Abstract
Actinobacillus pleuropneumoniae is the etiologic agent of swine pleuropneumonia, a major production-limiting disease in the pig industry. In the current study, 2,171 lung specimens obtained from pigs housed in 870 Spanish pig farms in regions of substantial pig production were examined. Conventional microbiology, coupled with species-specific polymerase chain reaction, identified 127 biovar 2 isolates, accounting for 25.3% of all A. pleuropneumoniae (n = 502) detected. Most isolates (79%) were recovered as pure primary cultures or as the predominant bacteria from lungs exhibiting lesions typical of acute swine pleuropneumonia. Coagglutination testing identified the isolates as belonging to serovars 2 (4.7%), 4 (4.7%), 7 (68.5%), and 11 (1.6%); however, 26 isolates were nontypeable. All biovar 2 isolates showed genes of the apxII operon alone, which encodes the corresponding ApxII exotoxin, leading to a different gene pattern for isolates in serovars 2, 4, and 11 compared with those of biovar 1. From this survey, it can be concluded that A. pleuropneumoniae biovar 2 infections are common in pigs in Spain, and they may be a common cause of respiratory disease in swine.
Actinobacillus pleuropneumoniae causes porcine pleuropneumonia, a contagious disease of pigs that has an economic impact on industrialized pig production world-wide. 14 Actinobacillus pleuropneumoniae isolates are classified on the basis of the β-nicotinamide adenine dinucleotide (β-NAD) requirement for growth into biovar 1 (β-NAD dependent) and biovar 2 (β-NAD independent), 16 and each biovar is further divided into serovars. Identification of the A. pleuropneumoniae serovar involved in a disease outbreak may help trace the source of the infecting strain through seroepidemiologic studies. Serovar determination is also necessary for effective vaccination by allowing appropriate serovar-specific immunization. Initially, 12 biovar 1 serovars (named 1–12) were recognized and studied extensively. Subsequently, 2 additional serovars among biovar 2 isolates that do not share common antigens with the known biovar 1 serovars were described 6,15 as serovars 13 and 14. Biovar 2 strains that share antigenic determinants with biovar 1 serovars 2, 4, 7, and 9 also have been recognized. 1 Description of the particular serologic and toxin gene profiles of a number of Australian A. pleuropneumoniae isolates led to the characterization of a new serovar (serovar 15) within the biovar 1 group. 2 Thus, a common serovar system that covers all 15 serovars described thus far, regardless of the biovar, is currently in force. 15
Epidemiologic data indicate that virulence of A. pleuropneumoniae is strongly correlated with its ability to produce 1 or 2 exotoxins (Apx toxins). 8 It is possible to group biovar 1 strains showing identical toxin gene patterns by means of polymerase chain reaction (PCR). 9 The resulting groups correlate with serovars and with the virulence observed for each individual strain in the field. 1 ApxI and ApxII are produced by the highly pathogenic serovars 1, 5, 9, and 11; ApxII and ApxIII are produced by serovars 2, 4, 6, 8, and 15; less-pathogenic serovars 7, 12, and 13 produce ApxII; serovars 10 and 14 produce ApxI; and serovar 3 produces ApxIII. 9
The occurrence, regional distribution, and dominant serovars of A. pleuropneumoniae biovar 1 are monitored continuously. 12 However, to our knowledge, there has been no detailed investigation dedicated to A. pleuropneumoniae biovar 2 strains involved in outbreaks of swine pneumonia. Biovar 2 serovars are often not reported in seroprevalence studies, and little is known about the distribution of biovar 2 serovars. Biovar 2 strains of A. pleuropneumoniae were first isolated in the late 1970s from cases of necrotic swine pneumonia and were described as Pasteurella haemolytica-like organisms; these isolates were later classified into the genus Actinobacillus. 16 The isolation of A. pleuropneumoniae biovar 2 has been reported from the early 1980s onward, both in some European countries and in the United States. 6,7,15 More recently, biovar 2 field isolates have been implicated in episodes of clinical disease and outbreaks, with high mortality in fattening pigs. 10 Disease caused by A. pleuropneumoniae biovar 1 is common in growing pigs in Spain, and serovars 2, 4, and 7 are the most prevalent. 12 However, neither the occurrence nor the serovar distribution of A. pleuropneumoniae biovar 2 in Spain has been reported. The present study reports the progression of the relative frequency, serovar distribution, and apx toxin gene patterns of A. pleuropneumoniae biovar 2, isolated from lungs of pigs in Spain during a 7-year period from 2002 to 2008.
The samples examined consisted of 2,171 lung specimens submitted to the Veterinary Diagnostic Centre of Laboratorios HIPRA (Girona, Spain) during the period from January 2002 to December 2008. Samples were obtained from pigs housed in 870 pig farms located in the 4 major Spanish swine-producing autonomous regions (Aragón; Castilla and Leon; Catalonia; and Murcia) distributed around 20 provinces in northern and eastern Spain. The pigs ranged in weight from 30 to 90 kg. Sample submissions were accompanied with a case history, which included respiratory illness in all cases, often followed by sudden death. After disinfection of the lung surface, lung tissue was aseptically swabbed for bacterial isolation. Columbia (supplemented with 5% sheep blood) and chocolate agar plates were inoculated simultaneously. A streak of a NAD-producing Staphylococcus aureus (ATCC 6538) on the Columbia agar plate surface was systematically used as the nurse strain. The plates were incubated at 37°C for 18 hr. Suspect bacteria were purified on Columbia blood agar. Bacteria other than A. pleuropneumoniae were identified to species level by conventional phenotypic methods. Primary identification of A. pleuropneumoniae was based on the Christie, Atkins, Munch-Peterson (CAMP), catalase, urease, and oxidase tests. Bacterial growth features (NAD requirement and β-hemolysis) and appearance (Gram stain) were also considered. To confirm the identity of suspected A. pleuropneumoniae biovar 2, single multiplex PCR using a commercially available PCR mixture a was performed. The PCR assay permits simultaneous identification and serovar group allocation of related A. pleuropneumoniae into 4 groups—GI (biovar 1 serovars 1, 9, and 11); GII (biovar 1 serovars 5a, 5b, and 10); GIII (biovar 1 serovars 2, 4, 7, 8, and 12); and GIV (biovar 1 serovars 3, 4, 6, and 7)—and biovar 2. The performance and suitability of the PCR test for species-specific detection of A. pleuropneumoniae biovar 1 and 2 sequences were established previously. 5 Coagglutination (CoA) with a panel of type-specific antisera to serovars 1–12 b of A. pleuropneumoniae was used for antigen-based typing, as described previously. 13 A serovar was assigned for those isolates that gave a single, serovar-specific reaction within 30 sec. Isolates showing cross-reactivity with 2 or more serovars were considered to be nontypeable (NT). When 2 or more isolates in the same serovar were recovered from an outbreak, only one of the isolates was considered. The apx toxin gene patterns of selected biovar 2 isolates, 13 biovar 1 reference strains (corresponding to serovars 1–12 and 15), and the type strains N-273 (biovar 2 serovar 13) and 3906 (biovar 2 serovar 14) were determined by a previously described apx toxin gene PCR assay. 9 The oligonucleotide primers used in the current study were those developed to amplify, in 5 separate amplification reactions, the activator and structural genes apxICA, apxIICA, and apxIIICA, and the secretion genes apxIBD and apxIIIBD. All product fragments were separated in a single electrophoresis run by measuring differences in length between the individual amplified gene fragments.
Conventional microbiologic analysis of the 2,171 lung samples (coupled with PCR tests on cultures suspected of being biovar 2) led to the isolation and identification of 502 A. pleuropneumoniae isolates. The remaining 1,669 samples were negative to A. pleuropneumoniae. Biovar 1 isolates accounted for 74.7% (n = 375), whereas biovar 2 represented 25.3% (n = 127) of the isolates. Biovar 2 strains isolated were retrieved mostly as pure primary cultures (79%) or as the predominant isolates in mixed cultures together with Pasteurella multocida (9 samples), Streptococcus suis (7 samples), Arcanobacterium pyogenes (4 samples), biovar 1 A. pleuropneumoniae (4 samples), or Haemophilus parasuis (2 samples). Actinobacillus pleuropneumoniae biovar 2 isolates showed copious growth, β-hemolysis, and NAD independence after overnight incubation. Preliminary identification of suspected colonies was straightforward. All biovar 2 isolates scored positive in CAMP, urease, and oxidase tests and scored negative for catalase. The PCR assay based on the omlA gene coding for an outer membrane protein of A. pleuropneumoniae worked according to the validation criteria of the kit manufacturer. A band of the size of the molecular weight marker (provided with the PCR test) allowed assignment of all of the amplicons generated to the typing group corresponding to A. pleuropneumoniae biovar 2. Gross lesions in 116 (91%) of the 127 lung specimens that were positive for A. pleuropneumoniae biovar 2 were consistent with those commonly caused by A. pleuropneumoniae in pigs with acute pleuropneumonia: either suppurative bronchopneumonia (occasionally with small abscess-like nodules) or necrotizing pleuropneumonia, both accompanied by fibrinous pleuritis.
The CoA assay assigned a single serovar to 101 (79.5%) of the 127 biovar 2 isolates tested. Serovars 2 (6/101), 4 (6/101), 7 (87/101), and 11 (2/101) were recorded, with serovar 7 as the most frequent (68.5%). The remaining strains were NT (26/101) because they showed either cross-reactivity to various combinations of serovars 2–4, 6–9, and 11, or no agglutination reaction (Table 1). Cross-reactivity made reading many of the tests difficult. The main cross-reactions were observed between serovars 2 and 4, 4 and 7, and 9 and 11. Serotyping of biovar 1 isolates showed that serovar 4 was the most frequent (42.1%), followed by serovars 2 (24.3%), 9 (9.1%), and 5 (8.8%). Biovar 1 serovars 1, 3, 7, 10, 11, and 12 were isolated sporadically throughout the study and accounted for 39 strains (10.4%). There were 20 NT strains in the biovar 1 cluster (5.3%). The apx toxin gene PCR analysis on 74 selected biovar 2 isolates, representing all serovars recorded (2, 4, 7, and 11), revealed that all 74 isolates possessed genes of the apxII operons exclusively. Hence, isolates in CoA serovars 2, 4, and 11 had different gene patterns compared with their counterparts in biovar 1 reference strains, whereas isolates in CoA serovar 7 in both biovars had identical gene patterns. Reference strain N-273 (biovar 2 serovar 13) showed genes of the apxII operon only, whereas strain 3906 (biovar 2 serovar 14) showed the gene pattern for ApxI toxin, which was unique among all biovar 2 strains analyzed.
Relative frequency and serovar distribution of Actinobacillus pleuropneumoniae biovars 1 and 2 isolated from pneumonic lungs of pigs over a 7-year period in Spain.*
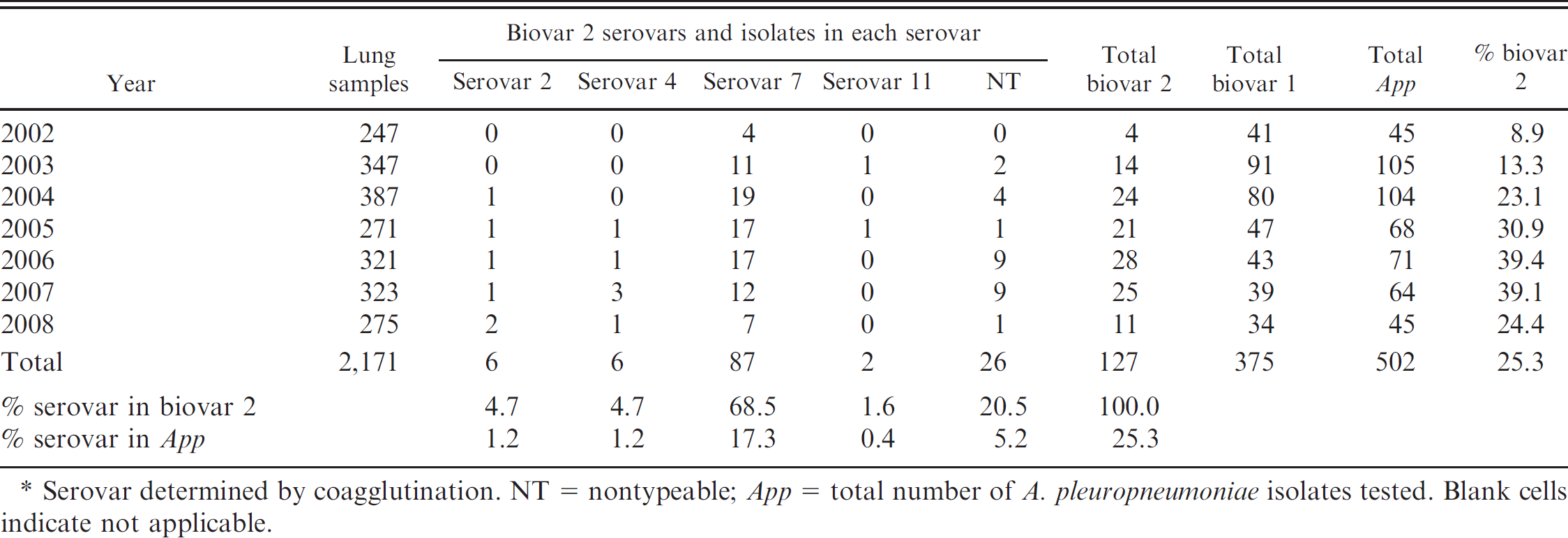
Serovar determined by coagglutination. NT = nontypeable; App = total number of A. pleuropneumoniae isolates tested. Blank cells indicate not applicable.
The present survey revealed that A. pleuropneumoniae biovar 2 infections are frequent in pigs in Spain, and they are a common cause of respiratory disease. Previous reports describe significant proportions of A. pleuropneumoniae infections involving biovar 2 strains in Belgium (12.5%). 4 Note that the growth or biochemical characteristics of atypical A. pleuropneumoniae isolates may lead to the misidentification of some isolates. In addition, biovar 2 strains may go unnoticed by laboratories that are not familiar with this group, because of its NAD-independent growth. It is, therefore, extremely important to consider gross pulmonary lesions, clinical history, and bacteriologic findings so as to facilitate correct diagnosis of swine pleuropneumonia caused by biovar 2 strains. Results of the current study also illustrate how PCR analysis of pure cultures suspected of being A. pleuropneumoniae biovar 2 can be a useful addition to the diagnostic strategy for identity confirmation.
One methodologic limitation of the current study is that biovar 2-positive samples were not tested further for possible coinfections with viruses or Mycoplasma spp. Consequently, it cannot be concluded that the pneumonic lesions observed were caused exclusively by A. pleuropneumoniae biovar 2 isolates. However, A. pleuropneumoniae is considered a primary swine pathogen because of its strong capacity to establish infection and cause disease on its own. Most of the biovar 2 isolates in the present study were recovered from pure primary cultures, indicating that in the absence of other pathogenic bacteria, they were the causative agents of the lesions observed. Note, however, that biovar 2 isolates in the current study contained neither the apxI nor the apxIII genes. Instead, they all showed the apxII gene pattern associated with less-pathogenic serovars, such as biovar 1 serovars 7 and 12. 8,9,11 However, experiments using mutated bacteria demonstrated that ApxII toxin is at least as important as ApxI toxin for the development of clinical and pathologic symptoms. Experimental data from challenge trials also support the significance of putative virulence factors other than Apx toxins in the development of acute lesions caused by A. pleuropneumoniae. 3
The serovar distribution of 71 Spanish A. pleuropneumoniae biovar 1 isolates has been reported previously. Serovars 2, 4, and 7 were predominant, whereas few isolates were serovars 1, 3, 6, 8, 9, and 12; biovar 2 strains were not described. 12 These previous results are generally consistent with the findings presented in the present study for the most prevalent serovars in biovar 1. However, 25.3% of all A. pleuropneumoniae isolated in the present study were biovar 2 strains. Note that the present study involved only passive collection of data from diagnostic material, rather than an exhaustive prevalence survey. Consequently, the data cannot be interpreted as prevalence rates. However, the findings suggest that natural biovar 2 infection in pigs is fairly common in Spain. Remarkably, 2 isolates that reacted strongly to biovar 1 serovar 11 were found. Indirect hemagglutination testing confirmed this cross-reactivity (data not shown). This finding, together with the relatively high proportion of NT biovar 2 strains reported in the present study (20.5%) deserves particular attention because it may indicate higher variability in field isolates of A. pleuropneumoniae biovar 2 than suspected previously. Regarding apx toxin gene patterns, discrepancies were found when comparing isolates and reference strains located in serovars 2, 4, and 11. Unlike reference strains, biovar 2 isolates in those serovars lack the gene for the ApxII toxin. The same divergent results from a previous report 1 lead to a different gene pattern for Swiss and Italian biovar 2 serovars 2 and 4.
In summary, the occurrence and yearly isolation rate in Spain of A. pleuropneumoniae biovar 2 strains from the lungs of pigs dying of pleuropneumonia are reported. To our knowledge, the current study is the first report of A. pleuropneumoniae biovar 2 isolates from infected pigs in Spain. Further characterization of these isolates, particularly determining their antigenic composition and assessing their in vivo infection capabilities, will elucidate their role in the course of the observed clinical disease.
Acknowledgements. The competent technical assistance of Carmen Sánchez is gratefully appreciated. Dr. Patrick Blackall is thanked for revising the manuscript.
Footnotes
a.
Adiavet® APP multiplex PCR kit, Adiagene, Saint-Brieuc, France.
b.
Biovac, Angers, France.
