Abstract
A 5-year-old Australian stock horse in Monto, Queensland, Australia, developed neurological signs and was euthanized after a 6-day course of illness. Histological examination of the brain and spinal cord revealed moderate to severe subacute, nonsuppurative encephalomyelitis. Sections of spinal cord stained positively in immunohistochemistry with a flavivirus-specific monoclonal antibody. Reverse transcription polymerase chain reaction assay targeting the envelope gene of flavivirus yielded positive results from brain, spinal cord, cerebrospinal fluid, and facial nerve. A flavivirus was isolated from the cerebrum and spinal cord. Nucleotide sequences obtained from amplicons from both tissues and virus isolated in cell culture were compared with those in GenBank and had 96–98% identity with Murray Valley encephalitis virus. The partial envelope gene sequence of the viral isolate clustered into genotype 1 and was most closely related to a previous Queensland isolate.
Keywords
Murray Valley encephalitis virus (MVEV; family Flaviviridae, genus Flavivirus) is an important human pathogen. The principal vector is the freshwater mosquito Culex annulirostris. 16 The virus is considered to be endemic in northern Australia and is periodically reactivated or reintroduced into southern and eastern Australia, but the epidemiology is complex, involving the interplay of vertebrate host, vector, and environmental factors. Although infections occur in a variety of vertebrate hosts, amplification is thought to occur principally in wild birds (waders), especially the Nankeen night heron (Nycticorax caledonicus). 16
Horses are known to be susceptible to a number of mosquito-borne flaviviral encephalitides, including infections by West Nile virus (WNV) and Japanese encephalitis virus. 17 Kunjin virus, a subtype of WNV, 9 has been implicated as the cause of a single case of naturally occurring encephalomyelitis in a horse from the state of Victoria in Australia. 1 MVEV is known to infect horses, based on serological evidence, 7 and there is considerable anecdotal evidence to suggest the virus can cause neurological disease. 22
Experimental infections of horses with MVEV have had varied outcomes, depending on the dosage and route of inoculation. Intracerebral challenge with high doses of virus has produced encephalitis in horses.7,18 Small doses of peripherally inoculated MVEV have either failed to produce infections 7 or have resulted in short-lived viremia and mild, transient clinical signs. 13 The current report describes a case of severe, naturally occurring encephalomyelitis in a horse, attributed to MVEV.
During April 2008, a 5-year-old Australian stock horse mare from Monto, Queensland, Australia (24°52′S, 151°07′E) developed neurological signs. When examined by the veterinarian on day 1, the horse was in a mildly stuporous state but could be roused by stimulation. Mild ataxia and forelimb proprioceptive deficits were noted. Appetite was depressed, but vital signs were normal. Treatment with parenteral a and oral b nonsteroidal anti-inflammatories was instigated. By day 3, there was a degree of facial paralysis, affecting the upper and lower lips but not the eyelids. By day 6, the horse was laterally recumbent and unable to lift its head. At this stage, the horse was euthanized with a barbiturate c overdose, and a field necropsy was conducted. The only abnormality noted at necropsy was brain swelling, which decreased after the dura mater was opened.
Tissues, including brain and cervical spinal cord, were fixed in 10% neutral buffered formalin, and then paraffin embedded, sectioned at 5 µm, and stained with hematoxylin and eosin, using standard techniques. Histology revealed moderate to severe subacute, nonsuppurative encephalomyelitis. Changes were more severe in the hippocampus, midbrain, medulla, and cervical spinal cord than in the cerebrum. The minimum change at all levels of the brain was perivascular cuffs in both gray and white matter. Cuffs were of variable thickness and consisted of a mixture of lymphocytes and histiocytes (Fig. 1). In the spinal cord, severe inflammation was largely restricted to the gray matter (Fig. 2). Diffuse and focal gliosis were present throughout the brain and spinal cord. In the more severely affected areas of midbrain and medulla, swollen, chromatolytic, and necrotic neurons were present, with glial satellitosis (Fig. 3). There were occasional small foci of hemorrhage and malacia in the cerebral white matter, medulla, and spinal cord (Fig. 2). There was also mild nonsuppurative meningitis. No abnormalities were detected in other tissues.
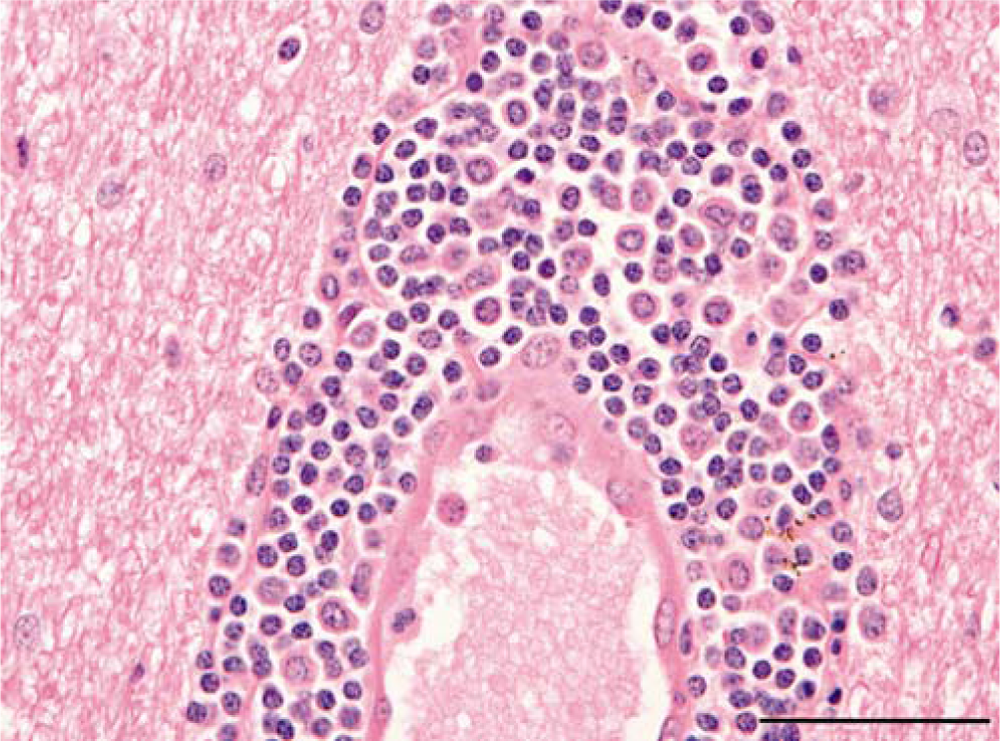
Five-year-old Australian stock horse; cerebral white matter. Dense perivascular cuff of lymphocytes and histiocytes. Hematoxylin and eosin. Bar = 50 µm.
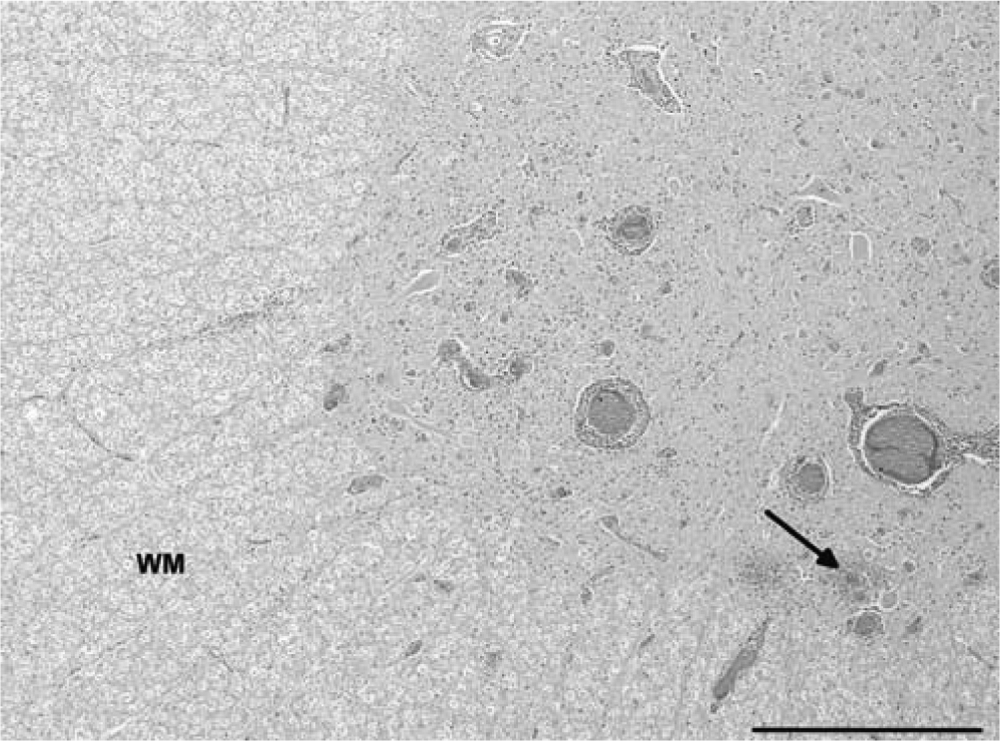
Five-year-old Australian stock horse; cervical spinal cord. Perivascular cuffing, diffuse and focal gliosis, and focal hemorrhage (arrow), largely confined to the gray matter. White matter (WM) is relatively unaffected. Hematoxylin and eosin. Bar = 500 µm.
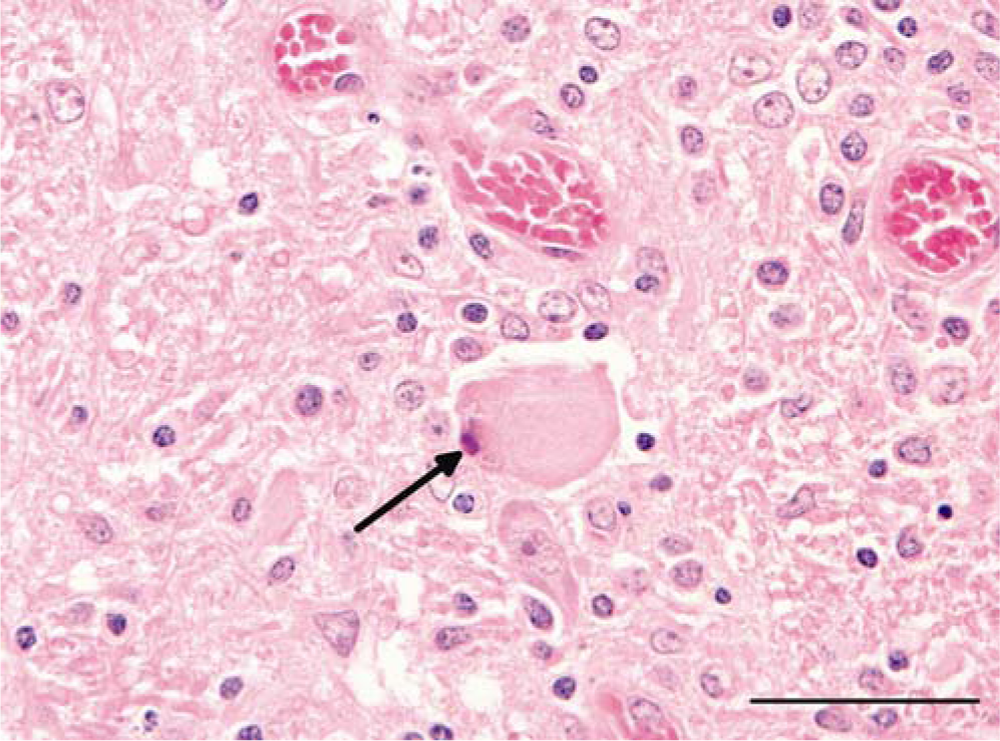
Five-year-old Australian stock horse; medulla oblongata. Satellitosis of a chromatolytic neuron with pyknotic nucleus (arrow); diffuse gliosis. Hematoxylin and eosin. Bar = 50 µm.
Sections of cerebrum, brain stem, cerebellum, and spinal cord were stained with a mouse monoclonal antibody d directed against a conserved epitope of the nonstructural (NS) 1 protein of MVEV, using methods described elsewhere. 2 Granular and diffuse antigen staining was detected in a small number of degenerate neurons, neuronal processes, and the neuropil of spinal cord (Fig. 4). No antigen was detected in cerebrum, cerebellum, or brain stem. Most lesions did not contain antigen staining.
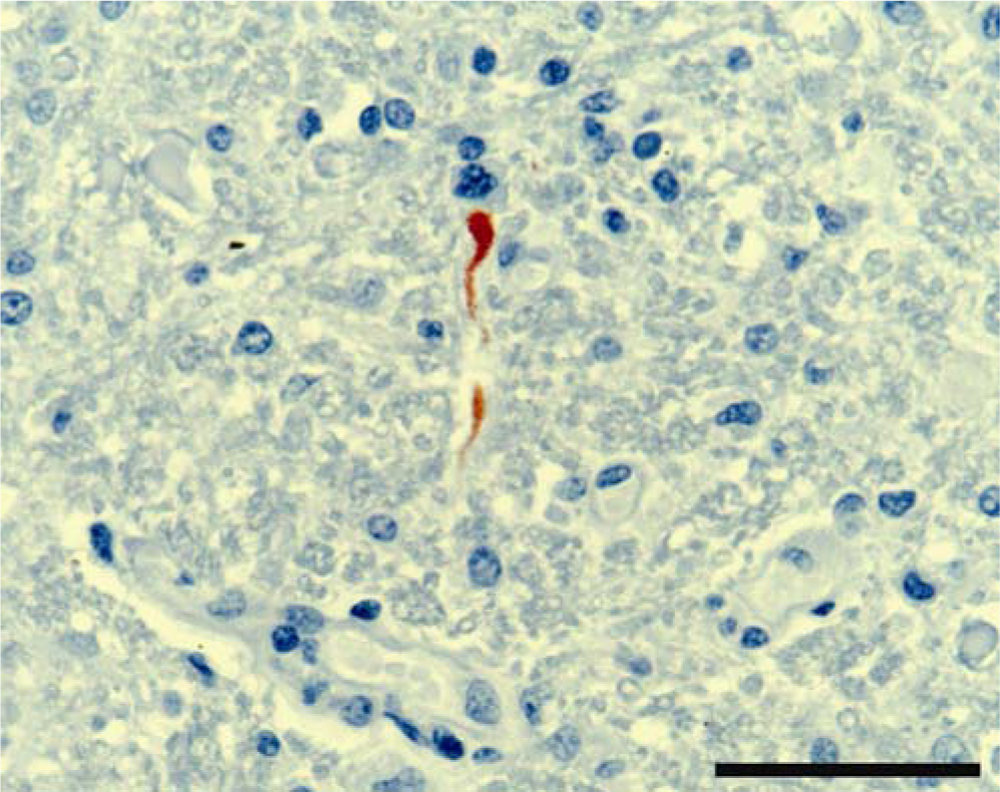
Five-year-old Australian stock horse. Cervical spinal cord, showing viral antigen labeling of a degenerate neuronal cell body and process. Peroxidase immunohistochemistry using 4G4 antibody, counterstained with hematoxylin. Bar = 100 µm.
Samples of blood, cerebrospinal fluid, brain, spinal cord, and facial nerve were tested by reverse transcription nested polymerase chain reaction assays (RT-nPCRs) for a range of arboviruses, including alphaviruses, 21 flaviviruses, 14 and orbiviruses. The latter used an in-house, developmental method still undergoing validation in equids. For flaviviruses, primers FU1PM, cFD3PM, FU2, and cFD4PM of a published method 14 were used. Primers FU2 and cFD4PM were modified to include redundancies and to enable reaction with a wider range of flaviviruses than the primers described in the original method. The primer sequences used were as follows:
FU1PM (TACAACATGATGGGVAARAGWGARAA),
cFD3PM (ARCATGTCTTCYGTBGTCATCCA),
FU2 (modified) (GCTGATGACACMGCYGGMTGG GAYAC), and
cFD4PM (modified) (AYNACRCARTCRTCYCCRCT).
All samples were negative for alphaviruses and orbiviruses. Positive results for flavivirus were obtained from all samples. Brain and spinal cord samples yielded much brighter amplicon bands than samples of cerebrospinal fluid or nerve, and the reaction from the blood sample was very weak. The flavivirus-positive samples were further tested with an in-house, developmental RT-nPCR specific for Kunjin virus; all samples tested negative.
Three samples of cerebrum, 1 sample of cervical spinal cord, and 1 sample of facial nerve were inoculated into yellow fever mosquito Aedes aegypti (AA) C6/36 cells incubated at 28°C for 2 weeks. This was followed by subculture into BSR cells, a clone of baby hamster kidney (BHK)-21 cells, for 1 week at 34°C with supplementation of 5% CO2. The AA and BSR cell lines were maintained in a minimum essential medium supplemented with 10% fetal bovine serum e and the following antimicrobials: penicillin (100 IU/ml), streptomycin (100 µg/ml), and amphotericin B (2 ng/ml). Virus was isolated from all samples except facial nerve. Cells showed cytopathic effect characterized by severe cytoplasmic vacuolation, stringing, rounding-up, and detachment. Supernatants from positive cell culture samples tested PCR-positive for flavivirus.
The RT-PCR amplicons (415 bp) were purified f and used to template dye termination DNA sequencing reactions using a dideoxynucleotide sequencing kit g and the nested flavivirus PCR primers (FU2 [modified] and cFD4PM [modified]). Amplicons were sequenced in both the sense and antisense directions. The reactions were resolved by the Griffith University DNA Sequencing Facility (Nathan, Queensland, Australia), and the resulting chromatograms were proofread and aligned using sequencing software. h The flavivirus group amplicon from virus isolated in cell culture was determined to have an identical nucleotide sequence to those obtained directly from tissue specimens. The consensus nucleotide sequence has been recorded as GenBank accession no. JN206679. This sequence was compared to those in GenBank using the search engine BLAST (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). The highest identity (96%) was with the NS5 gene of MVEV AF161266. BLAST in GenBank of the translated sequence resulted in a 98% similarity with the NS5 protein of MVEV AAC58777 and AAF05296. No other matches were indicated with the nucleotide search. Matches with lower identities (<90%) were noted with other flaviviruses such as Alfuy virus, Japanese encephalitis virus, Usuto virus, WNV, and Kunjin virus.
Further RT-PCR testing of virus culture extract was performed as previously described 11 to produce a 462-bp amplicon of the envelope gene. The amplicon was sequenced (GenBank accession no. JN119766) and aligned with analogous gene regions of reference sequences using ClustalW, as implemented in MEGA 4. 24 Subsequent phylogenetic analysis indicated that the equine MVEV isolate belongs to genotype 1 (Fig. 5), with highest levels of nucleotide sequence identity to the Queensland strain 857-2002, isolated from Burketown in 2002. 12 Close genetic relationships also were inferred with other Queensland and Western Australian strains isolated between 1989 and 2002 (98.3–99.6% nucleotide sequence identity). The results suggest that this strain might have been introduced into southeast Queensland from an endemic focus in either northern Queensland or northwestern Australia.
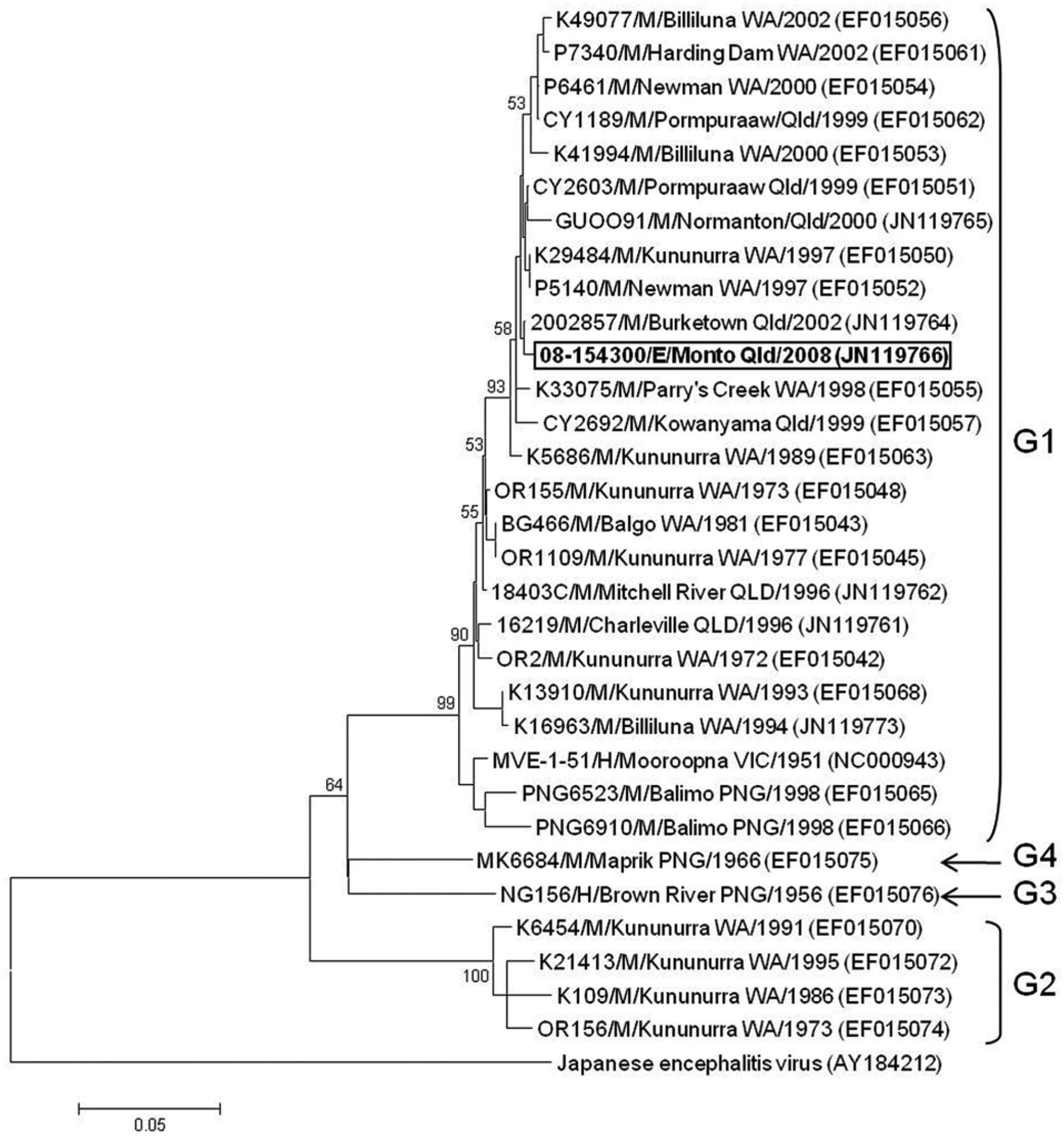
Phylogenetic relationships of the equine isolate of Murray Valley encephalitis virus (MVEV; highlighted) with reference MVEV strains. The tree was constructed from a 462-nucleotide region of the envelope gene using the neighbor-joining method and the maximum composite likelihood model of evolution (MEGA software version 4.0). Japanese encephalitis virus is shown as the outgroup. Percentage bootstrap values from 1,000 replicates are indicated, with a cutoff value of 50%. The scale bar represents 0.05 nucleotide substitutions per site. Details of MVEV strains are shown as strain name/species of origin (H = human; M = mosquito; E = equine)/place and state of origin/year of isolation. GenBank accession numbers are bracketed, and genotype designation is indicated. NSW = New South Wales; PNG = Papua New Guinea; QLD = Queensland; VIC = Victoria; WA = Western Australia.
Outside of the endemic area, there is a strong association between outbreaks of Murray Valley encephalitis (MVE) and certain weather patterns, including excessive rainfall in eastern watersheds. 20 Monto, in the north Burnett region of southeast Queensland, is not within the endemic area. Although higher-than-average rainfall was recorded in Monto during the summer months (January and February 2008) preceding this event, the factors responsible for the current equine case remain unknown. No human cases of MVE, and a single human case of Kunjin virus infection, were recorded from Queensland during 2008 (Australian Government, Department of Health and Ageing: 2005, National Notifiable Diseases Surveillance System. Available at: http://www.health.gov.au/internet/main/Publishing.nsf/Content/cda-surveil-nndss-nndssintro.htm. Accessed May, 25, 2011).
The diagnosis of encephalomyelitis due to MVEV in the current case was based on a combination of consistent histopathology, demonstration of flavivirus antigen within neurons and neuropil, virus isolation, positive RT-PCR, and nucleotide sequencing. The nonsuppurative encephalomyelitis in the horse in the present study has the histological hallmarks of a viral infection of the central nervous system: neuronal degeneration, reactivity of the glia, and perivascular cuffing with lymphocytes and histiocytes. 17 Although there are guidelines for differentiating some of the viral encephalitides of horses based on morphology,3,4 there are no published descriptions of the neuropathology of MVEV infections in horses. Moderate to severe nonsuppurative encephalomyelitis and meningoencephalitis were reported in 2 horses that died in southern Australia in 1974 during an epidemic of human MVE, 7 but infection with MVEV could not be confirmed in either horse. Horses with naturally occurring WNV infection have nonsuppurative polioencephalomyelitis, with the most severe lesions located in the brainstem and spinal cord. 4 In the current case of MVE, the most severe lesions were also located within the brainstem and spinal cord, but preferential involvement of gray matter was evident only in the latter. Other features common to the 2 infections include petechial and ring haemorrhages 4 and a relative scarcity of flavivirus antigen detectable by immunohistochemistry. 3 In the current case, despite widespread distribution of severe lesions, antigen was confined to 1 section of the spinal cord, and it was sparse within this location. This would indicate that the viral antigen was probably largely cleared or blocked by local central nervous system antibody at the time of death.
The differential diagnosis of viral encephalitides in Australian horses has, until recently, been a short list. Many of the recognized viral equine encephalitides are exotic to Australia; such encephalitides include the alphaviruses Eastern equine encephalitis virus, Western equine encephalitis virus, and Venezuelan equine encephalitis virus; Rabies virus; Equine encephalosis virus; Borna disease virus; and other tick-borne encephalitides such as Louping ill virus. 8 For Australian horses, there are infrequent records of encephalomyelitis attributable to Equid herpesvirus 1, 23 Hendra virus, 6 and Kunjin virus. 1 Japanese encephalitis virus has caused subclinical infections in horses in Torres Strait (north of the Australian mainland), 10 is spreading into previously nonendemic areas, 5 and should be considered in future outbreaks of equine neurological disease.
At the time the horse in the current case was diagnosed with encephalitis, Hendra virus infection was considered to be a cause of predominantly fulminating respiratory disease in horses. 19 In June 2008, the 10th recognized outbreak of Hendra virus infection manifested primarily as neurological disease. 6 The serious zoonotic risk of Hendra virus infection now dictates a much more cautious approach to postmortem sampling in horses with neurological disease (Queensland Government: 2011, Guidelines for veterinarians handling potential Hendra virus infection in horses, version 4.1. Available at: http://www.dpi.qld.gov.au/4790_13371.htm. Accessed May 26, 2011). This zoonotic risk complicates the future diagnosis of flaviviral, and indeed any infectious encephalomyelitis, in Queensland horses. There is heightened awareness of the potential for flaviviruses to spread, emerge, and appear in new geographic locations, 15 and the current confirmed equine case of MVE could likely be the forerunner of further such cases.
Footnotes
Acknowledgements
The authors are grateful to the staff of Biosecurity Sciences Laboratory and of Tropical and Aquatic Animal Health Laboratory for help in processing samples from this case. Immunohistochemistry tests were performed by the staff of the Histology Lab, CSIRO Australian Animal Health Laboratory. The authors thank Kalpana Agnihotri, Ibrahim Diallo, Brad Pease, and 2 anonymous reviewers for useful comments on the manuscript.
a.
Flunixil (flunixin meglumine, 50 mg/ml), Troy Laboratories (Australia) Pty Ltd., Glendenning, New South Wales, Australia.
b.
Equibutazone (phenylbutazone, 1 g/sachet), Virbac (Australia) Pty Ltd., Peakhurst, New South Wales, Australia.
c.
Lethabarb (pentobarbitone sodium, 325 mg/ml), Virbac (Australia) Pty Ltd., Milperra, New South Wales, Australia.
d.
4G4, kindly donated by Associate Professor Roy Hall, University of Queensland, St. Lucia, Queensland, Australia.
e.
Sigma-Aldrich, St. Louis, MO.
f.
QIAquick kit, Qiagen Pty Ltd., Doncaster, Victoria, Australia.
g.
BigDye v3.1, Applied Biosystems, Foster City, CA.
h.
Sequencher v4.8 software, Gene Codes, Ann Arbor, MI.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
