Abstract
Astroviruses are important pathogens of human beings and animals. Feline astroviruses have been identified by electron microscopy in the feces of either asymptomatic or symptomatic cats, and experimental infection with one isolate was shown to induce enteric clinical signs and virus excretion, thus suggesting a possible role as enteric pathogen. However, due to the lack of specific diagnostic assays, feline astroviruses are not included in the diagnostic algorithms of feline infectious diseases, and their role as feline pathogens remains unclear. The present report describes a dual infection by Feline astrovirus and Feline panleukopenia virus in a 4-month-old cat with severe gastroenteritis and panleukopenia. Upon sequence analysis, the Feline astrovirus strain was found to be genetically related to astroviruses identified in human beings and felids.
Astroviruses (AstVs; family Astroviridae) are small round viruses with a positive single-stranded RNA genome and a polyadenylated tail. Astroviruses have been identified in human beings and in a variety of terrestrial and marine mammals, as well as in several avian species, and they are classified into 2 genera, Mamastrovirus (AstVs of mammals), and Avastrovirus (AstVs of avians). 16,22 The genome (6.4–7 kilobases) consists of 3 open reading frames (ORFs). ORF1a and ORF1b encode nonstructural proteins, a serine protease, and a RNA-dependent RNA polymerase (RdRp), respectively. Open reading frame 2 encodes the capsid precursor. 16 In human beings and animals, AstVs are usually associated with mild to severe forms of enteritis. Some avian strains have also been associated with fatal intestinal and extraintestinal diseases. 16
Feline astrovirus (FAstV) was first identified by electron microscopy (EM) from the stool of a domestic kitten with diarrhea. 8 Subsequent EM-based investigations demonstrated that FAstV may be common in the stool of cats with and without diarrhea. 14,21 Experimental infection with a FAstV strain in specific pathogen–free (SPF) kittens induced enteritis and viral excretion for several days. 7 However, FAstV is not included in the diagnostic algorithms of feline enteric diseases, and its role as a primary pathogen or synergistic agent remains unclear. In order to fill this information gap, FAstV has been included in the panel of feline enteric viruses for routine testing of cases of feline enteritis in the authors’ laboratory.
Feline panleukopenia virus (FPV) infection was diagnosed in a 4-month-old cat hospitalized with severe gastroenteritis. The cat had not been vaccinated for FPV and developed lethargy, anorexia, fever (39.8°C), persistent vomiting, dehydration, and diarrhea. Two days after the onset of the signs, the animal was presented to the veterinary clinic, and appropriate re-hydrating therapy and antibiotic treatment were started. Blood cell count revealed marked leukopenia (1.8 × 103cells/mm3), and a parvoviral infection was suspected. The animal’s condition slowly improved, and the cat completely recovered after 7 days. At the time of hospitalization, a rectal swab from the kitten was collected and submitted to the authors’ laboratory for virological investigations. The diagnostic panel included FPV, Feline rotavirus (FRV), Feline calicivirus (FCV), and Feline coronavirus (FCoV), using either conventional or quantitative polymerase chain reaction (qPCR) and reverse transcription (RT)-PCR. 3–5,15 The animal tested negative to FRV, FCoV, and FCV but positive to FPV, thus confirming the clinical suspicion. Unexpectedly, the sample also tested positive in RT-PCR for AstV based on a broadly reactive set of AstV-specific primers consisting of a mixture of 2 forward primers, 5′-GARTTY GATTGGRCKCGKTAYGA-3′ and 5′-GARTTYGATTGGR CKAGGTAYGA-3′, and 1 reverse primer, 5′-GGYTTKAC CCACATNCCRAA-3′, targeting the ORF1b region. 2 The presence of astroviral RNA was confirmed by RT-PCR using primers specific for human AstVs: Mon 269 (5′-CAACTCAGGAA ACAGGGTGT-3′) and Mon 270 (5′-TCAGATGCATTGTCA TTGGT-3′), targeting conserved amino acids motives in the capsid region (ORF2). 18 A one-step RT-PCR procedure a was used for reverse transcription, and the reactions were performed using the following thermocycling conditions: reverse transcription at 50°C for 60 min and 94°C for 2 min; 35 cycles at 94°C for 1 min, 50°C for 1 min, 68°C for 1 min, with a final step at 68°C for 10 min. The PCR products were resolved in 1% agarose gels. Bands were purified b and sequenced. Purified amplicons were analyzed with a commercial sequencing kit and an automated electrophoresis system. c The sequences were assembled using BioEdit software package version 7.0 6 and compared to cognate sequences in the genetic databases using BLAST (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) and FASTA (http://www.ebi.ac.uk/Tools/sss/fasta/nucleotide.html) web-based programs. Phylogenetic reconstruction was carried out with various algorithms of distance correction and the neighbor-joining method, supported with bootstrapping over 1,000 replications. Distance analysis and phylogenetic inference were carried out using MEGA software package 3.0. 13
Upon sequence analysis (373 bp in the RdRp and 447 bp in the capsid gene; GenBank accession nos. GU980969 and GU980970), the strain detected in the current study (designated Fe/443/07A) displayed the highest nt identity in the RdRp (93.4%) and capsid protein gene (92.7%) to a cheetah AstV. 1 Identity to human AstVs was lower (<74.9% and < 83.6% nt) in the RdRp and ORF2 regions, respectively. Identity to FAstV (strain Bristol) was 86.3% nt in the capsid gene. No ORF1 sequence of feline viruses was available to compare within the databases. Upon phylogenetic analysis, strain Fe/443/07A clustered with mammalian AstVs within a lineage comprising human, cheetah, and feline AstVs (Fig. 1A, 1B).
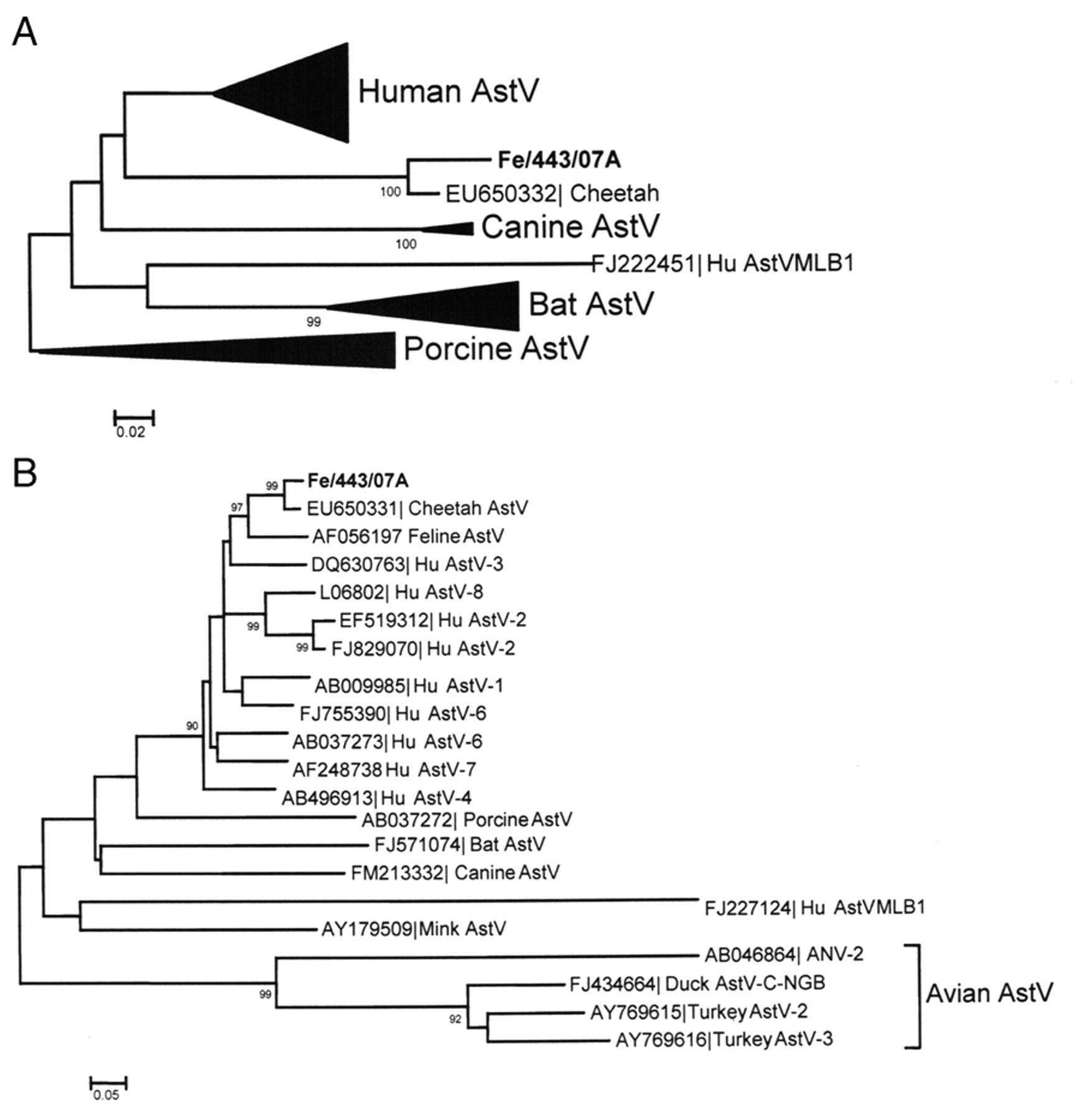
Phylogenetic trees based on partial sequences of
Due to the limited number of studies and reports, the role of AstV as an enteric pathogen in cats is unclear. Natural infection by AstV has been reported in domestic cats with vomiting, diarrhea, anorexia, dehydration, and fever. The animals completely recovered from the disease 7–10 days after the onset of signs. 7,8 An enteric disease has been reproduced experimentally in SPF kittens using the FAstV strain Bristol. 7 In the current study, a dual infection by FAstV and FPV was diagnosed in a cat presenting with severe gastroenteritis. Feline panleukopenia virus is an immunosuppressive agent causing severe leukopenia, enteric lesions, and neurological signs. Mixed viral infections in cats have been documented in several studies. 12,23 Possible interactions and synergistic mechanisms of enhancement of viral pathogenicity may not be ruled out. For instance, co-infection by Canine parvovirus (highly related to FPV) has been hypothesized to trigger the pathogenicity of coronavirus in dogs. 20
Many cases of enteritis in cats remain of unknown etiology. In all studies reporting on FAstVs, EM was used to reveal the presence of AstVs. It is safe to assume that the prevalence of FAstV based on EM has been underestimated, considering the limited sensitivity of EM and considering that AstVs may be easily confounded with other small round viruses (such as caliciviruses and enteroviruses). 19 The RdRp region represents the most conserved part of the astroviral genome and has been exploited to develop pan-AstV primer pairs, 2 which have been successfully used to diagnose FAstV infection in cats. Gathering information on the genetic diversity of FAstV will be important to design effective, more specific primer sets.
The high genetic relationship observed between the feline strain Fe/443/07A and the AstVs detected from captive cheetahs supports a previous hypothesis that stray cats may act as a reservoir of AstV infection for cheetahs. 1 Transmission of viruses (FPV, Felid herpesvirus 1, FCoV, and FCV) between cheetahs and domestic cats has previously been described. 11,17 Astroviruses are considered to be species-specific viruses, but interspecies transmission and/or recombination between either related or unrelated species has been hypothesized, involving cows and red deer, human beings and marine mammals, and human beings and/or ruminants/minks. 10,16,22
Interestingly, in the capsid gene (ORF2), FAstVs appear to resemble human AstVs more closely than other mammalian viruses, thus suggesting a relatively recent intersection in their evolution. 9 Also, a feline-like AstV has recently been detected in a symptomatic child, suggesting a possible direct cross-species infection (Walter JE: 2002, Genetic characterization of astroviruses associated with diarrhea among children in a periurban community of Mexico City. PhD diss. Medical School, University of Pecs, Pecs, Hungary). In the present study, the feline strain Fe/443/07A was recognized by primers Mon 269 and Mon270 that have been designed to recognize specifically human AstVs. 18
Although a number of viruses diverse from FPV have been identified in the feces of cats, these viruses are not included in the diagnostic algorithms of gastroenteric diseases of cats, thus portraying, often, an incomplete picture. Whether and to what extent AstVs may impact feline health, alone or in mixed infections with other enteric pathogens, deserve further study.
Footnotes
a.
SuperScript® III One step, Invitrogen Ltd., Paisley, United Kingdom.
b.
QIAquick® gel extraction kit, Qiagen Inc., Valencia, CA.
c.
BigDye® Terminator v3.1 Cycle Sequencing Kit, ABI Prism 3100 Genetic Analyzer; Applied Biosystems, Foster City, CA.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
