Abstract
Pseudorabies is caused by Suid herpesvirus 1, a member of the Alphaherpesvirinae subfamily. Although pigs are the natural host of Pseudorabies virus (PRV), the virus has a broad host range and may cause fatal encephalitis in many species. The United States obtained PRV-free status in 2004 after the virus was eradicated from domestic swineherds, but the virus is still present in feral swine populations. The current report describes PRV infection in 3 dogs that were used to hunt feral swine. The dogs developed clinical signs including facial pruritus with facial abrasions, dyspnea, vomiting, diarrhea, ataxia, muscle stiffness, and death. Two were euthanized, and 1 died within approximately 48 hr after onset of clinical signs. The salient histologic changes consisted of neutrophilic trigeminal ganglioneuritis with neuronophagia and equivocal intranuclear inclusion bodies. Pseudorabies virus was isolated from fresh tissues from 2 of the dogs, and immunohistochemistry detected the virus in the third dog. Virus sequencing and phylogeny, based upon available GenBank sequences, revealed that the virus was likely a field strain that was closely related to a cluster of PRV strains previously identified in Illinois. Though eradicated from domestic swine in the United States, PRV is present in populations of feral swine, and should therefore continue to be considered a possible cause of disease in dogs and other domestic animals with compatible clinical history and signs. Continued surveillance is necessary to prevent reintroduction of PRV into domestic swine.
Introduction
Pseudorabies virus (PRV), also known as Aujeszky’s disease, is caused by Suid herpesvirus 1, a member of the Alphaherpesvirinae subfamily. In pigs, the natural host of PRV, the clinical picture is dependent upon age at exposure, immune status, route of infection, and strain virulence. Although PRV infections in neonates and young pigs initially may present as respiratory disease, young pigs rapidly succumb to neurologic disease. In contrast, respiratory disease is the most common manifestation in older pigs, with neurologic disease developing only due to high titered inoculum of virulent viral strains. In addition, infection of pregnant gilts or sows frequently results in resorption, abortion, or birth of mummified fetuses or stillborn neonates. In pigs surviving acute infection, latency develops, primarily in neuronal tissues, but also in lymphoid tissues. Stress may induce recrudescence of latent virus, and the virus may be shed in nasal and oral secretions, semen, and vaginal secretions. The virus may be transmitted by nose-to-nose contact, coitus, artificial insemination, fomites, or transplacentally. Infectious levels of virus remain in the carcasses of dead animals. 31
Pseudorabies virus can affect a wide range of species including dogs, cats, cattle, sheep, goats, foxes, hedgehogs, polecats, opossums, jackals, some nonhuman primates (rhesus macaques, marmosets), chickens, pigeons, geese, ducks, buzzards, sparrow hawks, rabbits, guinea pigs, rats, and mice. 35 Human beings are not known to be susceptible. In species other than swine, PRV typically causes fatal neurologic disease, often associated with localized pruritus. 31
Pseudorabies virus is global in distribution, but it has been eradicated from several countries including Austria, Sweden, Denmark, United Kingdom, Canada, and New Zealand. 31 The virus was declared eradicated from domestic swineherds in the United States in 2004, 39 though it continues to circulate among feral swine. 20 However, the actual prevalence of PRV infection in domestic animals since that time is unknown. There have been unpublished reports of pseudorabies infection in Florida hunting dogs (Mark Cunningham, personal communication, 2008; www.promedmail.org, archive no. 20081118.3637). The objective of the current report is to characterize an outbreak of PRV infection in 3 Oklahoma hunting dogs.
Materials and methods
Tissue sampling
Postmortem examinations were performed at the Oklahoma Animal Disease Diagnostic Laboratory (Stillwater, Oklahoma) on dogs 1 and 3. The head of dog 1 had been submitted to the rabies laboratory and was not available for examination. The carcass of dog 3 had been frozen and thawed. Samples collected at necropsy included heart, lung, liver, kidney, stomach, esophagus, small intestine, and large intestine (dog 1), and trigeminal ganglia, brain, facial skin, heart, lung, liver, kidney, urinary bladder, stomach, small intestine, large intestine, adrenal gland, spleen, pancreas, and hind limb muscles (dog 3). Tissues were immersed in 10% neutral buffered formalin and allowed to fix for 24 hr. Representative sections of tissues were trimmed, embedded in paraffin, sectioned at 4 µm, processed routinely, and stained with hematoxylin and eosin.
Immunohistochemistry
Immunohistochemistry was performed on sections of the trigeminal ganglia from dog 3 at the Veterinary Diagnostic Center (University of Nebraska–Lincoln, Lincoln, Nebraska). Briefly, sections of formalin-fixed paraffin-embedded tissues were cut at 4 µm. Slides were deparaffinized and stained on an automated immunohistochemical stainer. a Primary antibody was a rabbit polyclonal antiserum (RB132) against PRV at a dilution of 1:10,000. 24 Positive and negative controls consisted, respectively, of a slide containing known positive tissue along with slides of test samples using an irrelevant primary antibody. The slides were pretreated with Fixative 1 b at ambient temperature for 8 min followed by 12-min incubation with protease III. b Before application of the primary antibody, a blocking step using antibody diluent b for 24 min was applied. Primary antibody incubation was for 32 min at 37°C. Secondary antibody, alkaline phosphatase, and substrate were proprietary. c Tissues were counterstained with hematoxylin for 8 min and coverslipped with glass coverslips.
Virus isolation and identification
Virus isolation and identification were performed at the Diagnostic Center for Population and Animal Health (Michigan State University, Lansing, Michigan). Homogenates of fresh brain tissue samples from dogs 1 and 2 were prepared, clarified by low speed centrifugation (600 × g; 10 min, 4°C), and filtered through disposable filters with an average pore diameter of 0.45 µm. Aliquots of the filtrates were inoculated onto confluent monolayers of Crandell feline kidney (CRFK) cells grown on chamber slides. d The growth medium consisted of Eagle minimum essential medium (EMEM), supplemented with 10% fetal bovine serum (FBS) and 10 µg/ml of ciprofloxacin. e The inoculum was allowed to adsorb for 1 hr at room temperature. The monolayers were then washed once, re-fed with EMEM containing 2% FBS and 10 µg/ml of ciprofloxacin, and incubated at 37°C in an atmosphere of 5% CO2 in air.
When cytopathic effects (CPE) were observed in the inoculated cells, the medium was removed and the slides were dipped sequentially in phosphate buffered saline (PBS) and distilled water. The cells were then fixed in 30% acetone in PBS for 10 min at room temperature and dried overnight at room temperature. The slides were then stained with 75 µl/well of anti-PRV conjugate f for 30 min at 37°C in a humidified chamber. They were subsequently rinsed in 0.1 M PBS (pH 7.2, fluorescent antibody wash buffer), soaked for 10 min in fluorescent antibody wash buffer, counterstained with 0.05% Evans blue in H2O, and viewed under a microscope equipped with a ultraviolet (UV) light source. g
DNA extraction
DNA from fresh tissue from dogs 1 and 2 and from virus cell culture supernatant was extracted using a commercial kit h according to the manufacturer’s protocol.
PCR
DNA was initially tested for the presence of PRV DNA using a real-time PCR protocol from the University of Illinois (Scherba, personal communication). Samples for PCR were pooled to limit financial costs. Primers that target the glycoprotein D (gD) gene, forward primer (US6-1190F) 5′-ACACCGACGAGCTAAAAGCG-3′ and reverse primer (US6-1252R) 5′-ATCATCATCGACGCCGGTA-3′, were each used at 0.45 µM concentration in a 25 µl final reaction mix made with a commercial kit. h The MGB probe, (US6-1211T) 5′-AGCCCGGTCCGTAGC-3′, was used at 0.2 µM final concentration. Five microliters of extracted DNA was used as template. The assay was run in an automated real-time PCR system i with a denaturation step at 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min.
DNA extracted from the virus isolate was also used as template for the amplification of a portion of the PRV glycoprotein C (gC) and glycoprotein E (gE) genes. A 712-bp fragment of the gC gene was targeted with forward primer (PRV-F1), 5′-TTTCCTGATTCACGCCCACGCT-3′, and reverse primer (PRV-R1), 5′-ACGTGTAGACGCCCTCGGTCT-3′. 20 Cycling conditions for gC amplification were as follows: denaturation at 95°C for 5 min; 8 cycles of 95°C for 50 sec, 67°C for 50 sec, and 72°C for 1 min; 37 cycles of 95°C for 50 sec, 63°C for 50 sec, and 72°C for 1 min; and a final extension at 72°C for 5 min.
For gE PCR, a 493-bp target was amplified using the forward primer (gE-nF), 5′-CCGCGGGCCGTGTTCTTTGT-3′, and reverse primer (gE-nR), 5′-CGTGGCCGTTGTGG GTCAT-3′. 23 Cycling conditions for gE PCR were as follows: denaturation at 95°C for 5 min, followed by 45 cycles of 95°C for 50 sec, 60°C for 40 sec, and 72°C for 50 sec, and final extension of 72°C for 5 min. Polymerase chain reaction for both gC and gE was carried out with a commercial kit, h using 0.5 µM final primer concentrations, 5% dimethyl sulfoxide, and 5 µl of DNA template, in a 50-µl final reaction volume. The PCR products were analyzed by electrophoresis on 2% agarose gels and visualized by UV transillumination.
Partial gC gene sequencing and phylogenetic analysis
Amplicons of the expected size were cut from the gel and purified using a commercial extraction kit. h The PCR products were sent together with both forward and reverse primers to the Research Technology Support Facility of Michigan State University for automated DNA sequencing. BLAST 1 analysis of sequence data was performed to search for significant similarities to sequences in the GenBank database. Sequence assembly and multiple alignments of nucleotide sequences using the ClustalW method 38 were conducted with the Lasergene biocomputing software. j The phylogenetic tree was constructed with the Treecon software package, 41 using the neighbor-joining method 33 and 500 bootstrap analyses.8,11 Sequence alignment data generated with Lasergene were converted to the Treecon format using the ForCon software (Raes J, Van de Peer Y: 1999, ForCon: a software tool for the conversion of sequence alignments. In: EMBnetnews. Available at: http://www.embnet.org/files/shared/EMBnetNews/embnet_news_6_1.pdf).
The following are the GenBank accession numbers and origin of the sequences used for construction of the phylogenetic tree: PRV11243 (AF176492, U.S.), PRV43 (AF176479, U.S.), PRV10501 (AF176490, U.S.), PRV7652 (AF176484, U.S.), PRV10649 (AF176491, U.S.), PRV9164 (AF176489, U.S.), PRV12486 (AF176495, U.S.), PRV12481 (AF176494, U.S.), PRV12271 (AF176493, U.S.), PRV8033 (AF176486, U.S.), PRV4411 (AF176481, U.S.), PRV8044 (AF176487, U.S.), PRV OK2010 (virus isolated in this study, U.S.), Indiana S (D49436, U.S.), F3303 (EU622054, Brazil), IB341-86 (EU622076, Brazil), NIA3 (D49437, Northern Ireland), 25 Becker (M12778, Europe), 13 576-Hun (GQ259114, Hungary), 29 613-GER SN (GQ259118, Germany), Yamagata S-81 (D49435, Japan), 25 614-GER BW (GQ862778, Germany), 2882-ESP (GQ259123, Spain), 536-FRA (GQ259099, France), and FA (AF403051, China). 18 The available GenBank sequences of the isolates from Brazil, 12 Germany, France, and Spain 29 lack the first 35 bases of corresponding sequence to PRV OK2010. The PRV OK2010 gC sequence and GenBank-derived corresponding sequences of PRV strains from the United States, Brazil, Europe, and China were used in the construction of a phylogenetic tree. The PRV OK2010 sequence was deposited in GenBank under the accession number JF767011.
Toxicological analysis
Thin-layer chromatography k drug analysis was performed at the Oklahoma Animal Disease Diagnostic Laboratory on samples of urine from dogs 1 and 3. Samples were tested for diphenhydramine, dimenhydrinate, ketamine, amphetamine, methamphetamine, codeine, morphine, acetaminophen, strychnine, pseudoephedrine, caffeine, diazepam, nicotine, and chlorpromazine. An enzyme-linked immunosorbent assay (ELISA) test card l was performed on urine from dog 1 to test for marijuana, amphetamines, cocaine, barbiturates, benzodiazepines, methamphetamine, morphine, phencyclidine, and tricyclic antidepressants.
Results
History and clinical findings
Over the course of 2 weeks, 3 dogs from the same kennel developed similar clinical signs and were either euthanized or died. The dogs were used for hunting feral hogs in southern Oklahoma. Two dogs escaped from the kennel together and returned a day later. Approximately 8 hr after their return, dog 1 (1.5-year-old female Catahoula leopard dog) developed signs of severe facial pruritus and self-trauma. On presentation to the veterinarian, the dog was depressed, dyspneic, vomiting, and salivating with variable muscular stiffness and marked facial pruritus. A complete blood count showed mild anemia (31.6%, reference [ref.] interval: 37.0–55.0%), leukocytosis (26.4 × 109/l, ref. interval: 6.0–16.9 × 109/l), neutrophilia (22.8 × 109/l, ref. interval: 2.8–10.5 × 109/l), eosinophilia (1.8 × 109/l, ref. interval: 0.5–1.5 × 109/l), hyperproteinemia (10.8 g/dl, ref. interval: 5.4–8.2 g/dl), hyperglobulinemia (8.4 g/dl, ref. interval: 2.3–5.2 g/dl), hypoalbuminemia (2.4 g/dl, ref. interval: 2.5–4.4 g/dl), and hyperglycemia (131 mg/dl, ref. interval: 60–110 mg/dl). Results of parasite screening m (ELISA for Dirofilaria immitis, Ehrlichia canis, Anaplasma phagocytophilum, and Borrelia burgdorferi) were negative. The dog was treated symptomatically with diphenhydramine, n dexamethasone, o cefazolin, p maropitant citrate, q diazepam, r and phenobarbital. s Due to progression of clinical signs and the lack of response to treatment, the dog was euthanized (pentobarbital). o
Dog 2 (2-year-old female Australian Cattle Dog) developed clinical signs at approximately the same time as dog 1. Clinical signs included rapid onset of respiratory distress, salivation, and muscle rigidity. The dog was febrile (40.1°C). A complete blood count revealed leukocytosis (32.7 × 109/l, ref. interval: 6.0–16.9 × 109/l) with granulocytosis (26.4 × 109/l, ref. interval: 3.3–12.0 × 109/l), hyperproteinemia (8.8 g/dl, ref. interval: 5.4–8.2 g/dl), hyperglobulinemia (5.6 g/dl, ref. interval: 2.3–5.2 g/dl), and hyperglycemia (298 mg/dl, ref. interval: 60–110 mg/dl). Results of parasite screening m were negative. Symptomatic treatment was administered as above for dog 1. The dog failed to respond to treatment and was euthanized (pentobarbital) o within 24 hr of the appearance of clinical signs.
As neither dog 1 or 2 was vaccinated against rabies, the heads of both dogs were submitted for rabies testing. Both tested negative for this virus by fluorescent antibody testing at the Oklahoma State Department of Health, Public Health Laboratory (Oklahoma City, Oklahoma). The body of dog 1 was submitted for necropsy.
One week later, dog 3 (2.5-year-old male Catahoula leopard dog) escaped from the kennel and returned the same day with signs of facial pruritus. Over 48 hr, facial pruritus and self-inflicted abrasions became more severe; the dog was increasingly restless and developed bloody diarrhea and vomiting. Treatment included activated charcoal, t butorphanol tartrate, u methylprednisolone, q acepromazine, v and ketamine (unrecorded manufacturer). Dog 3 died approximately 48 hr after the onset of clinical signs and was submitted for necropsy.
Toxicosis was initially suspected, considering the acute onset of disease affecting multiple animals in the same kennel and the short course of illness. Pseudorabies was also considered, given the clinical signs and history of exposure to feral hogs.
Gross and histologic lesions
Postmortem examination of dog 1 revealed extensive subendocardial hemorrhage of the right ventricle. The head was not available for examination. In dog 3, variable portions of the skin over the right side of the face were moist and abraded (Fig. 1). Within the right cardiac ventricle were multiple nematodes consistent with D. immitis. The small intestine contained numerous nematodes consistent with Ancylostoma sp. admixed with red contents, and large intestinal contents were variably red. Mesenteric lymph nodes were prominent.
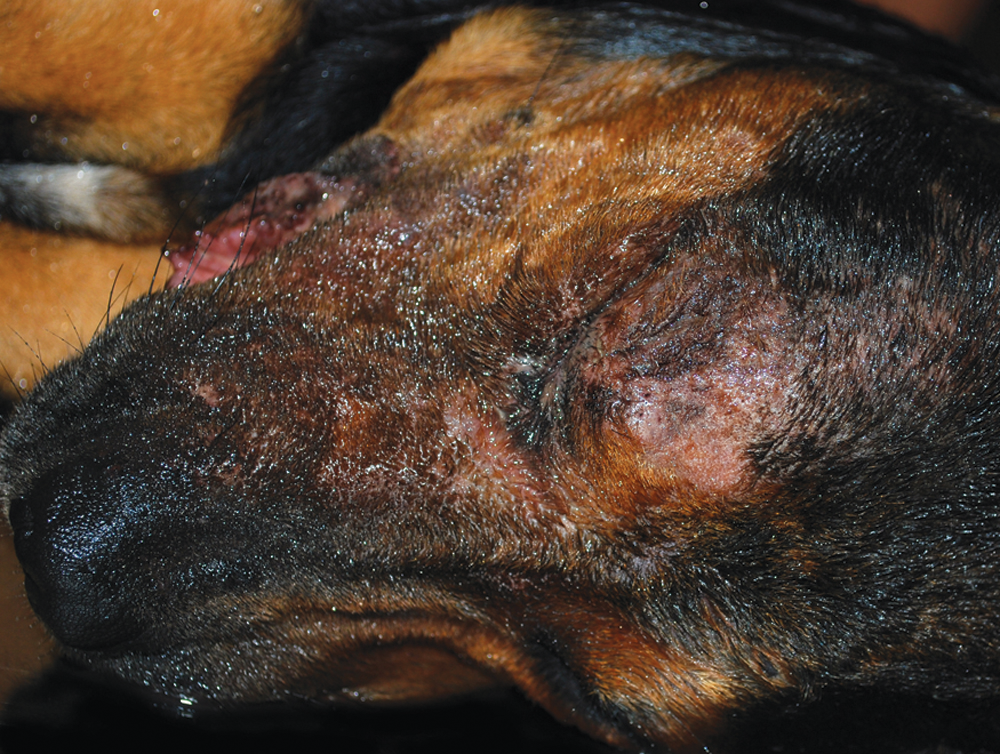
Moist and abraded skin over the right eye and muzzle of dog 3. Skin lesions are the result of self-trauma due to severe facial pruritus induced by infection with Pseudorabies virus.
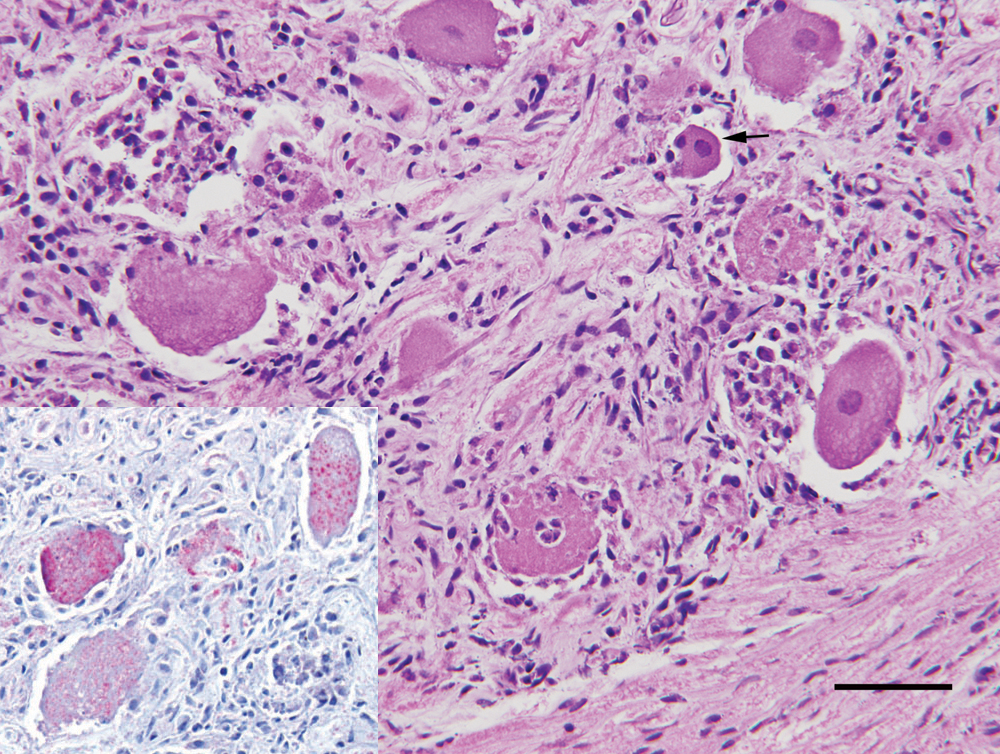
Trigeminal ganglion, dog 3. Inflammation associated with Pseudorabies virus (PRV) infection includes neutrophilic, lymphocytic, and histiocytic infiltrates. Ganglion cells are admixed with necrotic cellular debris, and neuronophagia is prominent. Within a single ganglion cell is an equivocal intracytoplasmic inclusion (arrow). Bar = 50 µm. Inset: immunostaining with antibodies against PRV demonstrated positive immunoreactivity in multiple ganglion cell bodies in the trigeminal ganglion.
Histologic examination of dog 1 confirmed subendocardial hemorrhage and demonstrated pulmonary edema and cecal trichuriasis. In dog 3, both trigeminal ganglia were infiltrated by neutrophils and fewer lymphocytes and macrophages. The right ganglion was more severely affected than the left. Ganglion cells were variably degenerated with multifocal neuronophagia. Scattered nuclei were equivocally distended by large, eosinophilic inclusion bodies with peripheral marginalization of the chromatin (Fig. 2). Few vessels within the medulla oblongata were cuffed by lymphocytes. Facial skin exhibited necro-ulcerative dermatitis; there was acute degeneration of the hind limb musculature; the lungs were moderately edematous; and the small intestine contained multiple strongylid nematodes (Ancylostoma sp.). Freeze–thaw artifact was moderate to marked, limiting interpretation of changes in the adrenal gland, myenteric plexuses, and cardiac plexuses.
Immunohistochemistry
Immunohistochemistry with PRV antibodies on trigeminal ganglia from dog 3 revealed many ganglion cells with positive cytoplasmic immunoreactivity and few ganglion cells with positive nuclear immunoreactivity (Fig. 2).
Virus isolation and identification
Brain homogenates from dogs 1 and 2 produced herpesvirus-like CPE in the inoculated CRFK cell monolayers. Staining of the fixed monolayers with a direct fluorescent antibody conjugate demonstrated the presence of PRV proteins in the infected cells (Fig. 3). The virus isolate was designated as PRV OK2010.
Pooled sample DNA from dogs 1 and 2 was found positive for PRV by real-time gD-specific PCR. Conventional PCR assays for the gC and gE genes also yielded products of the expected sizes. A 654-bp partial sequence of the gC gene of PRV OK2010 was derived and confirmed by BLAST analysis. Phylogenetic analysis, using available corresponding sequences from GenBank, showed that PRV OK2010 appears to be closely related to a cluster of PRV strains previously identified in Illinois (Fig. 4). 14 The Oklahoma strain forms a well-supported clade (bootstrap value of 82%) together with 11 U.S. PRV strains, NIA-3 strain from Northern Ireland, F3303 strain from Brazil, and Becker strain from Europe. Genetic divergence of the 2 other U.S. strains, Indiana S and PRV8044, from PRV OK2010 and its clade-mates was readily apparent.
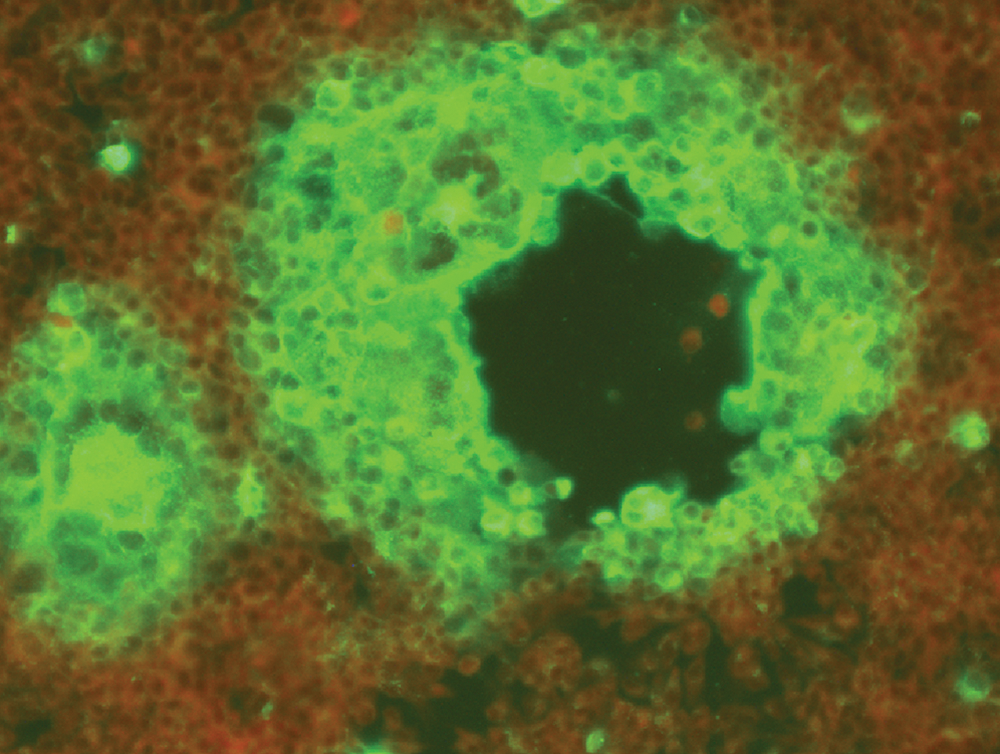
Direct fluorescent antibody staining of Pseudorabies virus isolated in Crandell feline kidney cells from nervous tissue homogenates.
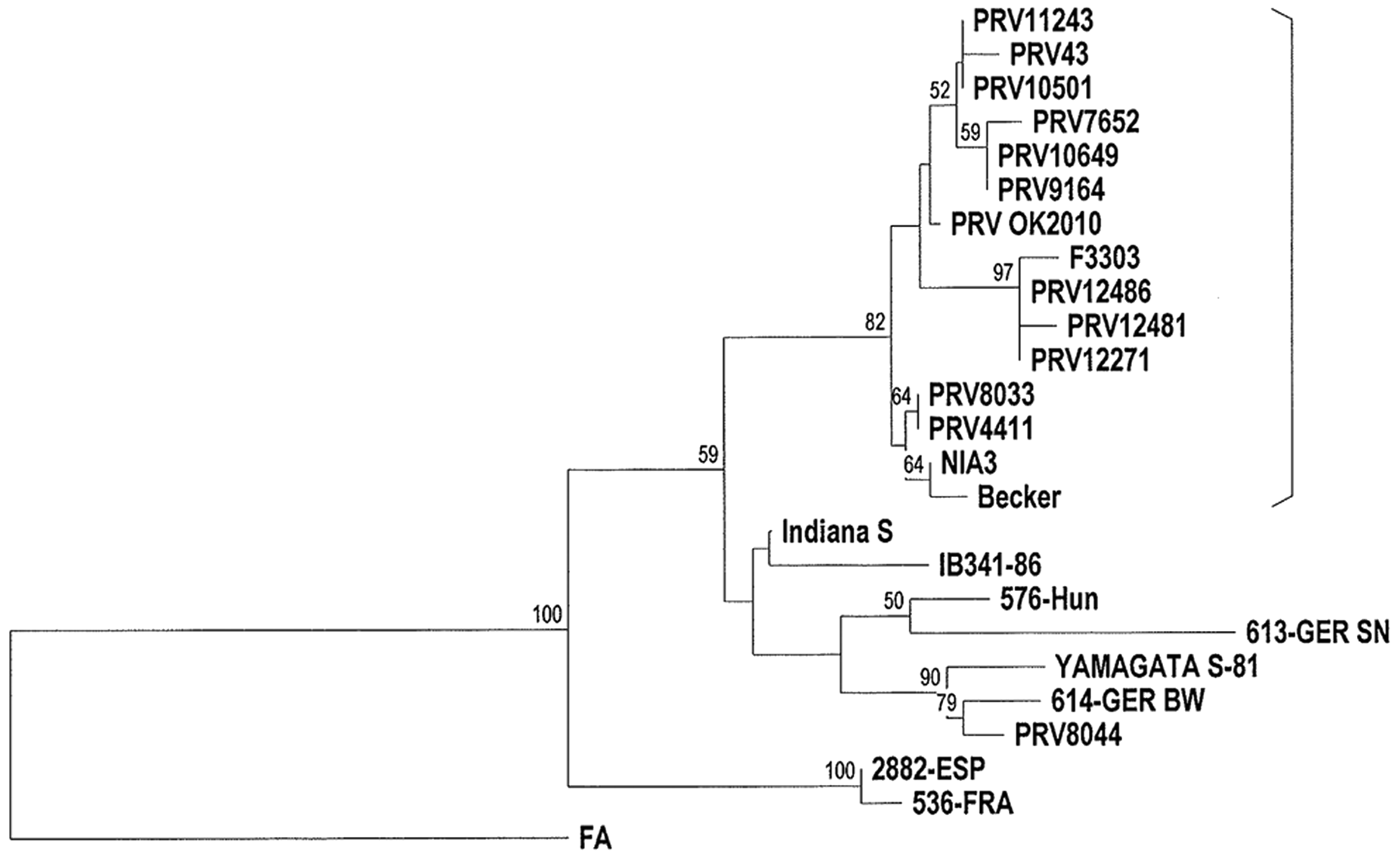
Phylogenetic tree based on the partial nucleotide sequence of the gC gene of Pseudorabies virus (PRV) OK2010 and corresponding sequences of selected PRV strains from GenBank. Origin of strains: PRV11243 (U.S.), PRV43 (U.S.), PRV10501 (U.S.), PRV7652 (U.S.), PRV10649 (U.S.), PRV9164 (U.S.), PRV12486 (U.S.), PRV12481 (U.S.), PRV12271 (U.S.), PRV8033 (U.S.), PRV4411 (U.S.), PRV8044 (U.S.), PRV OK2010 (U.S.), Indiana S (U.S.), F3303 (Brazil), IB341-86 (Brazil), NIA3 (Northern Ireland), Becker (Europe), 576-Hun (Hungary), 613-GER SN (Germany), Yamagata S-81 (Japan), 614-GER BW (Germany), 2882-ESP (Spain), 536-FRA (France), and FA (China). The well-supported clade at 82% bootstrap value is bracketed. The FA strain from China was selected as the outgroup sequence. Bootstrap percentage values are indicated at the nodes.
Toxicological analysis
Toxicologic analysis of the urine of dog 1 was strongly positive for barbiturates and suspicious for benzodiazepines and tricyclic antidepressants by ELISA testing. Diphenhydramine was detected by the thin layer chromatography analysis. Urine from dog 3 was positive for dimenhydrinate, diphenhydramine, and ketamine hydrochloride metabolites by thin layer chromatography analysis.
Discussion
Pseudorabies was suspected in the 3 dogs of the current report based on history of exposure to feral swine, clinical signs, and histologic lesions. Infection was confirmed with immunohistochemistry, virus isolation, PCR, and sequence analysis.
Prior to eradication of PRV in U.S. domestic swine operations, there were a number of reported cases of canine PRV infection in Florida,3,9 Georgia, Louisiana, Alabama, 9 and Missouri. 43 The most recent published report of PRV infection in dogs was in Florida in 2000, in 2 dogs that had fought with feral pigs. 3 Feral swine hunting dogs in Florida apparently continue to be intermittently infected with the virus (Mark Cunningham, personal communication, 2008; www.promedmail.org, archive no. 20081118.3637). Similar to these cases, the dogs described in the current report were likely exposed to PRV through contact with live and/or dead feral pigs. It is not known whether the dogs became infected by consuming carcasses of dead feral pigs (which were present on the premises) or whether they were exposed to the virus during hunting or fighting with live pigs. Subsequent to the outbreak, sanitary measures to limit PRV exposure were implemented, including removal of dead pigs from the premises. Additional dogs have not been infected. In most reported cases, dogs are infected with PRV due to consuming uncooked offal of infected pigs.2,4,10,26
The dogs in the present case manifested a variety of clinical signs consistent with pseudorabies, including the most classic clinical sign: facial pruritus. 31 Though considered classic, the actual incidence of facial pruritus in cases of canine PRV is variable, with 17.8% 22 to 52% 28 of dogs exhibiting this sign. The dogs of the current report also exhibited dyspnea, vomiting, diarrhea, ataxia, muscle stiffness, and death within approximately 48 hr after onset of clinical signs. Most dogs infected with PRV exhibit similar clinical signs, including dyspnea in 60% of cases, vomiting in 36%, ataxia in 76%, and muscle spasms in 36%. Additional clinical signs that are reported to occur with PRV infection include ptyalism in 100% of cases, restlessness in 84%, vocalization in 56%, and aggression in 36%. Dogs usually die within 96 hr of onset of clinical signs, often much sooner. 28
Differential diagnoses for dogs exhibiting these clinical signs include rabies infection, canine distemper, and various toxicoses including organophosphates, heavy metals, ethylene glycol, strychnine, inorganic arsenic, organomercurial compounds, and chlorinated hydrocarbons.2,22,28,36,37 Toxicologic testing for these substances was not performed. However, due to initial suspicion of toxicosis, thin-layer chromatography toxicologic testing was performed on urine of 2 of the 3 dogs. Testing was positive or suspicious for barbiturates, benzodiazepines, diphenhydramine, and tricyclic antidepressants (dog 1), and dimenhydrinate, diphenhydramine, and ketamine hydrochloride metabolites (dog 3). These toxicologic profiles were generally consistent with the treatment histories, though there was no record of tricyclic antidepressant administration to dog 1 or dimenhydrinate administration to dog 3.
Gross lesions in the dogs of the present report were minimal and included facial alopecia and edema, endocardial and epicardial petechiation and ecchymoses, and bloody intestinal contents. (Ancylostoma sp. endoparasitism could have at least contributed to the bloody intestinal contents in the dog with this finding.) Consistent with these gross findings, PRV infection often does not produce gross lesions in dogs,2,36 though affected dogs may have subcutaneous edema, 15 bloody intestinal contents,15,22 and endocardial and epicardial petechiation and ecchymoses. 43
Histologic examination of dog 3 revealed a severe neutrophilic trigeminal ganglionitis and mild lymphocytic cuffing of vessels in the medulla oblongata. This pattern of inflammation is typical of canine PRV infection, which commonly produces inflammation of the trigeminal, facial, glossopharyngeal, and vagal nerves, myelencephalon, and metencephalon. 32 Inflammation may be neutrophilic as in dog 36,7; however, inflammation is more commonly nonsuppurative, characterized by mononuclear cell perivascular cuffing, gliosis, neuronal degeneration, neuronophagia, and intranuclear inclusion bodies in neurons and astrocytes.6,7,10,21,26,32,36,37,43 The distribution of inflammation is consistent with the mechanism of viral spread. Following primary replication, the virus is taken up by sensory nerve endings and spreads by retrograde axonal transport first to the sensory nerve ganglia and subsequently to the central nervous system. 19 In addition to ganglioneuritis and encephalitis, PRV may also cause myocardial degeneration with ganglionitis of the cardiac autonomic plexuses 30 and degeneration of intestinal myenteric ganglia. 15
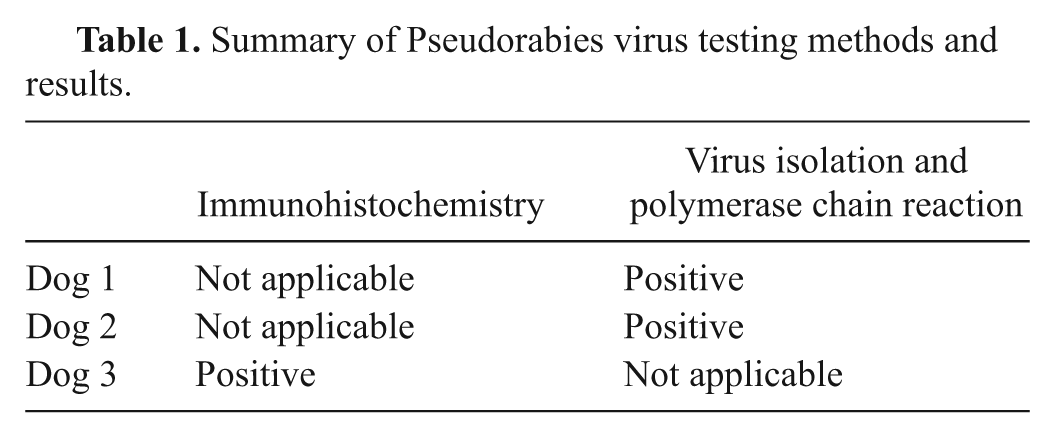
To test for PRV, fresh cerebellum and medulla from dogs 1 and 2 were used for virus isolation and identification (virus isolation and PCR results are summarized in Table 1). Fixed medulla and trigeminal ganglia from dog 3 were used for histopathology and immunohistochemistry (summarized in Table 1). Immunoreactivity was present within the cytoplasm of ganglion cells and few nuclei. This is consistent with a previous report of PRV immunoreactivity 32 and with the assembly mechanism of herpes virions, wherein replication occurs in the nucleus and final maturation occurs in the cytoplasm. 27 Virus isolation4,7,21 and immunohistochemistry10,17,32,37 are often used to diagnose PRV. Other reported tests include a rabbit bioassay,7,26 in situ hybridization, 32 electron microscopy, 21 PCR, 4,37 fluorescent antibody, 21 fluorescence in situ hybridization, 4 and DNA sequencing. 37 Tissues reported to be useful for virus isolation include haired skin from pruritic areas, salivary gland, pharyngeal mucosa, lung, adrenal gland, mesencephalon, medulla oblongata, and cerebellum.7,9,19 Currently, few veterinary laboratories in the United States routinely perform immunohistochemical or PCR testing for PRV on canine tissues (Sarah Cramer, personal communication, 2011).
Summary of Pseudorabies virus testing methods and results.

Though commercial swine operations in the United States were declared free of PRV in 2004, 39 feral hog populations remain variably infected with the virus. 20 The continued presence of PRV in feral populations raises concerns that the virus may be reintroduced to domestic swine,5,20,31 as direct contact between domestic and feral swine has been documented. 44 Multiple serologic surveys in several states have demonstrated variable seropositivity in feral swine herds: 30% in Texas (2004–2006), 44 20–61% in South Carolina (2006, 5 2002 16 ), and 34.8% in Florida (1980s). 40 Serologic survey of feral pigs in Oklahoma failed to demonstrate anti-PRV antibodies in the examined groups of pigs 34 ; however, more recent PCR-based testing has shown that feral swine in Oklahoma are infected with the virus. 20 The positive amplification of the gE gene from the PRV OK2010 isolate revealed that this strain did not originate from the gE-deleted vaccine virus that may be circulating among feral swine in certain parts of the United States.20,42
Since the gC gene of PRV contains considerable genetic variation, it is an ideal target for phylogenetic analysis.14,20,25 A partial sequence of the gC gene of PRV OK2010 was derived and confirmed by BLAST analysis. This sequence was used to construct a phylogenetic tree based upon available sequences in the GenBank database, which compared PRV OK2010 with PRV strains from the United States, Europe, Brazil, and China. Phylogenetic analysis revealed that PRV OK2010 appears closely related to a cluster of U.S. PRV strains previously identified in Illinois 14 with a well-supported clade that includes PRV strains from the United States, Northern Ireland, Brazil, and Europe. A similar grouping of the same U.S. and Northern Ireland viruses has been previously observed. 14 PRV OK2010 and its clade-mates demonstrated notable genetic divergence from U.S. strains Indiana S and PRV8044 (considered genetically similar to the Norden vaccine strain 20 ).
PRV OK2010 likely originated from the feral swine population, considering that the hunting dogs of the current report were frequently exposed to feral swine. Based on the viral phylogeny, it also seems probable that PRV OK2010 is a virulent field virus rather than a vaccine strain. In summary, these findings indicate pseudorabies should remain a differential diagnosis in U.S. dogs and other domestic animals that have a consistent history and clinical signs. Continued surveillance is necessary to prevent reintroduction of PRV into domestic swine.
Footnotes
Acknowledgements
The authors greatly appreciate the help of Ms. Belinda Greene for her key diagnostic suggestion, Ms. Brandy Janicek for excellent technical support, Dr. Gary Pickard for providing the primary antibody for immunohistochemistry, Dr. Vickie Cooper for providing positive immunohistochemistry control tissues, Dr. Tony Gilmour for providing clinical information, and Dr. Tony Confer and Dr. Timothy Snider for histologic consultation.
a.
Ventana BenchMark ULTRA®, Ventana Medical Systems Inc., Tucson, AZ.
b.
Ventana®, Ventana Medical Systems Inc., Tucson, AZ.
c.
Ventana ultraView Red®, Ventana Medical Systems Inc., Tucson, AZ.
d.
Nunc Lab-Tek™ II, Thermo Fischer Scientific, Roskilde, Denmark.
e.
Sigma-Aldrich, St. Louis, MO.
f.
NVSL, Ames, IA.
g.
Nikon Instruments Inc., Melville, NY.
h.
DNeasy® Blood and Tissue Kit, Taq Master Mix Kit, QIAquick® Gel Extraction Kit; Qiagen Inc., Valencia, CA.
i.
SmartCycler II, Cepheid Inc., Sunnyvale, CA.
j.
DNASTAR Inc., Madison, WI.
k.
Toxi-Lab®, Varian Inc., Lake Forrest, CA.
l.
Ontrack Testcard 9, Agilent Technologies Inc., Santa Clara, CA.
m.
SNAP® 4Dx® Test, IDEXX Laboratories Inc., Westbrook, ME.
n.
Baxter International Inc., Deerfield, IL.
o.
VetOne®, MWI Veterinary Supply, Boise, ID.
p.
West-Ward Pharmaceutical Corp., Eatontown, NJ.
q.
Cerenia®, Pfizer Inc., New York, NY.
r.
Hospira Inc., Lake Forest, IL.
s.
Qualitest Pharmaceuticals, Huntsville, AL.
t.
BOMAC Vets Plus Inc., Knapp, WI.
u.
Intervet/Schering-Plough Animal Health, Summit, NJ.
v.
RXV Veterinary Products, Westlake, TX.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
