Abstract
The objective of the present investigation was to differentiate between strains of Streptococcus equi subspecies equi implicated in abscess formation in vaccinated horses. Streptococcus equi isolates recovered from clinical specimens associated with equine strangles cases submitted to the University of Illinois Veterinary Diagnostic Laboratory were compared with S. equi isolates representing at least 12 lots of a commercial modified live vaccine (MLV) to determine whether the isolates obtained from the abscesses were vaccine or wild type. Genotyping techniques evaluated included enterobacterial repetitive intergenic consensus polymerase chain reaction (PCR), repetitive extragenic palindrome PCR, BOX element PCR, ribotyping, and pulsed-field gel electrophoresis (PFGE). Phenotypic evaluations were performed using the Biolog GP2 Microplate (hereafter, Biolog). In cases where Biolog and PFGE results did not coincide, a single nucleotide polymorphism located in the upstream regulatory region of szp gene was used to identify the S. equi strains. PFGE and Biolog successfully differentiated wild-type S. equi strains isolated from clinical submissions from isolates of the MLV. PFGE genotyping enabled further subtyping of the wild-type strains, whereas Biolog combined with szp sequencing was useful in differentiating the MLV strain from its wild-type progenitor. Deletion of a single guanine residue located in the upstream regulatory region of the szp gene appears to be conserved among vaccine isolates, and shows a 98.5% correlation to Biolog identification. This multiphasic approach can be used to answer specific diagnostic questions pertaining to the source of infection and/or outbreak, or to address quarantine concerns.
Introduction
Streptococcus equi subspecies equi, a Lancefield group C streptococcus, is the etiologic agent of equine strangles. 6 Strangles is an upper respiratory disease causing suppurative lymphadenitis of the regional lymph nodes including the retropharyngeal nodes, which, when swollen, can potentially result in obstruction of the airway. Serious complications may include septic spread of the organism forming abscesses at distant sites (bastard strangles) and the immune complex disease, purpura hemorrhagica. Upon clearing the infection, horses can harbor bacteria in guttural pouches for prolonged periods and may serve as asymptomatic carriers. 11
Beginning in 2002, increasing submissions to the University of Illinois Veterinary Diagnostic Laboratory (UIVDL; Urbana, Illinois) of nasal and abscess swabs from vaccinated horses suspected of strangles were observed. In acutely ill horses with clinical strangles, both dry and mucoid colony morphologies of S. equi could be recovered, often from the same abscess. The currently available modified live vaccine a (MLV) is an acapsular mutant with exposed surface antigens, which can be quickly recognized and eliminated by the host immune system. This MLV has a predominantly dry morphology but can produce encapsulated isolates from reconstituted vaccine as well as serial passages of dry morphology colonies on rich media. 16 Although movement of horses infected with wild-type isolates may be restricted by quarantine, movement of horses with vaccine-related infections is generally not. Hence, a rapid reliable method to differentiate strains is needed. Differentiation of S. equi strains using multilocus enzyme electrophoresis and multilocus sequence typing techniques has been largely unsuccessful because of the highly homogeneous nature of S. equi genome. 8,9 Sequence variation in the sem gene, which encodes the M protein (SeM) of S. equi, has been observed, and a molecular subtyping method for S. equi isolates exploiting these variations in sem has been proposed. 2 Large restriction fragment polymorphisms have also been used for typing of S. equi strains. 13 However, there are no widely accepted methods available for S. equi genotyping.
Because the dry phenotype is an unreliable marker of the vaccine strain, and as the S. equi strains are highly similar, 5 it soon became a diagnostic necessity to 1) differentiate vaccine strains from virulent wild-type strains, 2) differentiate the wild-type field strains from the wild-type progenitor of the vaccine strain, and 3) distinguish the vaccine strain from its wild-type progenitor. Thus, the objective of the current study of typing S. equi was expanded to encompass all aspects of differentiation. Several molecular methods including ribotyping, pulsed-field gel electrophoresis (PFGE), enterobacterial repetitive intergenic consensus (ERIC) polymerase chain reaction (PCR), repetitive extragenic palindrome (rep-PCR), BOX element (BOX) PCR, and DNA sequencing of the upstream regulatory region of the szp gene (szp sequencing) were, therefore, evaluated for the genetic differentiation of S. equi to distinguish wild-type isolates from vaccine isolates, and differentiate vaccine and wild-type isolates from the wild-type progenitor of the vaccine. Those methods that were discriminatory for vaccine and wild-type strains were compared with phenotypic data derived from the Biolog identification system to validate a rapid, inexpensive method of typing that is useful for diagnostic laboratories.
Materials and methods
Culture and Biolog assay
Between 2002 and 2008, a total of 403 equine nasal and abscess swabs or S. equi isolates submitted to UIVDL and isolates representing at least 12 lots of commercial MLV were plated on Columbia blood agar (BA), Columbia BA with colistin and nalidixic acid, trypticase soy agar with 5% sheep blood and amikacin (64 μg/ml), and MacConkey agar b and incubated overnight at 37°C and 5% CO2. Hemolytic colonies were subcultured onto BA and incubated overnight at 37°C and 5% CO2.
Gram staining, catalase testing, and serogrouping with the latex bead agglutination test c were performed to confirm the isolates as Lancefield group C streptococci. Isolates were streaked onto Biolog universal growth agar plates, d incubated overnight at 37°C, suspended in inoculating fluid, d and dispensed into Biolog GP2 Microplate d per manufacturer's instructions. Oxidation and reduction reactions of 95 amino acids, carbohydrates, and other substrates were measured at 690 nm as increasing tetrazolium dye absorbance by the reactants. Biolog data produce 95 substrate utilization patterns for each of the isolates tested. Streptococcus equi ssp. equi isolates were tested a total of 3 times with the Biolog identification system d and optical density values for wells of interest were averaged.
Genotypic characterization
Ribotyping. Streptococcus equi isolates were subjected to ribotyping e using the enzymes EcoRI, PstI, HinfI, PvuII, or AseI f following the manufacturer's instructions.
Genomic DNA extraction. Genomic DNA templates used in PCR reactions were extracted from plate cultures using a commercial kit g according to the manufacturer's instructions for processing gram-positive bacterial cultures.
Enterobacterial repetitive intergenic consensus polymerase chain reaction. Each 50-μl reaction mix contained 1× PCR buffer, 3 mM MgCl2, 4 U of Taq DNA polymerase h (5 U/μl), 200 μM deoxyribonucleotide triphosphates (dNTPs), h 1 μM ERIC1R primer i (5′-ATGTAAGCTCCTGGGGATTCAC-3′), 1 μM ERIC2 primer i (5′-AAGTAAGTGACTGGGGTGAGCG-3′), 30.2 μl of DNAse-free water, j and 200 ng of purified DNA template. Amplification parameters included 94°C for 3 min followed by 40 cycles of 30 sec at 94°C, 3 min at 51°C, and 2 min at 72°C, and a final extension step of 7 min at 72°C. 12
Repetitive extragenic palindrome polymerase chain reaction. The rep-PCR assay was performed using repPRO Uprime-Dt test kit, k which included Uprime-Dt primer 15 and all the reagents except Taq polymerase. 1 Each 50-μl rep-PCR reaction mix was prepared according to manufacturer's directions with 200 ng of total purified DNA template using the primer and reagents supplied in the kit and 2 U of Taq polymerase h under the following conditions: 95°C for 2 min followed by 31 cycles of 30 sec at 93°C, 1 min at 40°C, and 8 min at 65°C, and a final extension step of 8 min at 65°C.
BOX element polymerase chain reaction. Each 50-μl reaction mix contained 1× PCR buffer, 4 mM MgCl2, 4 U of Taq DNA polymerase h (5 U/μl), 200 μM dNTPs, h 1 μM each of BOX A primer i (ATACTCTTCGAAAATCTCTTCAAAC), BOX B primer i (5prime;-CACGAGAAACTAAAAGTAACTCATAT-3′), BR-1 primer i (5′-CTGACTTCGTCAGTTCTATC-3′), and BR-2A primer i (5′-TTTGTGACAAAACTCCAACA-3′), 27.2 μl of DNAse-free water, j and 200 ng of purified DNA template. 14 Amplification parameters were as follows: 94°C for 3 min followed by 40 cycles of 40 sec at 94°C, 1 min at 45°C, and 2 min at 72°C, and a final extension step of 5 min at 74°C. All amplicons were electrophoresed on 1.5% agarose gel f containing 0.5 μg/ml of ethidium bromide j for 50 min at 120 V.
Pulsed-field gel electrophoresis. Electrophoresis was performed using a modification of a protocol provided by Dr. Tanja Oppriessnig, University of Iowa, Ames, Iowa (personal communication). This protocol is similar to that reported previously. 13 Briefly, S. equi isolates were plated on BA, incubated overnight in 5% CO2 at 37°C, and resuspended in 1 ml of cell suspension buffer (100 mM Tris–HCl, 100 mM ethylenediamine tetra-acetic acid [EDTA], pH 8.0). Ten microliters of 25 mg/ml lysozyme l was added to 400 μl of the cell suspension, and the tubes were incubated for 30 min at 37°C. Then, 450 U of mutanolysin, l 20 μl of proteinase K, g and 400 μl of molten plug agar (1% certified low melt agarose, m 0.5 g sodium dodecyl sulfate j dissolved in 10 mM Tris-HCl, j 1 mM EDTA, j pH 8.0) were added to the suspension, mixed with a Pasteur pipette, and dispensed into PFGE plug molds. Plugs were allowed to solidify at room temperature for 15 min and stored at 4°C until further use. Each plug was incubated in 1.5 ml of cell lysis buffer (1% Sarcosyl j in 50 mM Tris–HCl, j 50 mM EDTA, j pH 8.0) and 80 μl of proteinase K g at 55°C water bath for 1.5 hr. Plugs were then washed 4 times in 50°C sterile distilled water for 15 min each, followed by 4 washes in 50°C TE (10 mM Tris–HCl, 1 mM EDTA, pH 8.0) for 15 min each. Restriction digests contained a thin plug slice in 175 μl of sterile water, 20 μl of appropriate buffer, and 50–100 U of restriction enzyme, f and were incubated at the appropriate temperatures (30μC for SmaI and 37μC for NotI) overnight. Pulsed-field gel electrophoresis was performed in a commercial unitm using 1% PFGE agarose gelsl in 0.5× Tris–borate–EDTA buffer at 6 V/cm; 120° included angle; 14°C; pulse times of 5–15 sec for 11 hr and 15–45 sec for 13.5 hr. Gels were stained for 1 hr with 0.5 μg/ml of ethidium bromide solution, destained in sterile distilled water, and photographed on an ultraviolet transilluminator.
szp sequencing. The PCR assay was performed with DNA polymerase f according to the manufacturer's directions. The following primer sequences were designed i and used in the assay: szp-1700F (5′-GCCAATTGAGTCAAGACAATCA-3′); szp-1700R (5′-GTGAGCAAGGCTTAGTCTCTCC-3′; Patterson et al., current study). The amplicons were subjected to DNA sequencing at the W. M. Keck Center for Comparative and Functional Genomics (University of Illinois at Urbana-Champaign) using the following primers: szp SEQ-F (5′-GCTCTAGCGATATAGCAGGG-3′); szp SEQ-R (5′-GCTGATACAGCT GCCGCACC-3′; Patterson et al., current study). Sequence alignments were performed using bioinformatics software. n
Statistical analysis
Hierarchical cluster analysis was performed on the Biolog absorbance data using centroid sorting method and Ward's linkage algorithm. Analysis of variance was performed on szp sequencing data and Biolog data using a commercial software. o
Results
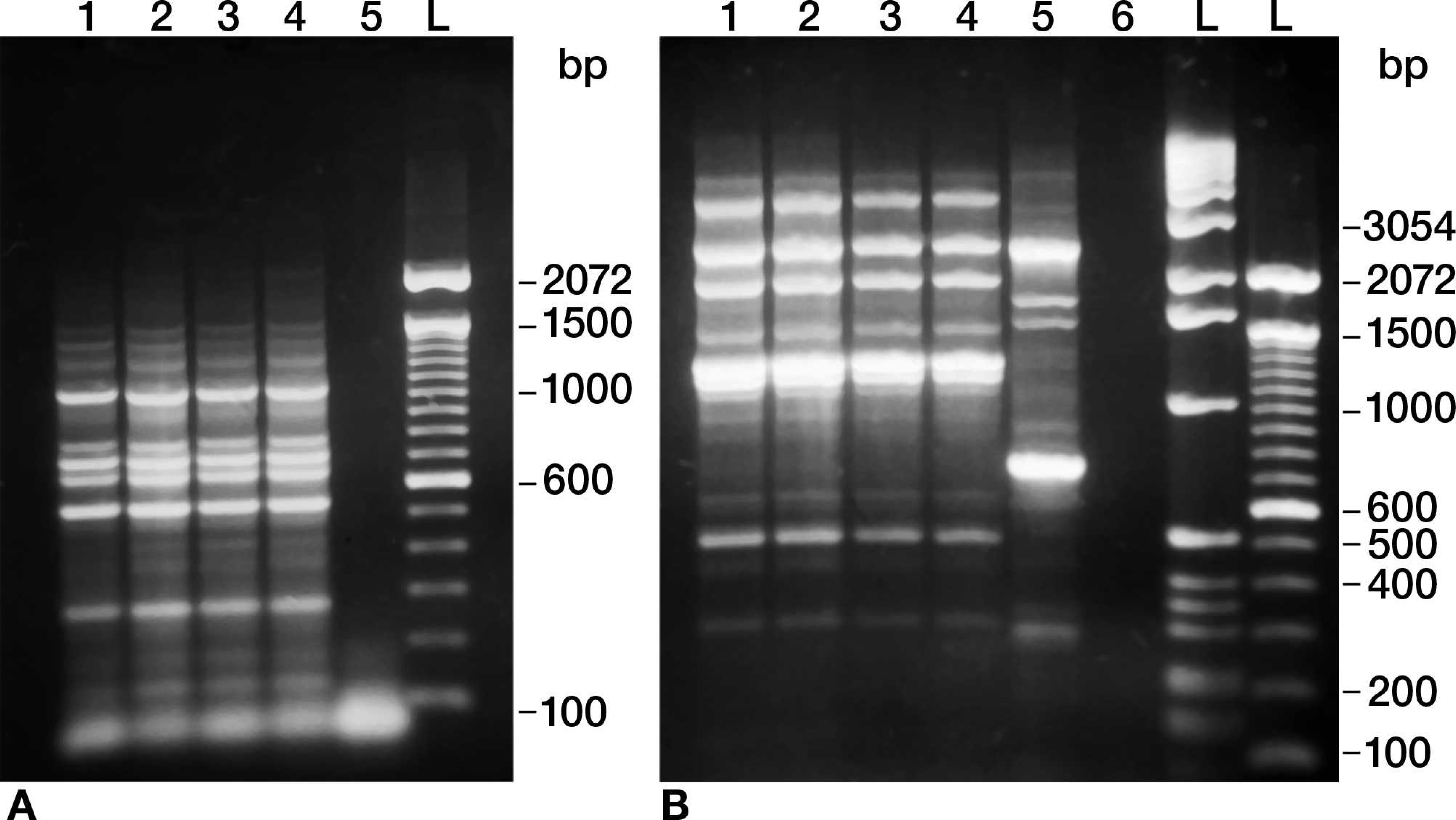
None of the random polymorphic DNA amplification techniques (i.e., ERIC PCR, rep-PCR, and BOX PCR; Fig. 1) or ribotyping with restriction enzymes EcoRI, PstI, HinfI, PvuII, or AseI (Fig. 2) were able to discriminate between any of the S. equi isolates tested. Ribotyping with PvuII consistently yielded 4 bands, and only a single band was obtained with the other enzymes tested.
Pulsed-field gel electrophoresis using SmaI and NotI restriction enzymes produced banding patterns that clearly distinguished vaccine from wild-type strains except the wild-type progenitor strain, as well as distinguished wild-type strains from each other (Fig. 3). The SmaI digests detected 13 different patterns, and the NotI digests detected 30 different patterns among the wild-type strains (not all restriction digest patterns are shown in the figures). Although SmaI digests identified a single banding pattern, NotI digests identified 2 different banding patterns, vaccine 1 and vaccine 2, for the vaccine strains used (Fig. 4). Only isolates exhibiting vaccine patterns with both SmaI and NotI restriction digests were designated vaccine-like.
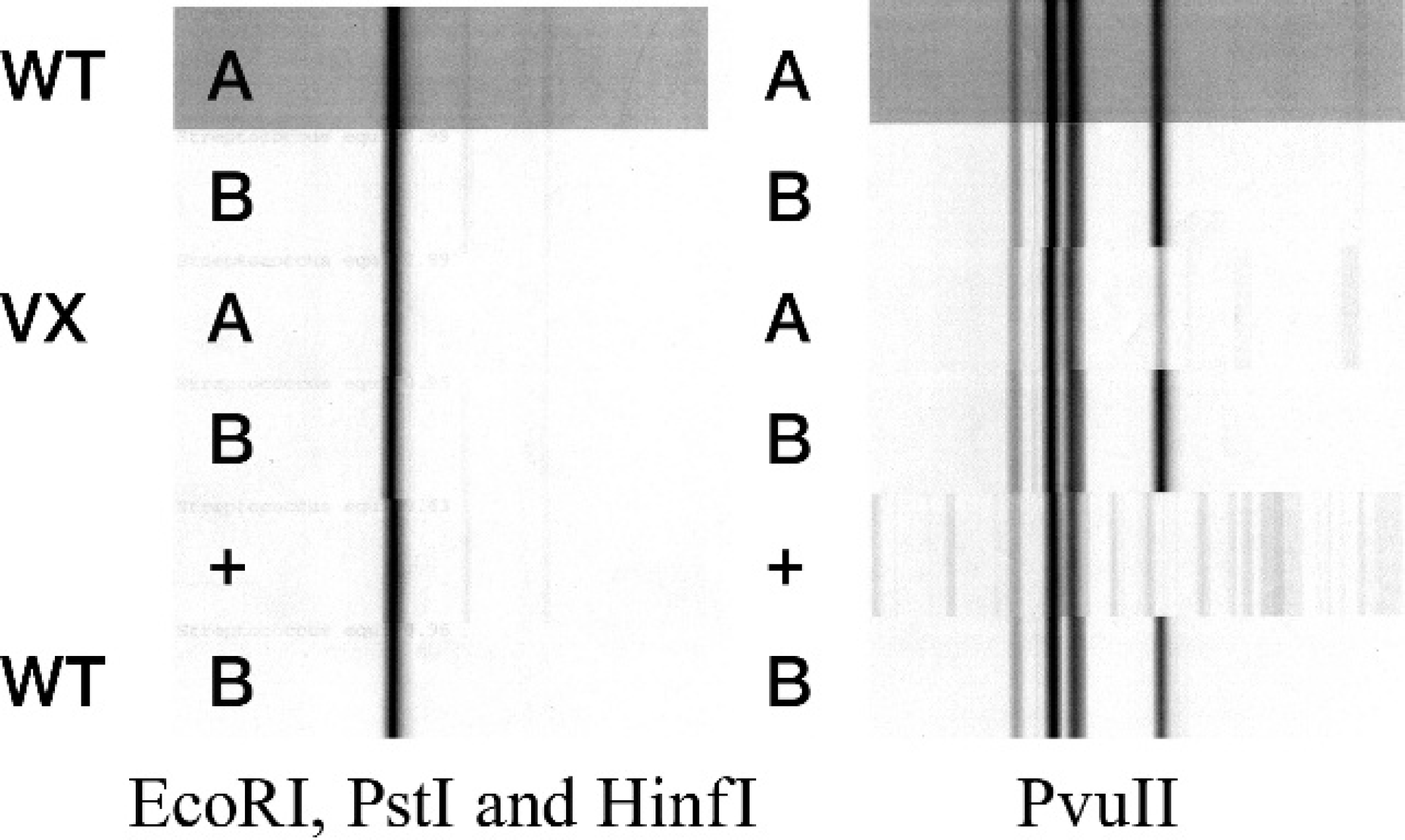
Ribotyping patterns of Streptococcus equi subspecies equi isolates. Wild type (WT) A and B = wild-type isolates; vaccine (VX) A and B = vaccine isolates; + = known S. equi isolate.
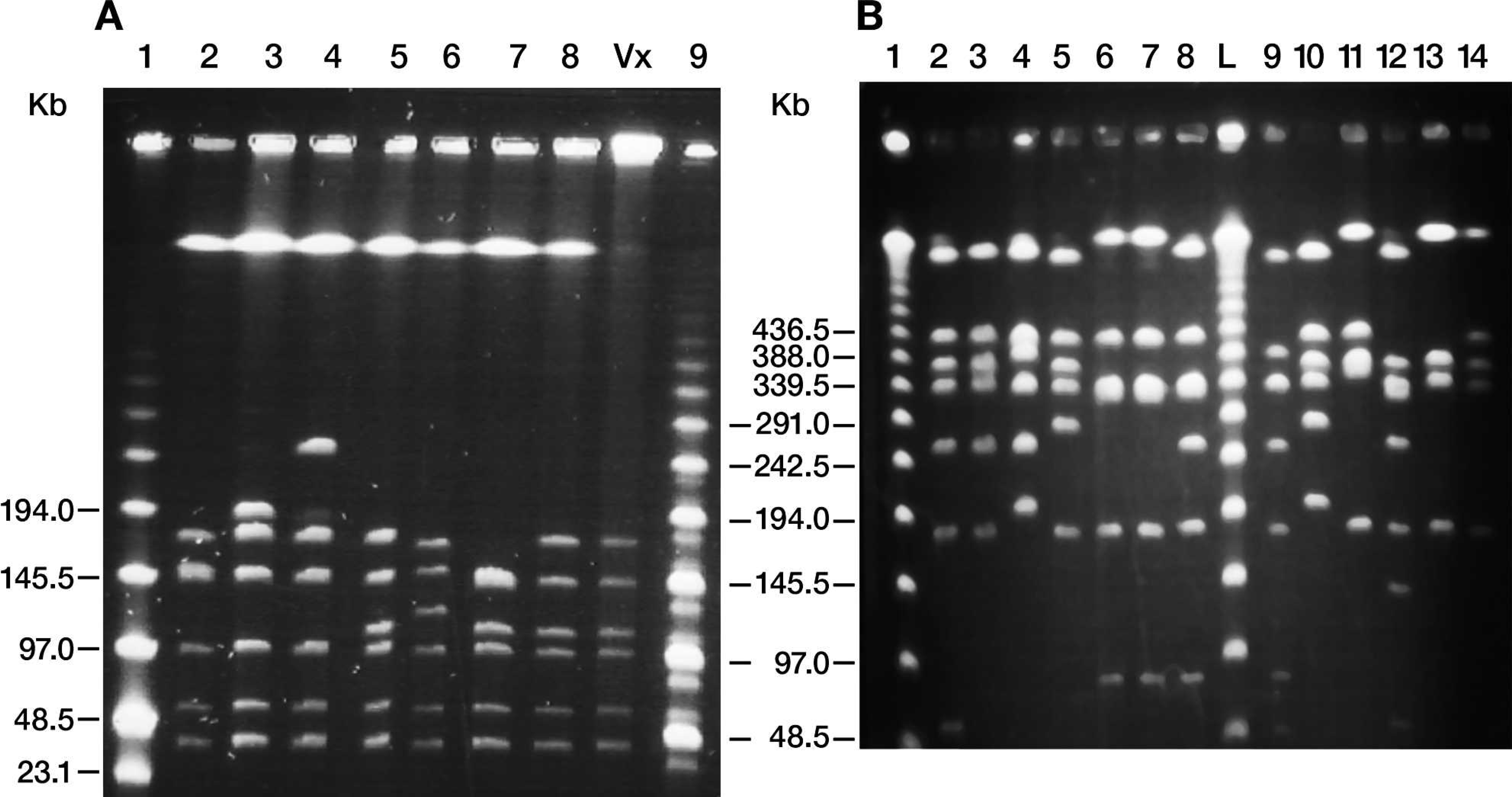
Pulsed-field gel electrophoresis (PFGE) patterns of Streptococcus equi subspecies equi isolates. A, digested with SmaI. Lanes 2–8: S. equi isolated from abscesses; lane Vx: S. equi isolated from vaccine; lane 1: low-range PFG marker; lane 9: mid-range I PFG marker.
In a preliminary investigation of Biolog data obtained from 126 isolates, histograms of the absorbance data from all substrates were developed and evaluated to eliminate consistently negative and redundant variables, which were excluded from further analysis. Four bimodal variables (glycogen, 3-methyl glucose, psicose, and methyl pyruvate) were identified to be suitable for distinguishing wild-type strains from vaccine strains. Both mucoid and dry colony morphologies recovered directly from several lots of MLV demonstrated consistently higher absorbance values for these 4 substrates compared with wild-type isolates. High usage of glycogen, 3-methyl glucose, psicose, and methyl pyruvate was, thus, considered to be a vaccine phenotype. In both vaccine and wild-type isolates, dry colony morphology phenotype often exhibited relatively higher utilization of glycogen than mucoid colony morphology phenotype. In addition, usage of psicose and methyl pyruvate were highly correlated (r = 0.747, P < 0.00001) to each other. Isolates exhibiting high utilization of psicose also exhibited high utilization of methyl pyruvate. Wild-type clinical isolates, including the wild-type progenitor of the vaccine strain, consistently demonstrated low absorbance values for these variables. Hierarchical cluster analysis using Ward's linkage algorithm as well as centroid sorting method was performed on substrate utilization data obtained from these 4 bimodal variables. Sensitivity and specificity of the Biolog data for these 4 variables (data not shown) from the 126 isolates were calculated using a 2 × 2 contingency table with PFGE as the gold standard and were found to be 93.5% (72/77; 95% confidence interval [CI]: 85.7–97.2) and 89.8% (44/49; 95% CI: 78.2–95.6), respectively.
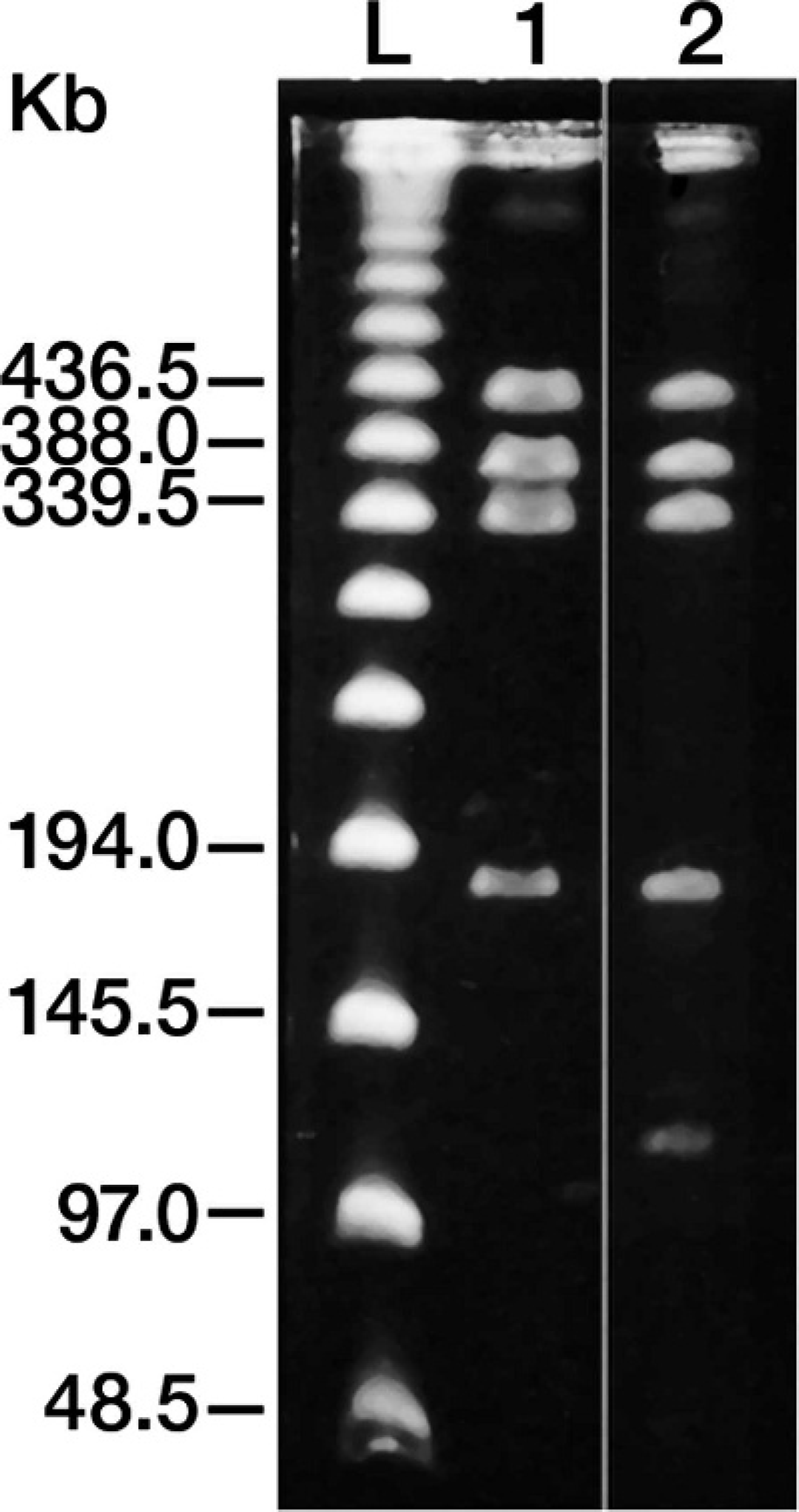
Two NotI restriction patterns associated with vaccine isolate. Lane 1: vaccine 1; lane 2: vaccine 2; lane L: lambda ladder marker.
Hence, data from these 4 substrates were used to analyze all 403 clinical S. equi isolates. Of the 403 isolates tested, PFGE identified 179 as wild type and 224 as vaccine, whereas Biolog identified 252 as wild type and 151 as vaccine. Sensitivity, specificity, and accuracy of the Biolog identification were calculated using PFGE as a reference standard. The sensitivity, specificity, and accuracy of the Biolog identification were 69.4% (95% CI: 63.5–74.8), 97.4% (95% CI: 93.4–99.0), and 79.9% (95% CI: 75.7–83.5), respectively (Table 1).
It was later discovered that a single nucleotide polymorphism (SNP) found in the upstream regulatory region of the szp gene could be used to resolve the discrepancy between PFGE and Biolog results (Patterson et al., unpublished). Hence, in cases where PFGE and Biolog results disagreed, the upstream regulatory region of the szp gene was sequenced to use this SNP to identify the isolates as wild-type or vaccine strain. The szp SNP consisted of a deletion of a single guanine (G) residue in vaccine strains. The wild-type strains have two G residues at this position. The presence or absence of this single G residue located 85 bp upstream of the putative start codon of the szp gene was found to consistently correlate (r = 0.957, P < 0.001) with vaccine or wild-type pheno-types predicted by Biolog.
Comparison of Biolog and pulsed-field gel electrophoresis (PFGE) data obtained from 403 Streptococcus equi subspecies equi isolates. *
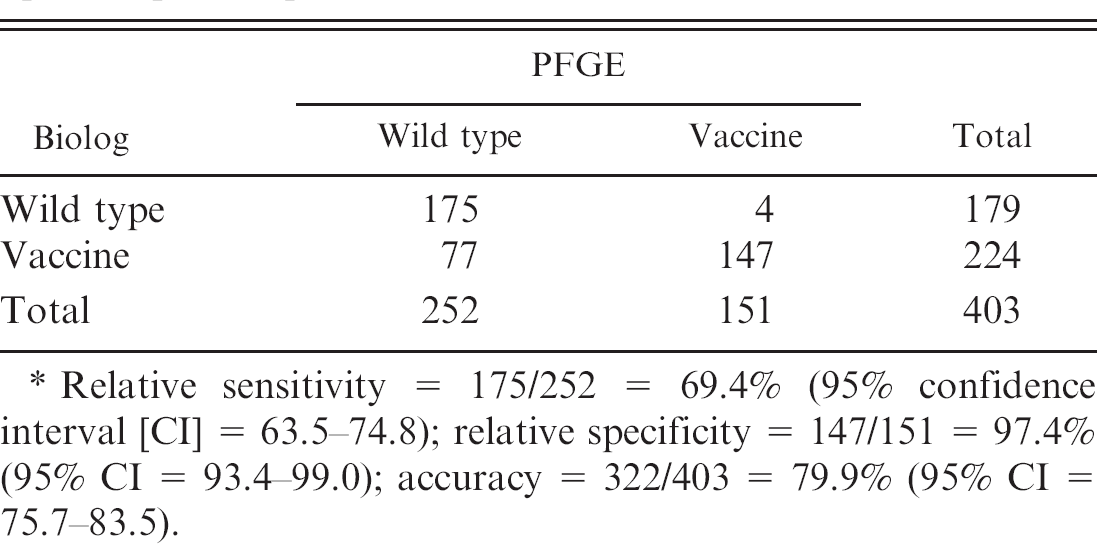

Relative sensitivity = 175/252 = 69.4% (95% confidence interval [CI] = 63.5–74.8); relative specificity = 147/151 = 97.4% (95% CI = 93.4–99.0); accuracy = 322/403 = 79.9% (95% CI = 75.7–83.5).
Because this single nucleotide deletion was characteristic of the vaccine strains, and did not occur in wild-type strains, including the vaccine progenitor strain used in the present study, szp sequencing was performed on 65 of the isolates that exhibited a wild-type Biolog profile and a vaccine PFGE profile. When more than 1 isolate obtained from the same animal exhibited similar Biolog profiles, only one of them was used for szp sequencing. Hierarchical cluster analysis and analysis of variance performed on szp-sequencing data and Biolog substrate utilization profiles obtained from these isolates demonstrated 98.5% agreement between Biolog and szp-sequencing profiles (Fig. 5). All of the isolates with Biolog vaccine profile, with the exception of 1, showed the single G deletion, whereas all the wild-type Biolog profiles exhibited 2 G residues at this location (Table 2).
Discussion
Beginning in 2002 an increase in clinical samples submitted to UIVDL from horses suspected of equine strangles was observed. These samples represented a wide geographic distribution within the United States with many of them originating from horses with a history of recent vaccination with MLV. a Streptococcus equi isolates obtained from horses with clinical strangles exhibited both mucoid and dry colony morphologies. Although the current vaccine consists of a nonencapsulated derivative of a live attenuated S. equi strain that primarily produces small dry colony morphology, upon serial passage these dry colonies may revert to mucoid phenotype. Both mucoid and dry colony morphologies have also been isolated from reconstituted vials of vaccine. 16 Thus colony morphology alone was not a reliable marker for differentiating wild-type from vaccine isolates. Because the nature of attenuation in the vaccine strain resulting from nitrosoguanidine treatment has not been genetically characterized, it became important to design biochemical or genetic methods that distinguish the strains. The current study describes and evaluates molecular and phenotypic approaches used to characterize S. equi isolates to distinguish wild-type from the vaccine strain and differentiate wild-type and vaccine strains from the wild-type progenitor of the vaccine. Initially, ERIC PCR, rep-PCR, and BOX PCR were tested to determine whether any of these methods or a combination of these methods would successfully differentiate the isolates. None of the DNA fingerprinting methods were able to discriminate between various wild-type strains or distinguish wild-type strains from vaccine strains. This may be a consequence of the highly homogeneous, clonal nature of the S. equi genome. 5 Similarly, the highly conserved 16S ribosomal DNA sequences resulted in ribotyping patterns that could not differentiate the various S. equi isolates. 4

Hierarchical cluster analysis of Biolog psicose and methyl pyruvate data from 65 Streptococcus equi subspecies equi isolates whose pulsed-field gel electrophoresis (PFGE) and Biolog profiles differed (Ward's linkage algorithm; squared Euclidean distances). Presence of the guanine deletion in the vaccine strains clusters with Biolog vaccine profile and absence of guanine deletion in wild-type strains clusters with wild-type Biolog profiles. The sem allele differences were unable to predict vaccine or wild type. PFGE/szp/Biolog classification: boldface = vaccine/wild type/wild type; italics = vaccine/vaccine/vaccine; boldface underlined = wild type/ wild type/inconclusive.
Comparison of Biolog and szp sequencing data obtained from 65 Streptococcus equi subspecies equi isolates that differed in their pulsed-field gel electrophoresis and Biolog profiles. *
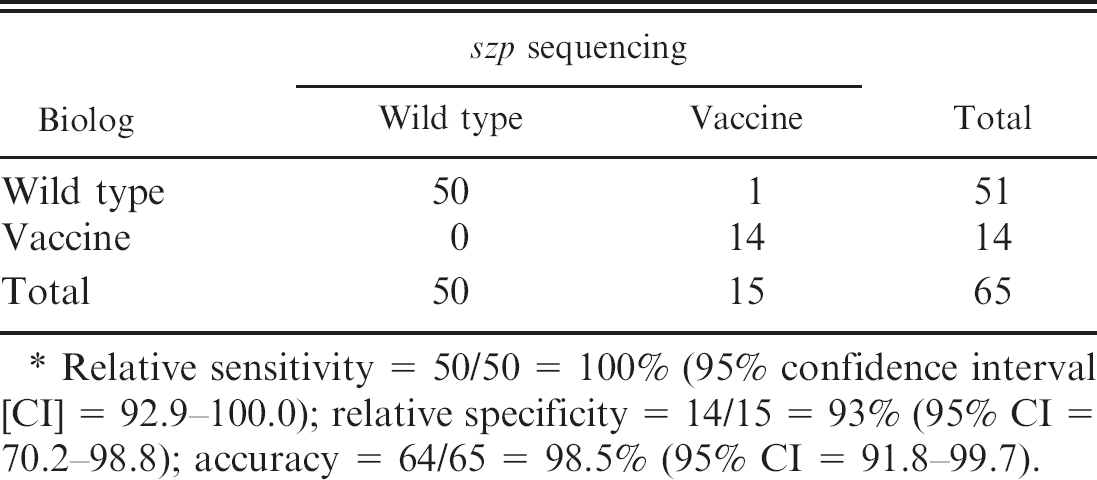
Relative sensitivity = 50/50 = 100% (95% confidence interval [CI] = 92.9–100.0); relative specificity = 14/15 = 93% (95% CI = 70.2–98.8); accuracy = 64/65 = 98.5% (95% CI = 91.8–99.7).
Large restriction fragment polymorphisms have been found to discriminate S. equi. 3,13 As a result, PFGE was used to address this issue, as PFGE was successfully used for genotyping S. equi 13 and has been considered the gold standard for genotypic analysis of many other bacteria. The PFGE data obtained from SmaI and NotI restriction digests demonstrated that there is a great deal of genetic variation among various wild-type isolates. This type of genetic variation in the form of small changes in the genomic DNA has been reported in a previous study 13 in which 7 restriction patterns were detected with NotI and 5 restriction patterns with SmaI among 20 field isolates obtained from the United States, Japan, and Ireland, and a single NotI restriction pattern for vaccine strain. In the present study, more field isolates were examined and a greater variety of restriction patterns were observed among wild-type isolates for both NotI and SmaI. Furthermore, 2 different NotI restriction patterns were observed for isolates obtained from vaccine vials (Fig. 4).
In addition, wild-type S. equi isolates and S. equi isolates obtained from vaccine vials had clearly distinguishable banding patterns. However, the wild-type progenitor strain could not be distinguished from the MLV strain by using PFGE alone as these 2 strains exhibited identical banding patterns with both NotI and SmaI. This is an important finding, as the wild-type progenitor of the MLV strain circulates in North America at low frequency and can be the cause of sporadic outbreaks of strangles.
To differentiate vaccine isolates from the wild-type progenitor strain, PFGE results were then compared with phenotypic data obtained from Biolog to look for correlation between the 2 methods. Data derived from the Biolog identification system showed good sensitivity and specificity when compared with PFGE as a gold standard. In addition, the Biolog data can distinguish wild-type strains, including the wild-type progenitor strain from isolates of the MLV, as evidenced by the excellent sensitivity and specificity of the Biolog assay when compared with szp sequencing. Isolates derived from vaccine vials, regardless of dry or mucoid phenotype, exhibited high absorbance values for utilization of psicose and methyl pyruvate on the Biolog. This was in sharp contrast to the wild-type progenitor, which showed very little absorbance for these substrates. In addition, the data indicated that increased usage of psicose and methyl pyruvate correlated perfectly with a SNP in the upstream region of szp, which also distinguishes the vaccine and wild-type strains. Together, with cost analysis shown in Table 3, findings suggest that the Biolog is a cost effective method for identification of the MLV strain in clinical isolates. However, neither Biolog nor szp sequencing can differentiate between wild-type field isolates for epidemiologic purposes.
Psicose is a rare low energy monosaccharide that yields only 0.3% of the metabolic energy of the equivalent amount of sucrose. 10 It remains unclear as to why the vaccine strain uses this sugar at an increased rate. It is likely that the increased utilization of this sugar may represent a defective carbohydrate metabolic pathway and potentially the dry pheno-type. This was supported by unpublished observations of this study that the vaccine strains grow slower than the wild-type progenitor strain and reach approximately 10% lower steady state concentrations. Further study of any mutations in vaccine strains that lead to this alteration in substrate utilization pattern would be beneficial both to characterize the vaccine strain as well as a rapid diagnostic test to differentiate the wild-type and vaccine strains.
Comparison of Biolog, pulsed-field gel electrophoresis (PFGE), and szp sequencing.
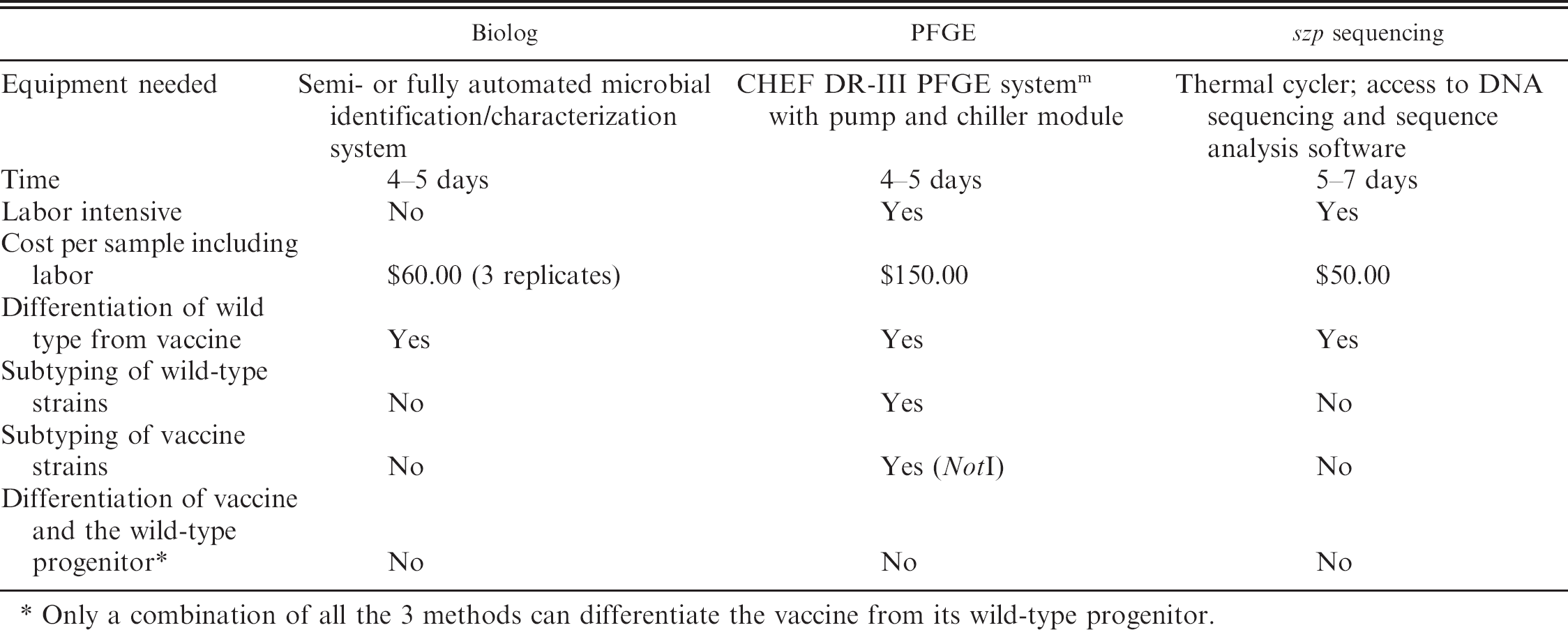
Only a combination of all the 3 methods can differentiate the vaccine from its wild-type progenitor.
Although the combination of PFGE and Biolog data showed excellent specificity, some S. equi strains identified as wild type by Biolog exhibited vaccine patterns by PFGE. These strains may represent vaccine revertants or isolates of the naturally circulating wild-type progenitor strain. After vaccination, a vaccine strain may revert to a virulent strain by compensatory mutations or by recombination with wild-type S. equi strains or possibly the closely related Streptococcus equi ssp. zooepidemicus. 7 These virulent reverted and/or recombined vaccine strains might be identified as wild type by Biolog due to their decreased utilization of psicose and methyl pyruvate, but no changes would be observed in their PFGE profiles unless a change occurs at sites recognized by SmaI or NotI restriction enzymes. Alternatively, deletions, insertions, or SNPs disrupting enzyme recognition sites may alter the PFGE pattern without disrupting the Biolog profile. Thus the disparity observed between PFGE and Biolog data was likely due to PFGE not differentiating vaccine strains from its wild-type progenitor strain. In other cases, mixed PFGE patterns with vaccine-like genotypes in one enzyme but wild-type–like patterns in another enzyme were observed. Some of these strains exhibited a vaccine-like Biolog profile and indeed the vaccine-associated SNP in the upstream region of szp was present.
Pulsed-field gel electrophoresis, Biolog, or szp sequencing can differentiate wild-type S. equi strains from vaccine strains; however, each method has important limitations (Table 3). Both Biolog and szp sequencing sensitively and specifically identify the MLV strain isolated in clinical cases. The PFGE method provides genotypic data that can be used to track outbreaks on a farm or between farms. None of the 3 methods can distinguish an isolate as a wild-type progenitor of MLV strain when used alone. This is an important limitation as the wild-type progenitor currently is in circulation in North America causing sporadic outbreaks of clinical strangles. Hence, only a combination of PFGE, Biolog, and szp sequencing can differentiate the MLV strain from its wild-type progenitor strain. In addition, such a combination approach may prove useful in monitoring vaccine strains for reversions that could potentially restore virulence. Thus, the current study illustrates the usefulness of genotypic and phenotypic approaches to differentiating S. equi strains and underscores the importance of incorporating a multipronged approach to answer epidemiologic and diagnostic questions.
Acknowledgements. The authors wish to thank Amy Stevenson, Tina Croix, Kristin Ellis, and Gregg Clabaugh of the University of Illinois Veterinary Diagnostic Laboratory for their assistance with S. equi cultures and Biolog assays.
Footnotes
a.
Pinnacle I.N.™, Fort Dodge Animal Health, Fort Dodge, IA.
b.
Remel Inc., Lenexa, KS.
c.
Pathodox™, Remel Inc., Lenexa, KS.
d.
BiologMicrolog™ System, Release 4.2, Biolog Inc., Hayward, CA.
e.
DuPont Qualicon, Wilmington, DE.
f.
Phusion®, Finnzymes Inc., Woburn, MA.
g.
QIAamp®, Qiagen Inc., Valencia, CA.
h.
Invitrogen Corp., Carlsbad, CA.
i.
Integrated DNA Technologies Inc., Coralville, IA.
j.
Fisher Scientific Co., Pittsburg, PA.
k.
Bacterial Barcodes Inc., Houston, TX.
l.
Sigma-Aldrich, St. Louis, MO.
m.
CHEF-DR® III, Bio-Rad Laboratories, Hercules, CA.
n.
DNASIS®, Hitachi Software Engineering America Ltd., South San Francisco, CA.
o.
STATISTICA, StatSoft Inc., Tulsa, OK.
