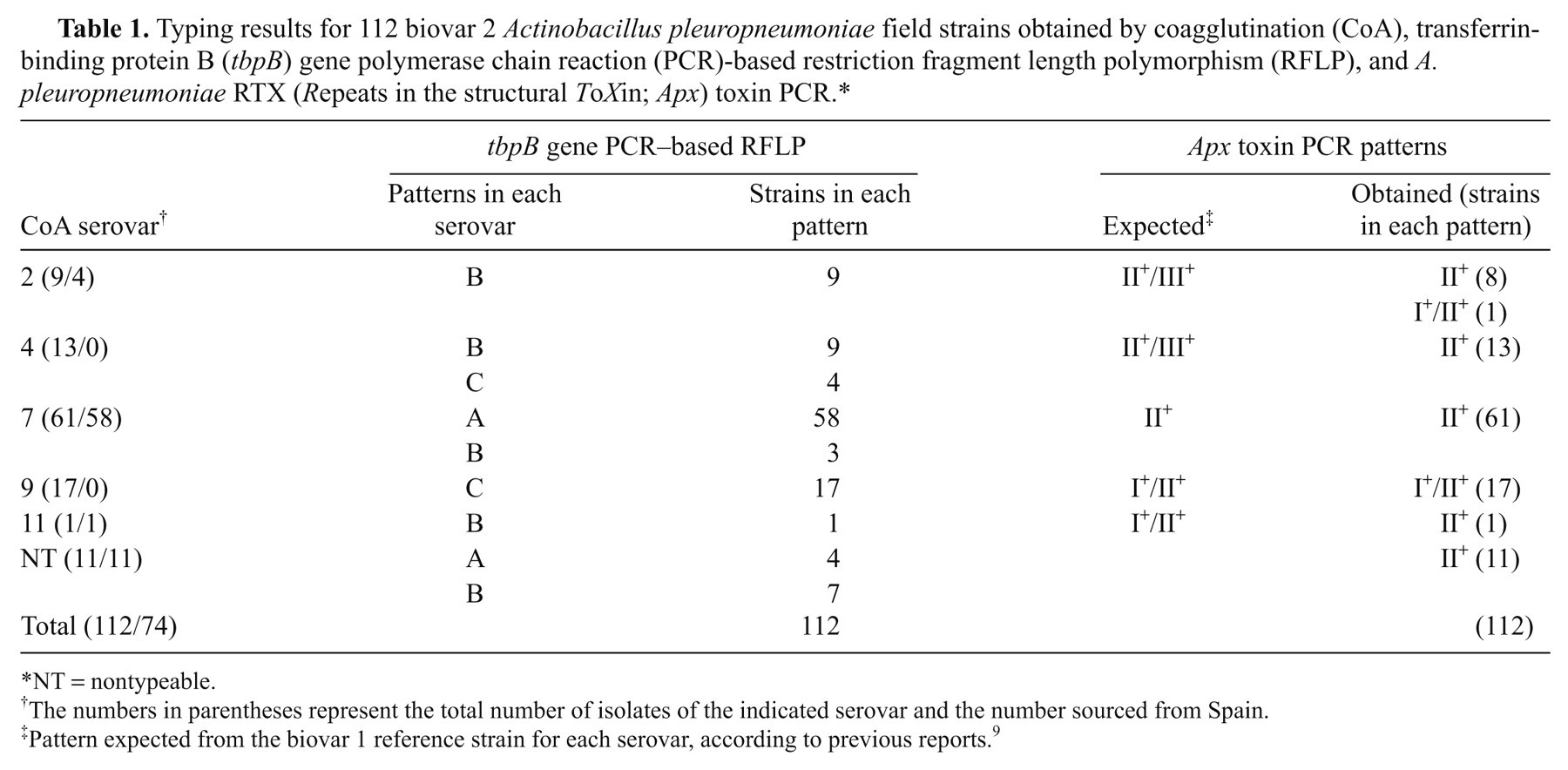
NT = nontypeable.
The numbers in parentheses represent the total number of isolates of the indicated serovar and the number sourced from Spain.
Pattern expected from the biovar 1 reference strain for each serovar, according to previous reports. 9
The results of PCR apx typing with 112 biovar 2 field strains are shown in Table 1. None of the strains carried genes from the apxIII operon. Strains in CoA serovars 2, 4, and 11 had gene patterns different from those of their biovar 1 counterparts, whereas isolates of CoA serovars 7 and 9 had gene patterns identical to those of their biovar 1 counterparts. Reference strains N-273 (biovar 2 serovar 13) and N-282 (biovar 2 serovar 2) carried out the genes of the apxII operon only. Strain 3906 (biovar 2 serovar 14) had the characteristic pattern associated with the ApxI toxin, and was unique in this respect among all the biovar 2 strains analyzed. Only 2 different gene patterns were obtained for the 112 field isolates tested, resulting in a low D value (0.28).
The tbpB gene PCR–based RFLP method generated 3 easily distinguished patterns (Table 2; Fig. 1). Strain N-273 was assigned to pattern A (fragments of 0.81, 0.48, and 0.35 kb), strain 45687 to pattern B (0.83, 0.78, and 0.06 kb), and strain 3906 to pattern C (0.97, 0.71, and 0.06 kb). All the field strains were tested with this assay and each gave a single pattern (Table 2). The patterns showed an influence of geography; all pattern A strains were Spanish and all pattern C strains were Italian. The D value for this method was 0.60.
Distribution of restriction patterns for 112 nicotinamide adenine dinucleotide (NAD)-independent unrelated field isolates, 3 NAD-independent collection strains, and 13 NAD-dependent reference strains of Actinobacillus pleuropneumoniae: analysis of the complete transferrin binding protein B (tbpB) gene with the AvaII enzyme.
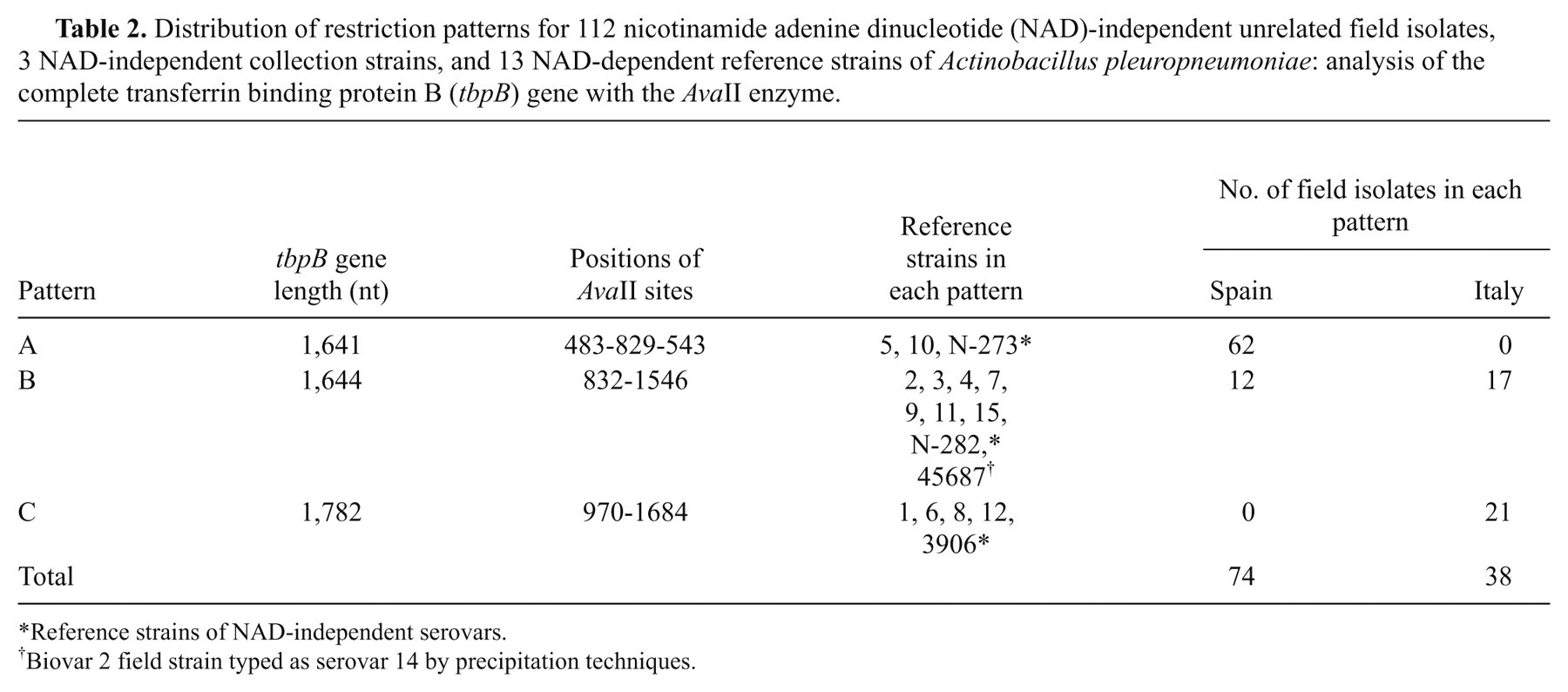
Reference strains of NAD-independent serovars.
Biovar 2 field strain typed as serovar 14 by precipitation techniques.
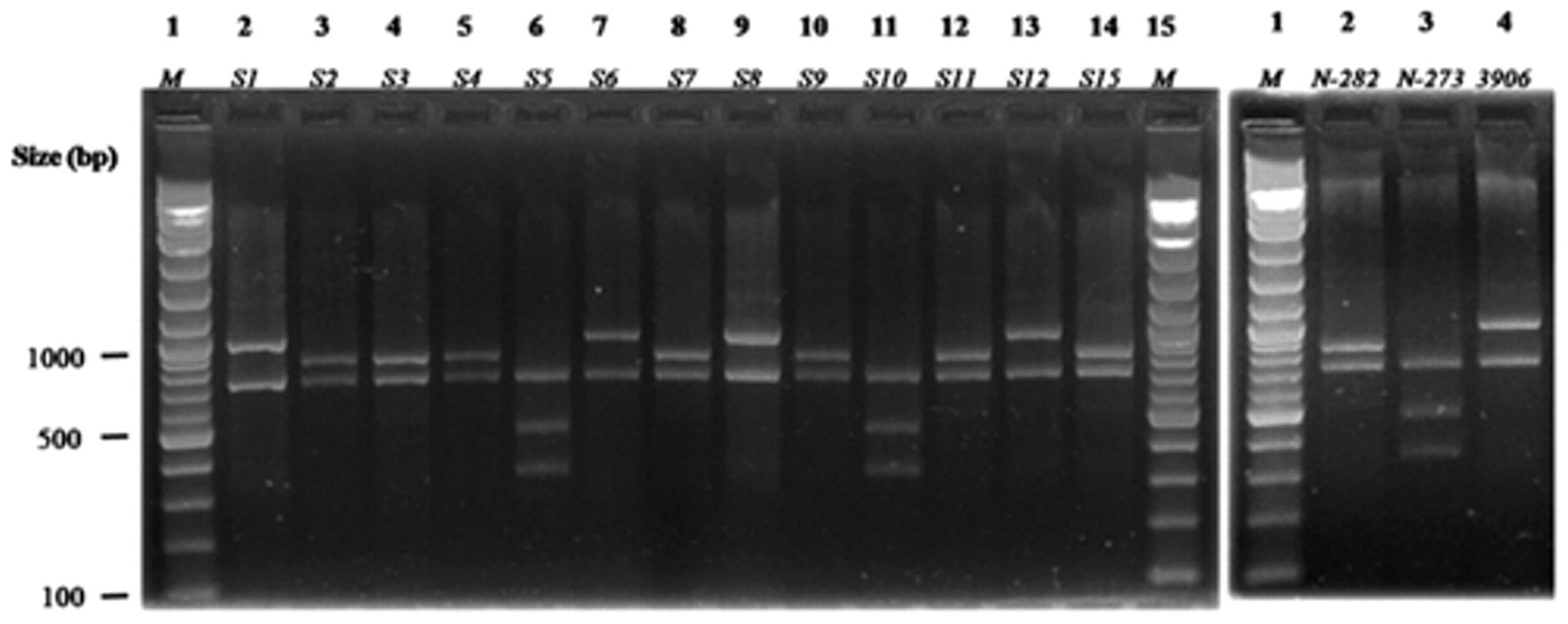
Banding patterns of the complete transferrin-binding protein B (tbpB) gene in field isolates of Actinobacillus pleuropneumoniae and reference strains after digestion with AvaII: left panel, lanes 2–14 correspond to biovar 1 serovars (S)1–12 and S15; right panel, lanes 2–4 correspond to biovar 2 S2 (N-282), S13 (N-273), and S14 (3906). The products were run on 2% agarose gel with a 100-bp ladder as the molecular size marker (M).
In the current study, 3 different methods (CoA typing, PCR apx typing, and tbpB gene PCR–based RFLP typing) were compared, for a sample of 112 A. pleuropneumoniae biovar 2 field isolates, covering a wide geographic and temporal range. The highest D value was obtained with the CoA assay, which may therefore be considered the most appropriate for typing. However, 11 isolates (9.8%) could not be assigned to a serovar by the CoA method, due to cross-reactions with more than one serovar or an absence of reaction. Many of the A. pleuropneumoniae biovar 2 isolates that could be classified by CoA assays (61/101), together with strain N-273 (serovar 13), reacted strongly with biovar 1 serovar 7. This finding is consistent with the previously reported reaction of reference serovar 13 antiserum with the reference serovar 7 antigen in the ring precipitation test and in diffusion tests. 2 The cross-reactivity presumably results principally from the identical chemical structures of the capsular polysaccharides, lipopolysaccharide, and antigenic O-polysaccharide moieties responsible for the serological specificities of these 2 serovars, as recently shown. 15 Hence, it can be concluded that some of the strains previously identified as biovar 2 serovar 7 on the basis of tests with an antiserum against biovar 1 serovar 7 may actually be serovar 13. In the current study, serovar 7 isolates and the N-273 strain were not tested with a serovar 13–specific antiserum, because no such antiserum was commercially available. However, a previous study reported the occurrence of A. pleuropneumoniae biovar 2 isolates that were originally assigned to serovar 7 in tests with antisera against serovars 1–12 of biovar 1 in the CoA test (as reported herein), but were subsequently reassigned to serovar 13 following precipitation-based tests (i.e., ring test and diffusion test) with type-specific antisera against serovar 13 (Maldonado J, Martínez E, Blanco M, et al.: 2006, Typing of NAD-independent Actinobacillus pleuropneummoniae by PCR and RFLP analysis. In: Proceedings of the 19th International Pig Veterinary Society Congress, vol. 2, p. 238. Copenhagen, Denmark).
Several molecular methods for typing A. pleuropneumoniae have been proposed, but most have been tested only with biovar 1 isolates. The PCR apx typing system described previously 9 made it possible to classify the 15 serovars into 6 groups. When this method was applied to the 112 biovar 2 isolates in the current study, only 2 types were found. The D value was 0.28, and this method was therefore less effective compared with CoA typing and tbpB gene PCR–based RFLP typing.
The tbpB gene, which encodes a 60-kDa protein subunit of the transferrin receptor, is highly divergent in the family Pasteurellaceae. It has therefore been proposed as an alternative tool for typing A. pleuropneumoniae biovar 1 isolates. 7 The method successfully differentiated the reference biovar 1 serovar 7 strain and the reference biovar 2 serovar 13 reference strain (N273; Table 2). Strain N273 and 58 field biovar 2 serovar 7 isolates clustered in PCR-RFLP pattern A, together with the reference strains for biovar 1 serovars 5 and 10 (Table 2). The remaining 3 biovar 2 serovar 7 isolates clustered in PCR-RFLP pattern B, together with biovar 1 serovars 2–4, 7, 9, 11, and 15 reference strains. Taken together, these results support the hypothesis that the A. pleuropneumoniae biovar 2 field isolates tested in the current study and sharing a PCR-RFLP pattern with strain N273 are actually serovar 13, whereas those sharing a PCR-RFLP pattern with serovar 7 from biovar 1 may actually be serovar 7. The use of cross-adsorbed serovar-specific antisera would be necessary to confirm this.
The CoA test assigned 9 biovar 2 field strains from Italy and Spain to serovar 2, and 13 to serovar 4. All but 1 of these strains contained the genes of the apxII operon only. However, biovar 1 strains of serovars 2 and 4 contained the genes of both the apxII and apxIII operons. A previous study reported similar results, with different gene patterns for Swiss and Italian strains of biovar 2 serovars 2 and 4. 1 Hence, the serovar 2 and 4 strains of biovar 2 circulating in Europe seem to have diverged from biovar 1 strains in terms of the organization of apx genes. The tbpB gene PCR–based RFLP assay further divided biovar 2 serovar 4 field strains and nontypeable strains into 2 clusters (Table 1), suggesting that this test may be a useful complementary tool for typing these isolates.
Seventeen biovar 2 field strains gave identical results in all 3 typing assays. The strains reacted strongly with antiserum against serovar 9 in the CoA test, had the apx gene pattern typical of serovar 9 in biovar 1, and clustered in PCR-RFLP pattern C. However, despite this apparent homogeneity, the reference strain for serovar 9 in biovar 1 clustered with a different PCR-RFLP pattern (pattern B), indicating some degree of genetic heterogeneity between the serovar 9 strains of both biovars.
In conclusion, the present study shows that the CoA method reveals more diversity than the other methods tested for the typing of A. pleuropneumoniae biovar 2 isolates, although a significant number of isolates were not typeable. There is evidence that some biovar 2 strains that are designated as serovar 7 may actually be serovar 13. Typing based on the apx profile was the least discriminatory method, recognizing only 2 patterns in the sample of 112 isolates. Furthermore, most (94) of the isolates had an apxII profile. Nevertheless, analysis of the apx gene pattern provides information for assessment of the virulence of biovar 2 field isolates. Finally, typing based on tbpB gene sequence diversity assigned all biovar 2 isolates to a defined RFLP pattern. The tbpB gene PCR–based AvaII RFLP assay is therefore a useful method that may be considered complementary to conventional serotyping. Furthermore, this method can be used to discriminate between strains with identical apx toxin gene arrangements.
Footnotes
Acknowledgements
The authors would like to thank Patrick Blackall (University of Queensland, St Lucia, Australia) for constructive criticism and advice, and Dr. Giuseppe Barigazzi (IZSLER, Parma, Italy) for providing isolates.
a.
Adiavet® APP multiplex PCR kit, Adiagene, Saint-Brieuc, France.
b.
Biovac, Angers, France.
c.
QIAamp® DNA mini kit, Qiagen SA, Courtaboeuf cedex, France.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors received no financial support for the research, authorship, and/or publication of this article.
