Abstract
Rhodococcus equi is an important pathogen of foals aged 1–6 months. Evidence exists that foals are exposed to a wide diversity of R. equi strains in their environment. However, limited data are available regarding the extent to which genotypic variation exists among isolates infecting individual foals. Therefore, electrophoresis of repetitive sequence–based polymerase chain reaction (rep-PCR) amplicons in an automated microfluidics chip format was used to genotype 9 virulent R. equi isolates obtained from distinct anatomic locations in a single foal. Four of the isolates were obtained from different regions of the lung, and 5 were from abscessed intra-abdominal lymph nodes (LNs). Six distinct genotypes were identified among the 9 isolates. None of the pulmonary isolates was identical; however, a pulmonary isolate was found to be identical to an isolate recovered from a small intestinal LN, and another pulmonary isolate was identical to an isolate from a mesenteric LN. These results indicate that foals can be infected with multiple strains of virulent R. equi. Furthermore, identical strains can be found in multiple, remote anatomic locations in an infected foal, and this can occur for >1 strain in the same foal. The automated system used in the current study provided a rapid, reproducible, and discriminating method for typing R. equi isolates.
Rhodococcus equi (order Actinomycetales, suborder Corynebacterineae, family Nocardiaceae) is an important pathogen of 1- to 6-month-old foals that causes a severe, purulent bronchopneumonia. 7 Often accompanying the pneumonia are extrapulmonary disorders, the most common being pyogranulomatous enterocolitis and intraabdominal lymph node abscesses. 7,12 Foals infected with R. equi can present with bronchopneumonia exclusively, extrapulmonary disorders exclusively, or both. 7 Molecular epidemiologic studies of R. equi have described considerable heterogeneity in the genotype of clinical and fecal isolates obtained from various sources such as farms and diagnostic laboratories. 1,10,14,16,18 However, detailed information about genotypic variation among multiple isolates from an individual foal is lacking.
Various methods for genotyping strains of bacteria have been developed, including repetitive sequence–based polymerase chain reaction (rep-PCR), 9,11 which uses oligonucleotide primers that target strain-specific noncoding repetitive sequences interspersed throughout the bacterial genome. 6,9,11 Repetitive sequence–based PCR has been used widely to genotype numerous bacterial species and has recently been adapted to an automated format. 6 This automated system a has been demonstrated to have a high degree of repeatability and discriminatory power in a wide array of bacterial genera; however, to the authors' knowledge, no published data exist using this system to evaluate isolates of R. equi. The objective of the current study was to describe the extent to which multiple pulmonary and extrapulmonary isolates from an individual foal were the same strain (i.e., were clonal) or whether multiple strains (i.e., polyclonal or mixed infection) can be found in a given host, using this automated rep-PCR genotyping system.
An approximately 70-day-old Quarter Horse foal was referred to the Large Animal Hospital at Texas A&M University (College Station, Texas) on March 10, 2006, with complaints of diarrhea, increased respiratory effort and rate, and fever. Thoracic radiography revealed multiple cavitary lesions and a diffuse bronchointerstitial pattern consistent with pneumonia caused by R. equi. A sample of tracheobronchial fluid yielded a pure culture of R. equi, and cytologic examination of the fluid was consistent with septic, suppurative inflammation with multiple pleomorphic bacteria identified both extracellularly and intracellularly. Ultrasonographic examination of the foal's abdomen revealed multiple nodular masses of heterogeneous echoic pattern that were interpreted as enlarged lymph nodes presumed to be infected with R. equi. After limited clinical response following a week of antimicrobial therapy, the foal was euthanized at the owner's request. Postmortem examination of the foal revealed multiple pyogranulomas in the thoracic and abdominal cavities. All regions of the lungs were affected, and 1 large lesion was filled with granulomatous material and gas (emphysematous bulla). Lymphadenopathy and lymph node abscessation in the abdominal cavity were extensive. Culture of these lesions confirmed disseminated R. equi pneumonia, lymphadenitis, and ulcerative colitis. Microscopic examination of tissues was consistent with pyogranulomatous pneumonia, enteritis, colitis, and multifocal peritonitis.
At the time of necropsy, samples were collected by 1 of the authors (KC) using aseptic technique from 9 sites with pyogranulomatous lesions: 1) left cranial lung, 2) small intestinal lymph node, 3) right caudodorsal lung, 4) left caudodorsal lung, 5) mesenteric lymph node, 6) a cecal lymph node, 7) a small intestinal lymph node, 8) a colonic lymph node, and 9) right craniodorsal lung. Swabs of these sites were streaked onto culture plates containing a modified nalidixic acid–novobiocin–actidione (i.e., cycloheximide)–potassium tellurite (NANAT) medium and incubated at 37°C for 48 hr; plates from each site yielded a pure culture of R. equi. A single colony from each plate was expanded on a modified NANAT plate, and then 3–5 colonies were transferred to tubes containing beads b for cryopreservation according to the manufacturer's recommendations and frozen at −80°C for subsequent genotyping. To confirm that all isolates were virulent R. equi, a multiplex PCR technique was performed on DNA extracted from each isolate using species-specific primers for R. equi based on the sequence of the choE (cholesterol oxidase) gene and primers for the virulence plasmid-borne vapA gene. 5 In addition to the 9 clinical isolates, strain ATCC (American Type Culture Collection) 33701 was included as a reference (positive control).
A bead of each isolate was streaked onto brain–heart infusion agar plates and incubated at 37°C for 48 hr. The plates were then visually inspected to ensure only the presence of R. equi. Next, DNA was extracted from each isolate using a 10-μ1 loop of plated culture and a commercial DNA isolation kit c following the manufacturer's instructions. All DNA was diluted to a concentration of between 25 and 50 ng/μ1, per the manufacturer's instructions.
The extracted DNA was amplified using a commercial kit d according to the manufacturer's instructions. The product insert for this kit indicates that it is recommended for members of the genera Rhodococcus and Nocardia. Briefly, 18 μ1 rep-PCR master mix 1, 2 μ1 of primer mix AA, 2.5 μ1 10X PCR buffer, e and 0.5 μ1 each of dimethy1 sulfoxide and DNA polymerase f were combined for a total of 23.5 μ1 per reaction for preparation of the master mix. Next, 23 μ1 of the master mix was added to 2 μ1 of genomic DNA (either from sample isolates of R. equi, the reference strain 33701, or the manufacturer's proprietary DNA used as a positive control for PCR amplification) for a total of 25 μ1 per reaction mixture for rep-PCR. A negative control also was included for each rep-PCR assay in which 2 μ1 of molecular-grade water was added to 23 μ1 of the master mix. The thermal cycling parameters were as follows: initial denaturation of 94°C for 2 min; 35 cycles of denaturation at 94°C for 30 sec, annealing at 65°C for 30 sec, and extension at 70°C for 90 sec; and a final extension at 70°C for 3 min.
The DNA amplicons were separated using electrophoresis and a microfluidics chip device g in a bioanalyzer. h Microfluidics technology uses a network of channels that are etched onto a polymer chip to move picoliter volumes of DNA along with intercalating dye through a matrix for separation and across a laser beam for detection via electrokinetic forces.
Analysis was performed with the appropriate software (version 3.3). a Similarity matrices were determined using the Kullback–Leibler correlation coefficient, and the unweighted-pair group method with arithmetic mean was used to create a dendrogram. A computer-generated virtual gel image based on results of the microfluidics was also produced using the software to better visualize rep-PCR patterns. a Isolates were considered genotypically distinct if there was <95% similarity between isolates from the similarity matrix and if there were at least 2 differences in rep-PCR bands. 17 Isolates were considered closely related (i.e., the same strain) if they were ≥95% similar and differed by only 1 band and were considered identical when similarity was >98% and all bands were the same.
The automated rep-PCR system a used in this study was highly discriminating: >12 bands were generated for all isolates, and multiple genotypes were readily distinguished. All steps from growing the bacteria and extracting DNA through DNA amplification and microfluidic genotyping were repeated for all isolates on a different day, and results were identical for each isolate, including the reference strain 33701 included as a positive control, substantiating reported repeatability of the technique. 6 The negative control yielded no bands, as expected.
Nine isolates (4 pulmonary and 5 extrapulmonary) from a single foal were used, and it was determined that 6 distinct genotypes were present (Fig. 1). Two pairs of isolates were identical, and each of the 2 pairs included a pulmonary (left cranial lung [strain A] or right caudodorsal lung [strain E]) and an intra-abdominal isolate (small intestinal lymph node [strain A] or mesenteric 1ymph node [strain E]). An isolate recovered from a small colon 1ymph node was the same strain but not identical to the isolates from the left cranial lung and small colon 1ymph node (Fig. 1, strain A). The reference strain ATCC 33701 also appeared to be grouped as strain A. An isolate from a colonic 1ymph node was similar to but differed by more than a single band from the isolates representing strain E (Fig. 1). All 4 of the pulmonary isolates were found to be genotypically distinct, as were 4 of the 5 intra-abdominal 1ymph node isolates. Results for the proprietary DNA used as a positive control were not included in Figure 1; however, it fell outside of the R. equi isolates when included in a dendrogram (i.e., below the isolate from the right craniodorsal lung).
Foals infected with R. equi often have been assumed to be infected with a single strain of the bacterium, although evidence exists that foals may be infected with more than 1 strain. 10,16 Unfortunately, the number of isolates from individual foals and the description of genotypic variation of these isolates are quite limited either because the topic of polyclonal infection was not the principal focus of a previous report 10 or because the ribotyping method used was not highly discriminating (i.e., ribotyping generated only 3–4 bands). 16 In the current study, 6 distinct genotypes were found within a single foal using rep-PCR, indicating that foals can be infected concurrently with a variety of virulent R. equi strains. Similar findings have been reported for isolates of Mycobacterium tuberculosis from individual human tuberculosis patients. 13,15 Depending on the frequency with which polyclonal infections occur among foals with R. equi, there are several important implications of the findings of the present study. Most commonly, R. equi infections are treated with macrolide antibiotics, namely, erythromycin, azithromycin, and clarithromycin. 3 Because isolates of R. equi can vary in their susceptibility to macrolides, 4 results of antimicrobial susceptibility from a given isolate obtained by tracheobronchial aspiration may not adequately or accurately reflect the susceptibility of all strains with which a foal may be infected. Thus, distinct strains of R. equi infecting a single foal that have different antimicrobial susceptibility patterns could have a significant impact on the successful outcome of treatment or even survival of the foal. Genotypic variation, however, does not necessarily infer variation in antimicrobial susceptibility, as reflected by the finding of variations in ribotypes of rifampin-resistant isolates of R. equi from an individual foal. 16 Infection with multiple genotypic strains of R. equi also can confuse or limit the application of molecular epidemiologic methods to link infections. The finding of multiple strains in an individual foal also further weakens the anecdotal concerns about the existence of a given farm having a specific strain that causes disease among resident foals.
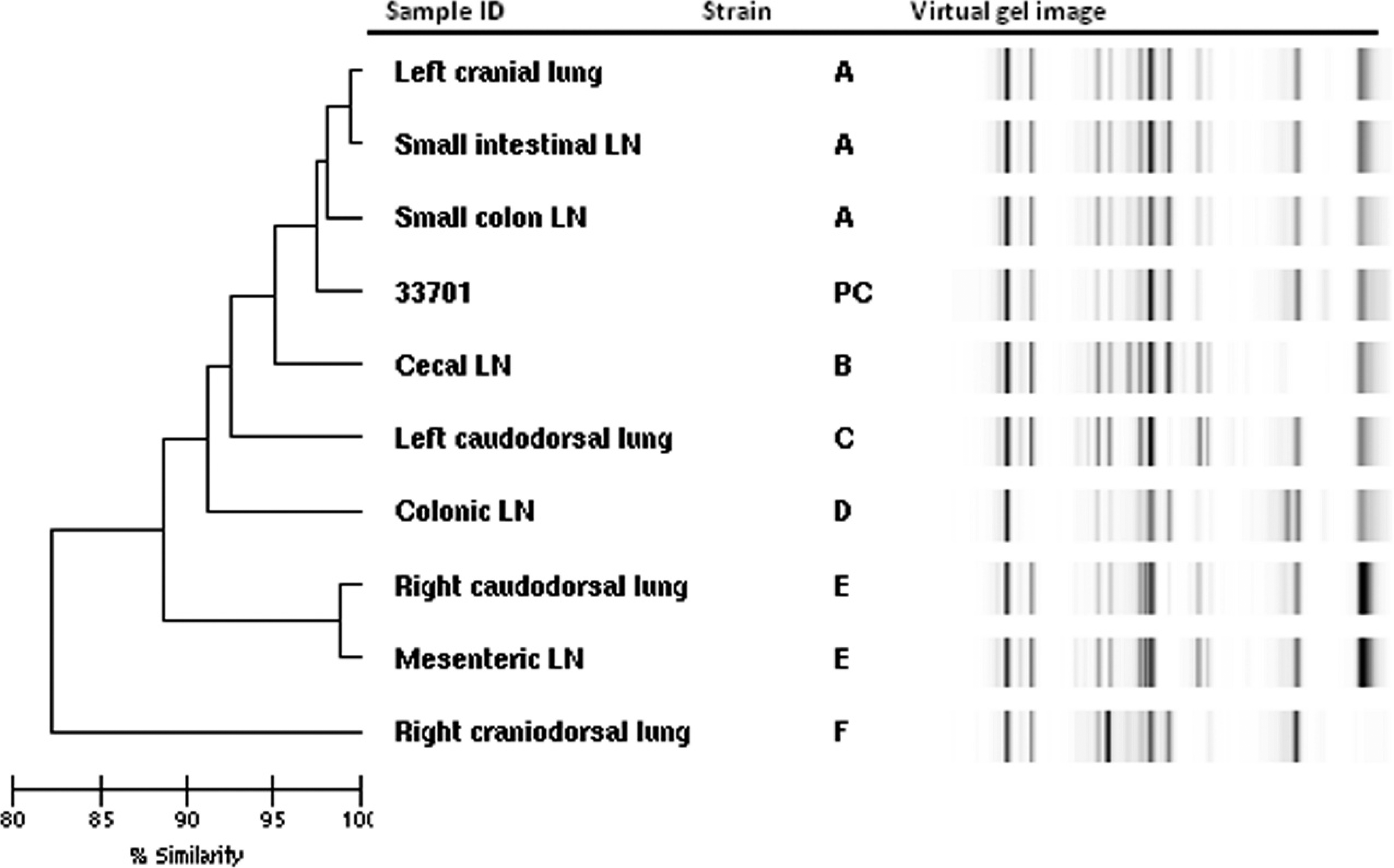
Dendrogram and virtual gel image demonstrating comparison of 9 Rhodococcus equi isolates from 9 anatomically distinct sites obtained from a single foal. Strains are identified by letters; PC represents the positive control reference strain ATCC (American Type Culture Collection) 33701. Six distinct strains were identified (see text for details); % similarity was determined using the Kullback–Leibler correlation coefficient. LN = 1ymph node.
There are a number of possible explanations for finding multiple strains of R. equi in a single foal. It is generally considered that foals become infected from inhaling infectious agents in their environment. 7,8 Evidence exists that there is considerable variation in the molecular fingerprints of genomic DNA from isolates of R. equi obtained from horses and foals and their environment. 1,10 , 14,16,18 The data suggest that essentially any strain of R. equi that possesses the virulence plasmid is capable of causing disease in susceptible foals. Thus, it is possible that a variety of virulence-plasmid–bearing isolates from its environment might simultaneously infect a given foal. Alternatively, it is possible that some of the genotypic diversity among isolates reflects genetic changes that occurred postinfection. Strain type A contained 3 isolates, including one that was >95% similar and differed by only 1 band from other strain A isolates, and strain C was >95% similar to members of strain E (Fig. 1). These slight genotypic differences could have resulted from either adaptive genetic changes or random mutations occurring after infection of the host. 2 However, the finding of several more distinct genotypes (Fig. 1) is consistent with polyclonal R. equi infection and suggests that polyclonal infections may be more common than postinfective genetic variations. Finally, it is also possible that multiple recurrent infections (rather than a single event of infection with multiple strains) can occur in foals. Further studies regarding the frequency with which polyclonal infections occur in foals are needed.
The finding of identical strains at remote anatomic locations, such as the left cranial lung isolate being identical to a small intestinal lymph node isolate, indicates that clonal dissemination can occur. The route of dissemination of R. equi infection in the host is unknown. It is possible that dissemination occurs hematogenously or via lymphatic circulation. Alternatively, it is possible that bacteria from pulmonary lesions reach the intestinal tract following expectoration and swallowing. It is also possible, but less likely, that regurgitated R. equi are aspirated and infect the lower respiratory tract or that simultaneous swallowing and inhalation of genotypically identical R. equi strains occurs.
The present study had several limitations. First, polyclonal infection was investigated and described for only a single foal. The extent to which polyclonal or mixed infection occurs in other foals is not known. Second, only a single isolate from each of the 9 anatomical locations sampled was available for genotyping. Based on the finding of genotypic heterogeneity between remote and distinct R. equi isolates in the current study, the possibility that all R. equi within a single lesion are clonal is questionable. Indeed, variation of isolates from a single culture has previously been described in human patients infected with Mycobacterium tuberculosis. 13 It is possible that the current study has underestimated the extent of diversity of isolates because other genotyping methods such as pulsed-field gel electrophoresis (PFGE) may be more discriminating than rep-PCR. For example, PFGE was more discriminating than rep-PCR for typing Staphylococcus aureus. 17 This limitation would seem to underscore the principal result in the current study that a variety of strains was isolated from the foal reported herein. Finally, genotyping of R. equi using the methods described requires access to the proper bioanalyzer and computer software. This limitation is likely offset by the advantages of convenience, reliability, and rapidity of the system, such that it can readily be established by diagnostic or reference laboratories to provide genotyping results that can be compared among laboratories.
In summary, the present report describes genotypic variation among isolates of R. equi from a single foal suggesting polyclonal infection, as well as identification of genotypically identical strains isolated from remote locations in the same foal. The automated system used was highly discriminating and provided results rapidly. Moreover, this system has been reported to be reliable for comparing results from different laboratories; thus, this method could serve as a method for comparing genotyping results for isolates of R. equi among laboratories. However, further testing including comparison of the newly developed automated rep-PCR method with other methods such as pulsed-field gel electrophoresis is needed to establish the clinical and epidemiological value of this rep-PCR method.
Acknowledgements
The authors wish to thank Dr. Suresh Pillai for the use of the DiversiLab software and Agilent bioanalyzer and Dr. Mei Liu for technical assistance. This work was funded by the Link Equine Research Endowment, Texas A&M University, and by the National Institutes of Health Summer Veterinary Medical Scientist Training Program.
Footnotes
a.
DiversiLab™ system, bioMérieux Inc., Durham, NC.
b.
CryiCare Bacteria Preservers, Key Scientific Products Inc., Stamford, TX.
c.
UltraClean™ Microbial DNA Isolation Kit, MO BIO Laboratories Inc., Carlsbad, CA.
d.
Bifidobacterium DNA fingerprinting kit, bioMérieux Inc., Durham, NC.
e.
GeneAmp® 10X PCR buffer, Applied Biosystems Inc., Foster City, CA.
f.
AmpliTaq® DNA polymerase, Applied Biosystems Inc., Foster City, CA.
g.
LabChip® device, Caliper Technologies Inc., Hopkinton, MA.
h.
Agilent 2100 Bioanalyzer, Agilent Technologies Inc., Santa Clara, CA.
