Abstract
A simple and portable assay for detection of Streptococcus equi subspecies equi has been developed based on amplification of S. equi–specific sequence using a thermophilic helicase-dependent reaction followed by visual detection of the amplicon in a disposable lateral flow cassette. An experimental kit (IsoAmp™ SE) was evaluated. Analytical sensitivity was 50 copies of S. equi genomic DNA per reaction. The IsoAmp SE assay had 100% specificity when applied to nasal swabs and washes. The assay was more sensitive than culture but less sensitive than nested polymerase chain reaction (PCR). The test requires neither expensive equipment nor extensive training of personnel, provides a practical alternative to culture or PCR assays for detection of S. equi in clinical samples, and expedites identification of atypical colonies of S. equi and Streptococcus zooepidemicus in the laboratory.
Introduction
Strangles caused by Streptococcus equi subspecies equi is the most prevalent contagious bacterial disease of horses in the world. Pyrexia, followed by nasal discharge and abscessation of the lymph nodes of head and neck, are characteristic symptoms.20,22 In some horses, abscesses are formed in body organs distant from the head and neck. 16 Susceptible horses usually acquire infection through direct horse-to-horse contact and indirectly via water sources or feed contaminated by nasal discharges. Shedding of S. equi usually begins after a latent period of 4–16 days and continues for 3–6 weeks after the acute phase. Some animals harbor S. equi in the guttural pouch for months after clinical recovery and may become long-term carriers that intermittently shed the organism.8,14,15 There are over 9.5 million horses in the United States, some of which are frequently moved long distances between racetracks and other competitive events and for breeding, sales, and training. Thus, there is potential for transmission of infectious agents including S. equi. Identification of horses persistently or intermittently shedding the infectious agent is crucially important to control strangles.18,28
Prevention and control of strangles is highly dependent on knowledge of the infection status of individual horses. Culture of nasal swabs and washes, or pus aspirated from abscesses, which is considered the “gold standard” in detection of S. equi, may take several days, and requires significant expertise since specimens often contain other beta-hemolytic streptococci. Moreover, identification of atypical S. equi requires additional biochemical tests. 20 Polymerase chain reaction (PCR) assay based on seM, the gene for the antiphagocytic M protein of S. equi, 24 is more rapid, sensitive, and specific than culture,11,14,23 but is technically complex, requires costly instrumentation, and reagents are not widely available. Also, shipment of samples to qualified laboratories delays decisions concerning movement and transportation of horses and increases the cost of testing.
Development of a rapid, specific, and affordable assay for a wider range of users may substantially expedite identification of S. equi in specimens from horses and improve control of the disease. Such an assay could be based on thermophilic helicase-dependent DNA amplification (tHDA). As for PCR, amplification of target sequence in tHDA is achieved by numerous cycles of DNA denaturation, primer annealing, and primers extension. However, tHDA uses heat-stable helicase to unwind complementary DNA strands instead of heat, allowing isothermal DNA amplification without a thermocycler. 27 Further simplification of the assay could be attained by using a disposable lateral flow cassette for detection of the amplicon.12,27 Such a platform has been recently developed and commercialized a in a kit format and utilized for identification of toxigenic Clostridium difficile in stool samples, 7 Helicobacter pylori, 17 Vibrio cholera, Bacillus anthracis, 26 and Human immunodeficiency virus-1. 21 The assay takes less than 2 hr and can be performed in a minimally equipped laboratory or office setting.
In the current study, the development and evaluation of a tHDA assay for detection of S. equi in specimens from horses is described. The assay is based on asymmetrical amplification of the 5′ end of Se18.9 specific to S. equi 25 followed by hybridization of the amplicon to the labeled probe and detection of the hybridization product in a disposable cassette.
Material and methods
Bacterial culture, DNA isolation, and sample preparation
Twenty isolates of S. equi and 35 isolates of Streptococcus zooepidemicus isolated over the last decade (2000–2010) in different geographical locations were grown at 37°C in Todd–Hewitt broth (THB) supplemented with 0.2% yeast extract or THB agar. Streptococcal DNA were isolated from 5–10 ml of stationary-phase cultures. Following centrifugation at 10,000 × g for 3 min, bacterial pellets were resuspended in 200 µl of phosphate buffered saline (PBS) supplemented with 200 units of mutanolysin b and incubated for 1 hr at 37°C. The DNA was isolated using a commercial kit c from bacterial lysates following the manufacturer’s instructions. Concentrations of DNA were determined using a spectrophotometer. d
Development of S. equi–specific thermophilic helicase-dependent DNA amplification
Sequence encoding the N-terminal region of mature Se18.9 25 of S. equi with least homology in S. zooepidemicus was used to design primers and probe using the Primer3 program. 19 Program parameters were set to select primers with melting temperature, GC, and size of amplicon as 65–71°C, 42–47%, and 80–120 base pairs (bp), respectively. The reverse primer, 5R (Table 1), was labeled with biotin so that the resulting amplicon would bind to streptavidin-conjugated color particles for visualization (Fig. 1). The tHDA probe (Table 1) was labeled with fluorescein isothiocyanate (FITC) as a tag to anchor the colored amplicon-probe hybrid to FITC-specific antibody on the T-line of the detection strip in the disposable lateral flow cassette a (Fig. 1). The primers and probes used in the current study were proprietary. e An IsoAmp™ SE experimental kit for detection of S. equi was prepared by complementation of a commercial tHDA kit a with primers 15F, 5R, and tHDA probe (Table 1). An excess of 5R primer was added to generate single-stranded DNA to bind to the hybridization probe in the final reaction product.
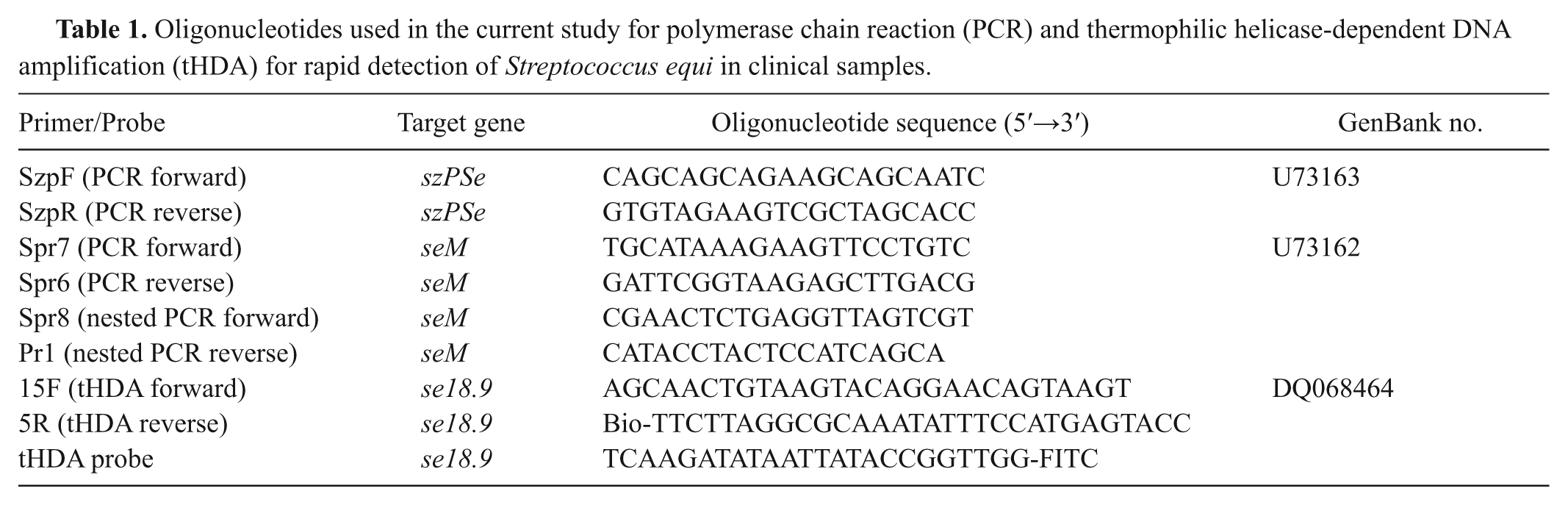
Oligonucleotides used in the current study for polymerase chain reaction (PCR) and thermophilic helicase-dependent DNA amplification (tHDA) for rapid detection of Streptococcus equi in clinical samples.
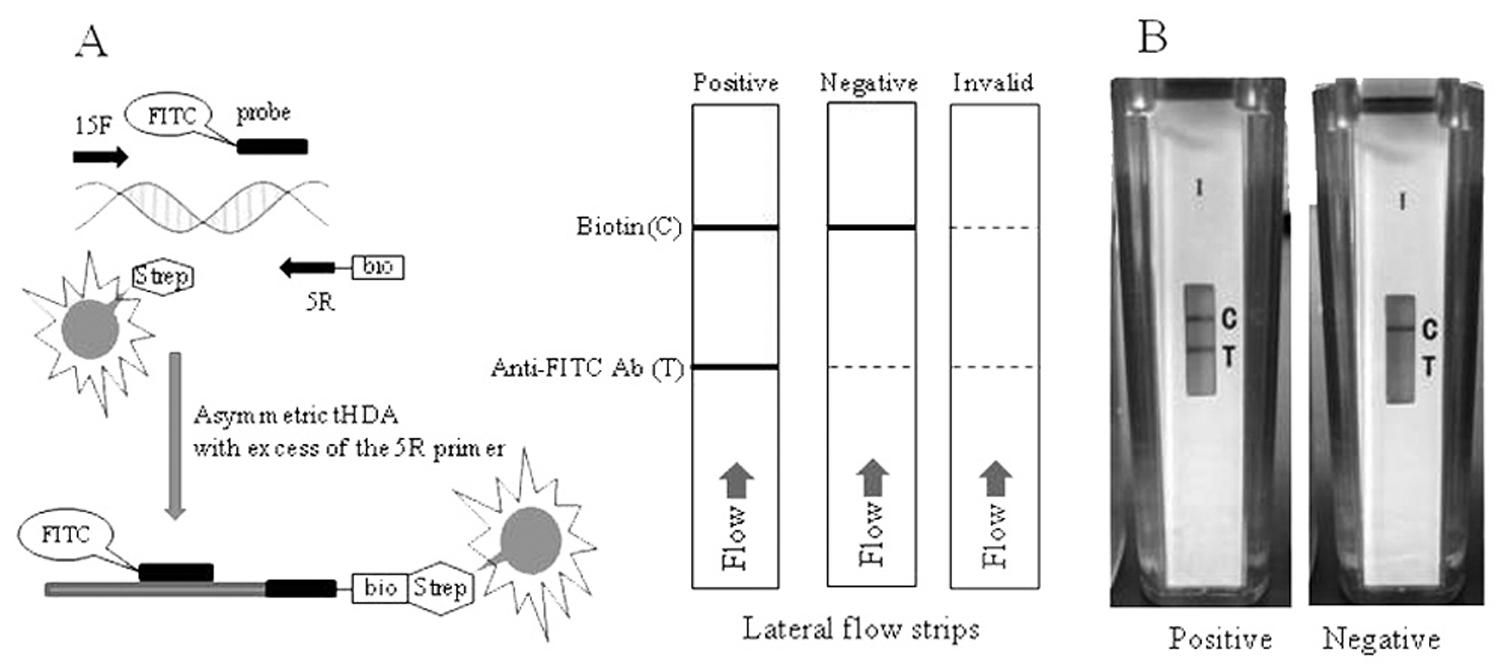
Schematic diagram of the thermophilic helicase-dependent amplification (tHDA) assay using the IsoAmp™ SE kit
a
and disposable lateral flow cassettes.
a
Analytical sensitivity of the IsoAmp SE kit
Ten, 100 copies of genomic DNA of 3 isolates of S. equi were used in triplicate to determine the analytical sensitivity of the kit. Twenty µl of streptococcal genomic DNA were mixed in a 0.2-ml tube with 16.5 µl of buffer mix, a 3.5 ul of enzyme mix, a and 10 µl of water. Following 90-min incubation at 65°C, the reaction tubes containing products of reaction were transferred into disposable cassettes for detection of amplicon by lateral flow capture assay (Fig. 1).
Analytical specificity of the IsoAmp SE kit
The analytical specificity of the assay was tested using 50 copies of genomic DNA from 20 S. equi isolates and 103 and 104 copies of genomic DNA from 35 isolates of S. zooepidemicus per reaction. The tHDA assay and visualization of results were performed as described above.
Evaluation of the IsoAmp SE kit for detection of S. equi in clinical specimens
Samples for tHDA and PCR were prepared from 38 clinical specimens as follows. One ml of nasal wash or 0.5 ml of PBS-containing material released from nasal swabs were centrifuged at 10,000 × g for 3 min. DNA was extracted from the pellets as described above using a commercial kit. c Twenty µl of DNA was used for the tHDA reaction as described above.
Culture and nested PCR were used as reference controls during evaluation of the helicase-dependent DNA amplification experimental kits. For culture, clinical specimens were plated on THB agar supplemented with horse blood and incubated in 5% (v/v) carbon dioxide in air at 37ºC for 48 hr and inspected at 24 and 48 hr for presence of mucoid, hemolytic colonies typical of S. equi. Additional fermentation tests were performed for non-mucoid colonies. Streptococcus equi was identified by its inability to ferment ribose, sorbitol, trehalose, and lactose.
Nested PCR was performed as follows. Aliquots (2 µl) of samples were added to 30 µl of 1× Taq DNA polymerase buffer f containing 1 unit of Taq DNA polymerase, 2 mM of magnesium chloride, 0.2 mM deoxyribonucleotide triphosphate, and 0.2 µM of primers Spr7 and Spr6 (Table 1). The PCR was performed on a thermocycler g using a program involving denaturation at 94°C for 2 min followed by 30 cycles of denaturation at 92°C for 1 min, primer annealing at 57°C for 30 sec, and primer extension at 72°C for 1 min. Following completion of the first round, 2 µl were transferred to a fresh reaction mix with nested primers Spr8 and Pr1 (Table 1), and a second round of PCR was performed using the same conditions. The PCR products were analyzed by ethidium bromide agarose electrophoresis. Amplicons produced by nested PCR of samples negative in tHDA and culture were purified and sequenced to confirm the presence of seM sequence of S. equi. Sequencing was performed by a commercial facility. e
Results of tHDA and culture were scored in comparison with nested PCR as a reference standard. Sensitivity, specificity, and diagnostic accuracy were calculated as ratios of (true positive/true positive + false negative), (true negative/true negative + false positive), and (true negative/true negative + false negative), respectively. True positive, false positive, true negative, and false negative were specimens that were PCR+/Test+, PCR–/Test+, PCR–/Test–, and PCR+/Test–, respectively.
Identification of atypical colonies of S. equi and S. zooepidemicus
Portions of colonies of non-mucoid isolates of S. equi or mucoid isolates of S. zooepidemicus were transferred using toothpicks into 0.5-ml tubes containing 15 µl of water. Three randomly selected colonies of each isolate of S. equi and S. zooepidemicus were used. Following incubation at 100°C for 10 min and centrifugation at 10,000 × g for 2 min, 10- and 5-µl aliquots of supernatants were tested by tHDA and PCR, respectively. Primers SzpF and SzpR universal for S. equi and S. zooepidemicus, and Spr6 and Spr7 specific for S. equi, were used for PCR (Table 1). The primers used in the present study are proprietary. h The tHDA and PCR assays were performed as described above.
Results
Development of S. equi–specific thermophilic helicase-dependent DNA amplification
Primers 15F and 5R (Table 1) were selected, following comparison of 20 primers, as the most specific and providing the highest amplicon yield in tHDA reaction. No homology to amplicon sequence has been detected in genomes of other microorganisms following a search of available data banks. An experimental kit, IsoAmp SE, was manufactured using these primers and probe. Figure 1 illustrates the workflow of the assay including tHDA and lateral flow detection of amplicons. Asymmetric amplification of the target sequence in the presence of an excess of primer 5R results in enrichment of the reaction with single-stranded, biotin-labeled DNA. Such DNA is effectively hybridized at incubation temperature to the probe forming a biotin/FITC-labeled complex. The hybrid moves along the detection strip of the cassette, binds with red latex beads coated with streptavidin, and is captured by anti-FITC antibodies in the test line. Streptavidin-coated beads that have not been bound by amplicon are captured on the biotin control line. When only the control line is detected, the assay is scored as a true negative (Fig. 1).
Analytical sensitivity and specificity of the IsoAmp SE kit
Ten, 100 copies of genomic DNA of 3 isolates of S. equi were used in triplicate to determine the analytical sensitivity of the kit. Results were strongly positive in all experiments with quantities of template equivalent to 50 and 100 copies of S. equi genomic DNA per reaction. Reducing the quantity to 25 copies led to inconsistent results that are probably due to minor variation in concentration of target or capture antibody on the detection strip. No responses were detected for 10 copies of genomic DNA per reaction.
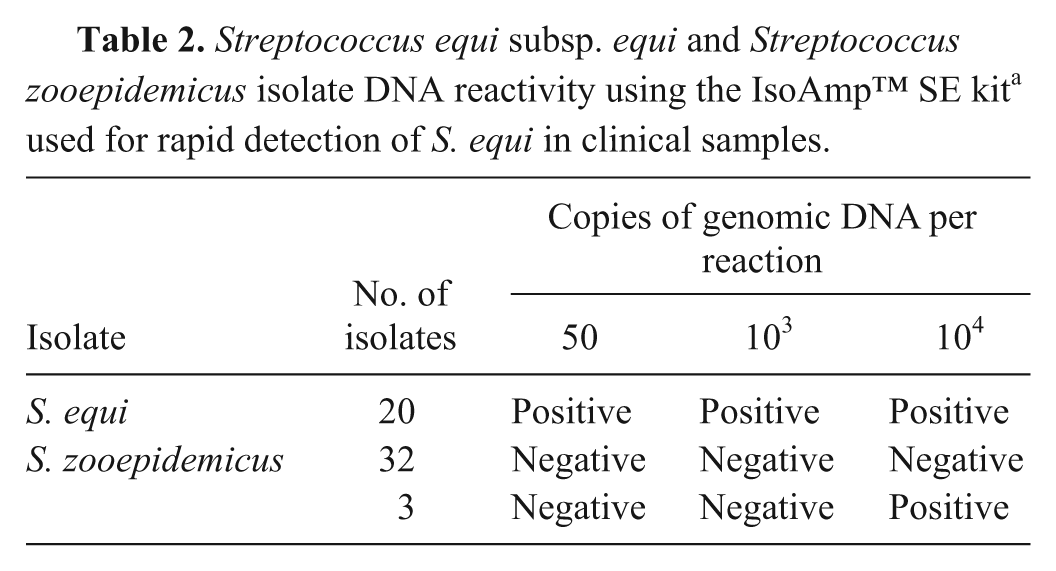
The analytical specificity of the assay was tested using DNA from 20 isolates of S. equi and 35 isolates of S. zooepidemicus. The DNA of S. equi isolates gave positive responses when 50 copies of genomic DNA were used in the assay (Table 2). No reactivity in tHDA was detected for 103 copies of genomic DNA for each of 35 isolates of S. zooepidemicus. However, DNA of 3 isolates of S. zooepidemicus gave positive responses at 104 copies per reaction.
Streptococcus equi subsp. equi and Streptococcus zooepidemicus isolate DNA reactivity using the IsoAmp™ SE kit a used for rapid detection of S. equi in clinical samples.
Evaluation of thermophilic helicase-dependent DNA amplification assay for detection of S. equi in clinical samples
Polymerase chain reaction, tHDA, and culture were compared for detection of S. equi in equine samples. Streptococcus equi DNA was detected by tHDA in 20 out of 38 samples that were also positive by nested PCR. No tHDA-positive samples were identified among PCR-negative samples showing that tHDA is as specific as PCR for detection of S. equi. Nevertheless, detection of S. equi DNA by nested PCR in 3 specimens negative by culture and tHDA revealed a higher sensitivity of the nested PCR. Sequence of amplicons produced by nested PCR of these specimens showed identity to the corresponding part of the seM gene of S. equi. Culture revealed S. equi in 17 out of 20 tHDA-positive specimens, whereas S. equi was not detected in tHDA-negative specimens. Thermophilic helicase-dependent DNA amplification was therefore as specific as culture but more sensitive in identification of S. equi in clinical samples (Table 3). Other Gram-positive flora including S. zooepidemicus contaminated all except 3 of the S. equi–positive and –negative specimens, but this did not interfere with assay specificity.
Comparison of thermophilic helicase-dependent amplification (tHDA) and culture for detection of Streptococcus equi in clinical samples from horses.*
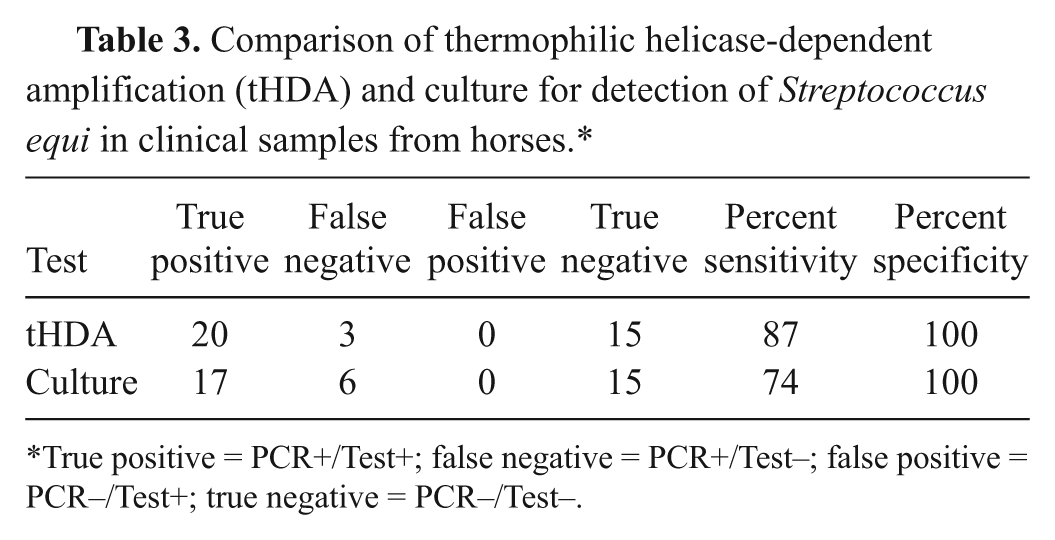

True positive = PCR+/Test+; false negative = PCR+/Test–; false positive = PCR–/Test+; true negative = PCR–/Test–.
Comparison of tHDA and culture using the nested PCR as a standard revealed that the tHDA assay had a sensitivity of 87%, a specificity of 100%, and a diagnostic accuracy of 83%. Culture was as specific as PCR and tHDA for detection of S. equi in specimens from horses but had inferior sensitivity (74%) and a diagnostic accuracy of 71% (Table 3).
Identification of morphologically atypical S. equi and S. zooepidemicus
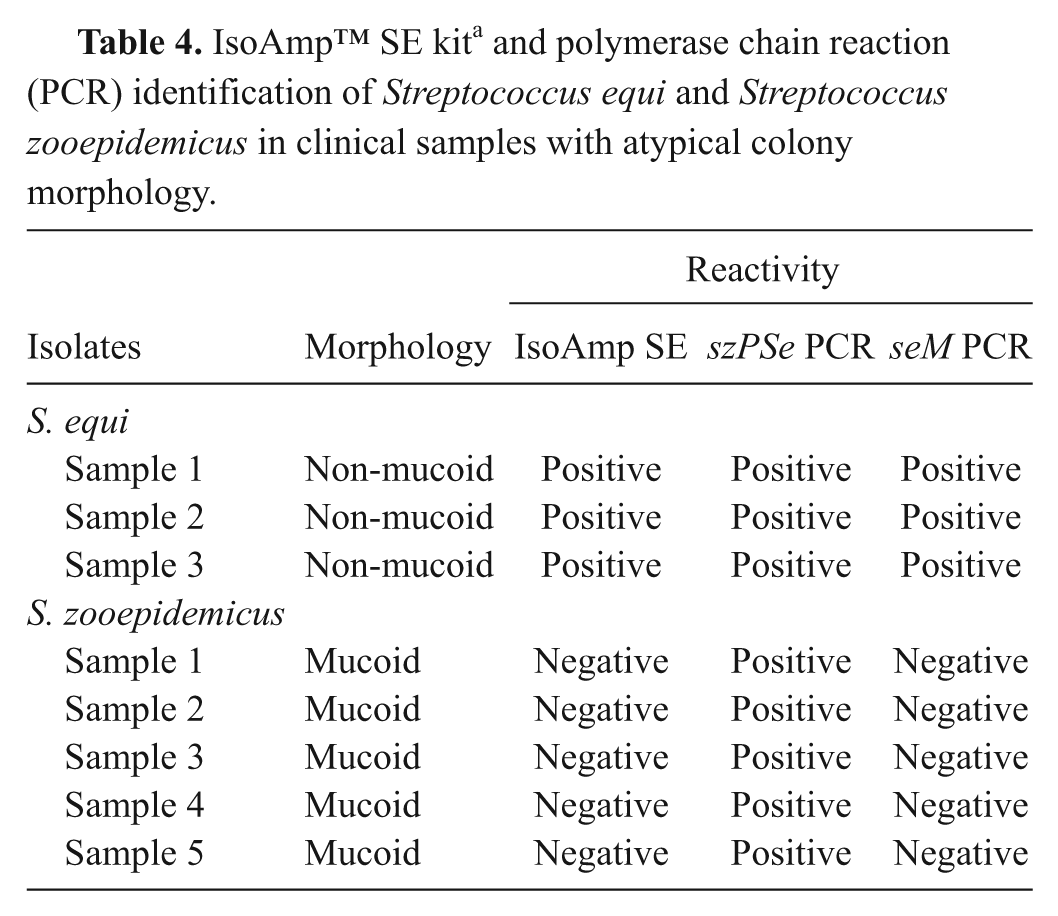
Nucleic acids extracted from a portion of colonies of 3 non-mucoid S. equi and 5 mucoid S. zooepidemicus were analyzed by tHDA and PCR. The presence of streptococcal DNA from colonies was confirmed by positive reactions in PCR with SzpF and SzpR primers specific for both S. equi and S. zooepidemicus (Table 4). Streptococcus equi isolates alone were positive in tHDA as well as in PCR when primers Spr6 and Spr7 specific for seM were used.
IsoAmp™ SE kit a and polymerase chain reaction (PCR) identification of Streptococcus equi and Streptococcus zooepidemicus in clinical samples with atypical colony morphology.
Discussion
In the current study, a tHDA-based assay (IsoAmp SE) for detection of S. equi in specimens from horses was developed and evaluated. The test required neither expensive equipment nor extensive training of personnel and may be used for point-of-care testing. The Se18.9 gene encoding an antiphagocytic virulence factor of S. equi that binds the H control factor of complement is secreted abundantly by S. equi, but not by the closely related S. zooepidemicus, which is commonly found in the nasal cavity of horses, 25 and was used as a target for the tHDA assay. Comparison of sequences of Se18.9 from 8 isolates of S. equi and 10 isolates of S. zooepidemicus showed that its 5′ terminal half is highly conserved among S. equi isolates of different geographical origin, whereas the ortholog in S. zooepidemicus is divergent and shares only limited homology with Se18.9 (data not shown). Primers and a probe selected from this sequence revealed high specificity to S. equi giving positive results in tHDA with all DNA from 20 randomly selected isolates of this microorganism. The DNA from 35 isolates of S. zooepidemicus collected during the last decade from different geographical regions showed negative results in tHDA when 103 copies of genomic DNA per reaction were used. However, additional experiments using 104 copies of genomic DNA per reaction resulted in positive responses for 3 out of 35 isolates of S. zooepidemicus creating the potential for false-positive results in cases of heavy contamination by some S. zooepidemicus isolates (Table 2).
The analytical sensitivity of the tHDA assay was 50 copies of genomic DNA of S. equi per reaction and lies between that for conventional PCR and nested PCR, as previously described for S. equi.3,14 The sensitivity reported for this platform (combination of tHDA and disposable cassette) and that previously developed 9 for identification of Staphylococcus aureus were the same. However, this sensitivity level is not the limit for this platform. For instance, the analytical sensitivity of the assay for detection of C. difficile was 20 copies of genomic DNA per reaction. 7
Comparison of the tHDA test with assays currently used for identification of S. equi reveals that the specificity of tHDA is similar to that of culture and PCR when applied to nasal swabs and washes. Sensitivity and diagnostic accuracy of tHDA were 13% higher than those of culture. Although evaluation of the tHDA applied to culture-negative clinical specimens from horses with different degrees of contamination did not reveal false-negative results (Table 3), a conclusion about the probability of false-positive results will require future trials on additional clinical samples.
A second potential application of the tHDA assay is rapid identification of atypical forms of group C streptococci periodically isolated from equine specimens. Culture is widely used for definitive diagnosis of strangles and should be routinely used to confirm diagnosis based on PCR. According to this technique, samples from horses are inoculated onto horse blood agar, incubated 24–48 hr, and inspected for hemolytic mucoid colonies characteristic of S. equi. The other Lancefield group C streptococci, S. zooepidemicus and Streptococcus dysgalactiae subsp. equisimilis, which frequently present in clinical samples, also form hemolytic colonies, but are usually non-mucoid. Ability to ferment lactose, trehalose, and sorbitol is an additional descriptor.13,20,22 Nevertheless, the correct identification of S. equi and other streptococci could be hampered in the case of isolates with atypical colony morphology and fermentation pattern. 10 Several molecular methods including sequencing of the intergenic region of the 16S-23S RNA genes, 6 multiplex PCR, 2 real-time PCR, 4 multilocus sequence typing, 29 pulsed field gel electrophoresis, 5 and random amplification of polymorphic DNA PCR1,5 have been evaluated for differentiation of S. equi and other streptococci. Although any one of these methods could be used for resolving ambiguous cases, the absence of essential reagents, equipment, and expertise in a diagnostic laboratory exclude practical application of these techniques. The current study demonstrated that simple sample preparation followed by tHDA could differentiate atypical S. equi and S. zooepidemicus in less than 2 hr.
In summary, a specific and sensitive assay for detection of S. equi in equine specimens based on tHDA reaction followed by visualization of the test result in a disposable cassette is described. The ability to perform this assay in a simple laboratory or even point-of-care by personnel with basic training could substantially reduce the time for making decisions concerning movement and/or isolation of horses. The assay could also be used in clinical laboratories to expedite identification of morphologically and biochemically atypical isolates of S. equi and S. zooepidemicus.
Footnotes
a.
IsoAmp®, BioHelix® Express Strip (BESt™) cassette type I, IsoAmp® II Universal tHDA Kit; BioHelix Corp., Beverly, MA.
b.
Sigma-Aldrich, St. Louis, MO.
c.
ZR Genomic DNA II Kit™, Zymo Research Corp., Irvine, CA.
d.
NanoDrop, Thermo Scientific, Wilmington, DE.
e.
Operon Biotechnologies Inc., Huntsville, AL.
f.
Invitrogen Corp., Carlsbad, CA.
g.
Sprint, Thermo Hybaid, Franklin, MA.
h.
Integrated DNA Technologies Inc., Coralville, IA.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The U.S. Department of Agriculture (USDA SBIR 2008-00036) and the Kentucky Science and Engineering Foundation (KSEF-148-502-08-228) supported the current research.
