Abstract
Pasteurella multocida is the causative agent of fowl cholera in domesticated and wild birds. The disease outcome is affected by various host- and pathogen-specific determinants. Several putative virulence factors have been proposed to play a key role in this interaction, including the ptfA gene, the products of which assemble to form type 4 fimbriae on the bacterial surface. One way to understand more precisely how ptfA contributes to pathogenesis is to gather molecular features of this gene in circulating avian P. multocida strains. Therefore, molecular characterization of the ptfA gene of P. multocida strains isolated from domestic poultry was performed using the combination of nucleotide sequence analysis and a newly developed allele-specific polymerase chain reaction assay. Two major ptfA alleles were identified among 31 strains, representing various serogroups and somatic serotypes. It was noteworthy that allele specificity and case severity of a subset of strains correlated with the available gross pathology data. Therefore, the acquisition of comprehensive clinical and epidemiological data together with molecular characteristics of individual strains will help to design and implement adequate preventive and intervention strategies.
Fow1 cholera (avian pasteurellosis) is a contagious disease that affects over 100 avian species and leads to significant economic losses in domestic poultry worldwide. 2 The course of infection can be either acute or chronic. In acute infections, the clinical signs are often restricted to the observation of sudden death. In chronic infections, the spectrum of clinical signs is mainly localized to the limb joints, the comb, the oviduct, the respiratory tract, the wattles, and the subcutaneous tissue of the head. 13
Fow1 cholera is caused by Pasteurella multocida, the strains of which can be classified into 5 capsular serogroups (A, B, D, E, and F). 3,14 Among avian strains of P. multocida, 4 capsular serogroups are represented: A, B, D, and F. Serogroup A strains cause the majority of fow1 cholera cases, whereas serogroups B, D, and F strains are less frequently associated with the disease. 15 To date, 16 somatic serotypes (1–16) 9 have been described in P. multocida, of which serotypes 1, 3, and 4 have been most commonly seen in birds. 4
The molecular basis of pathogenesis and protective immunity against P. multocida infections is not fully understood. Putative virulence determinants such as outer membrane proteins, proteins involved in iron uptake and acquisition, sialidases, toxins, and various adhesins may play a part in these mechanisms. 8 Adhesins have a crucial role in mediating colonization and invasion of the host. Thus, their presence on the bacterial surfaces is usually correlated with virulence. 10 Adhesins have been targeted in recent virulence genotyping studies, 7,21 which demonstrated that among fimA (fimbriae), hsf-1,2 (autotransporter adhesins), pfhA (filamentous hemagglutinin), tad (nonspecific tight adherence protein), and ptfA (subunit of type 4 fimbriae), only fimA, hsf-2, and ptfA are present in all or virtually all pathogenic isolates of P. multocida. These data indicate a possible role of the ptfA gene in the virulence of P. multocida in various hosts and disease conditions.
Genetic characterization of the ptfA gene is limited to a handful of strains isolated in parts of Asia and Australia, 5,20 showing the existence of 2 major sequence variants within the avian group of P. multocida. A better understanding of the importance of allelic polymorphism of the ptfA gene and its potential use in the assessment of virulence of field isolates in epidemiologic monitoring studies requires additional sequence information on strains isolated in other geographic areas. Exploring allelic polymorphism may also help to determine the key factors responsible for the different clinical outcomes of P. multocida infections in poultry. Moreover, ptfA might be an attractive component of next generation P. multocida vaccines. 1,6 Understanding the genetic variability and the occurrence of various alleles of the ptfA gene is likely to assist in the formulation of efficacious vaccines that elicit cross-protection against this serologically heterogeneous pathogen group and will thus facilitate the implementation of adequate preventive strategies against P. multocida in poultry flocks.
In order to gather relevant information on ptfA gene diversity, the ptfA gene in a subset of Hungarian avian P. multocida strains was sequenced. The phenotypic and genotypic characterization of these strains by enterobacterial repetitive intergenic consensus sequence–based (ERIC) polymerase chain reaction (PCR) has been described previously. 18 Some of these phenotypic and genotypic features are listed in Table 1. Twenty strains, belonging to various serogroups and somatic serotypes, were selected for the sequencing study of the ptfA gene. The strains represented different ERIC-PCR clusters that showed considerable correlation with their geographical origin and the host species. 18
Characteristics of Pasteurella multocida strains used in the current study.*
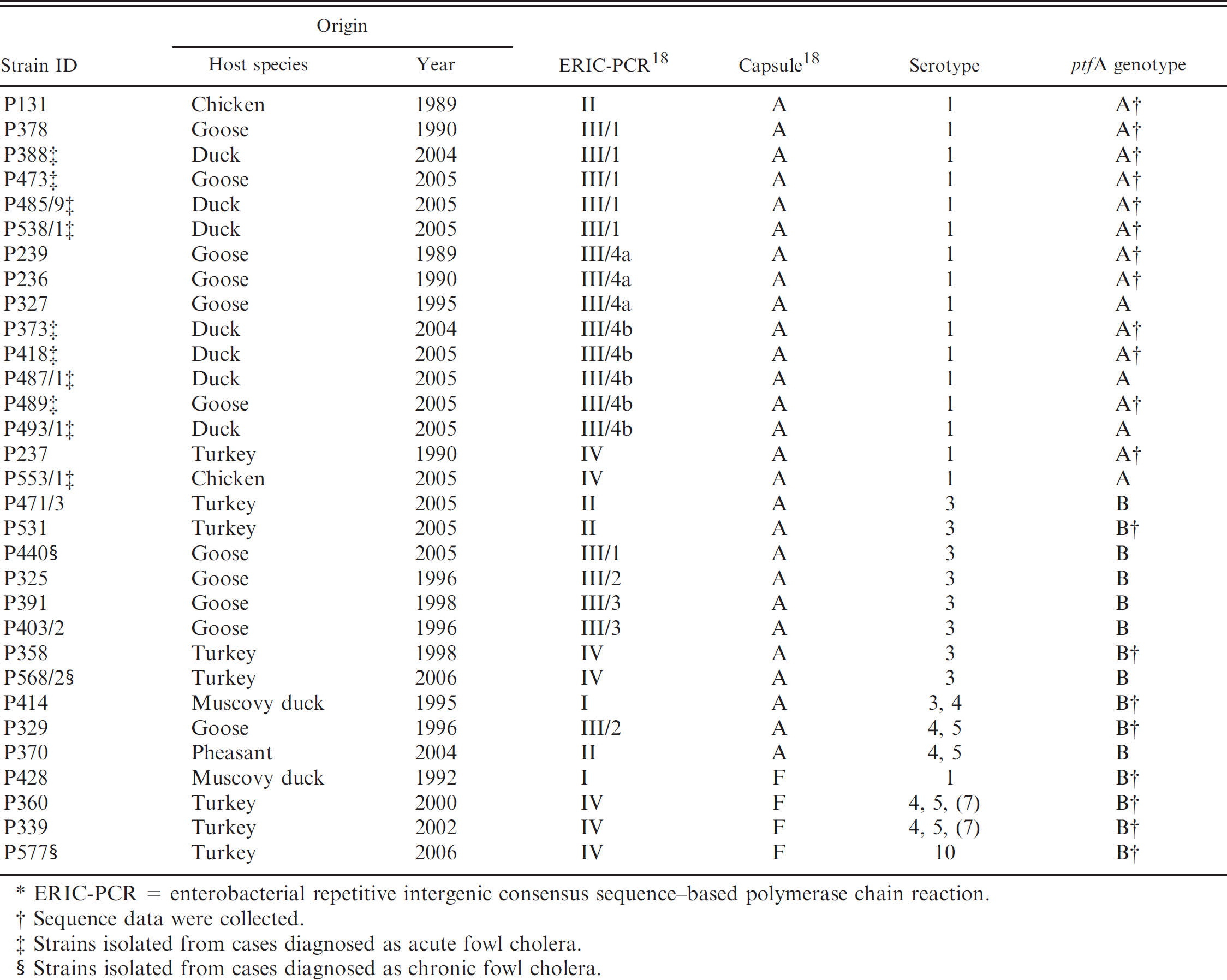
ERIC-PCR = enterobacterial repetitive intergenic consensus sequence–based polymerase chain reaction.
Sequence data were collected.
Strains isolated from cases diagnosed as acute fow1 cholera.
Strains isolated from cases diagnosed as chronic fow1 cholera.
All strains were cultivated on 5% sheep blood agar at 37°C for 24 hr. Genomic DNA was extracted using the Chelex method. Briefly, a bacterial colony was suspended in 1 m1 of autoclaved water at 0.5 McFarland standard (1–1.5 × 108 colony-forming units) and then pelleted for 1 min at 16,000 × g in a microfuge. After the removal of the supernatant, 50 μ1 of 5% Chelex 100 resin a suspension in water was added and mixed thoroughly. This step was followed by incubations at 5°6C for 30 min and then at 100°C for 8 min. After centrifugation (3 min at 16,000 × g), the supernatant was stored at −20°C. Five microliters of these DNA preparations were used in a 25-μ1 PCR reaction. Oligonucleotide primers flanking the ptfA gene were designed based on the P. multocida Pm70 (A:3) complete genome 12 that contained ampD-PM0084 open reading frame (ORF; ptfA homologous region)–hofB genes (GenBank accession no. AE006044). The ptfA forward primer was 5′-AATGCCAGTCCACTCGTTGT-3′ (nucleotides [nt] 978–998), and ptfA reverse primer was 5′-CGTCCTGAGGCAAGCGTGTT-3′ (nt 2043–2063). A reaction volume of 25 °1 containing template DNA, 12.5 pmol of each primer in 1X PCR buffer, b 2.5 mM MgC12, b 200 nM deoxyribonucleotide triphosphate (dNTP) mix, b and 1.25 U DNA polymerase b was prepared. DNA polymerase was prepared and subjected to 30 cycles (denaturation: 95°C, 30 sec; annealing: 50°C, 30 sec; extension: 72°C, 1 min). An initial denaturation step (95°C, 5 min) and a final extension step (72°C, 10 min) were also employed. The 1,085–base pair (bp)-long amplicons were purifiedc and subjected to direct sequencing using a commercial kit. c Dye-labeled products were run on a genetic analyzer capillary sequencer d at the DNA sequencing laboratory of the Biological Research Center (Szeged, Hungary).
Nucleotide sequences of the ptfA gene of the P. multocida strains were aligned with the few reference sequences available in GenBank (http://www.ncbi.nlm.nih.gov/) using the MegAlign program in the Lasergene package e and using the ClustalW multiple alignment algorithm. By visual inspection, 2 major sequence variants were identified, referred to as type A (designated for PBA815 strains A:1; GenBank accession no. AF154834) and type B (designated for Pm70 strains A:3; GenBank accession no. AE006044). Consistent with previous findings, 16 the 5′ end of the ptfA gene was highly conserved (not shown), while there were marked differences in the 3′ end of the gene, 5 including 60 nucleotide substitutions, which demarcated these 2 alleles (Fig. 1)
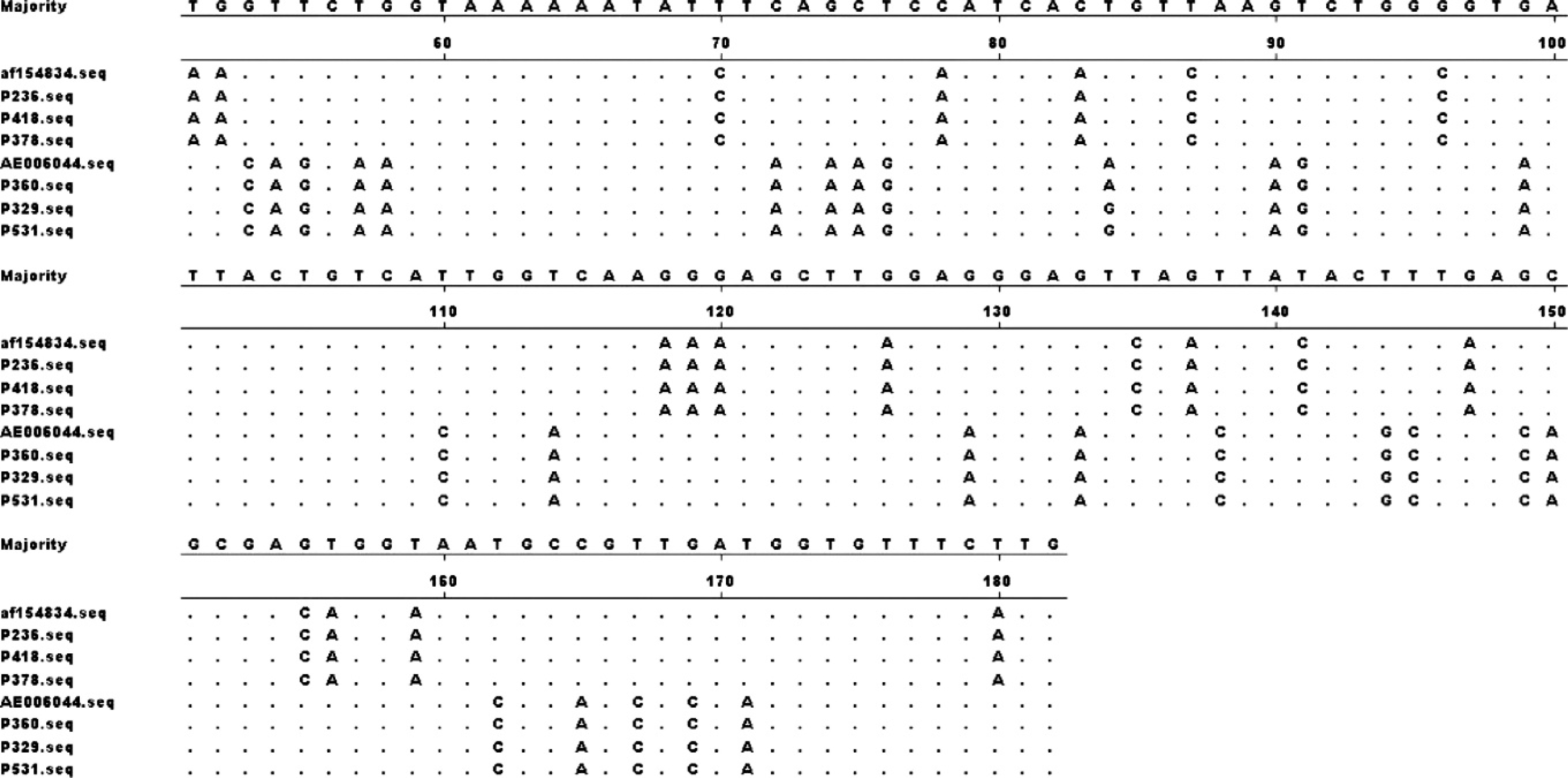
Nucleotide (nt) sequence alignment of the variable region (nt 200–380) of the ptfA gene from representative avian isolates with that of Pasteurella multocida A:1 (AF154834) and A:3 (AE006044) strains. The serotypes of avian strains were A:1 (P236, P418, P378), A:4,5 (P329), A:3 (P531), and F:4,5,(7) (P360).
The observed low genetic heterogeneity of the ptfA gene indicated that a simpler and less time-consuming allele typing method could potentially be developed as a substitute for nt sequencing. Such a method might be used in epidemiological investigations aimed at typing of individual virulence genes. Thus, an allele-specific PCR assay based on the available sequence data of the Hungarian strains and the reference P. multocida strains was developed. The primers were designed to the variable region of the ptfA gene. Primer sequences were as follows: 5′-TATCGTGCCGAGGTAGAAC-3′ (forward; nt 163–182 on both types of ptfA gene), 5′-CTTGACATTGATTGAGCTGAG-3′ (type A–specific reverse; nt 268–289 on ptfA type A allele; AF154834), and 5′-AAGAAACACCTTGAGCTG-3′ (type B–specific reverse; nt 360–378 on ptfA type B allele; ptfA homologous region [PM0084 ORF] of AE006044). A PCR reaction volume of 25 μ1 containing template DNA, 10 pmol of type A primer and 15 pmol of forward primer and type B primer in 1X PCR buffer f with 2 mM MgC12, f 200 nM dNTP mix, f and 1 U DNA polymerase f was prepared and subjected to 20 cycles of reaction with denaturation (95°C, 30 sec), annealing (60°C, 30 sec), and extension (72°C, 20 sec). The initial denaturation was at 95°C for 5 min, and the final extension was at 72°C for 10 min using a commercial thermocycler. g The expected lengths of amplicons were a 126-bp fragment for type A and a 217-bp fragment for type B strains (Fig. 2)
All 20 sequenced strains and 11 additional strains for which no sequence information was retrieved were successfully typed by the allele-specific PCR (Table. 1). None of the strains remained untypeable in the newly developed assay.
Recent studies 11,17,19 documenting the prevalence of P. multocida in different avian species indicate that serotype A:1 strains are responsible for the majority of economic losses. Previously, the authors 18 demonstrated that in Hungary A:1 strains mainly affect geese and ducks, while strains other than A:1 are relatively more common in other avian species. In the present study, a strong correlation was seen between type A of ptfA and serotype A:1 as well as between type B and serotype non-A:1 strains. Unfortunately, with this historical strain collection, limited data concerning the clinical manifestations of the particular P. multocida infections were available (Table. 1). Thus, it was not possible to make any specific conclusions regarding any strain-specific pathogenic potential. However, the observation that all strains isolated from birds with acute fowl cholera proved to be ptfA type A, while strains from cases of chronic fow1 cholera were classified as ptfA type B, seems to be of interest and deserves further investigation. Systematic collection of adequate clinical data would be important to aiding an understanding of the differences, if any, in virulence associated with allelic variants of the ptfA gene.
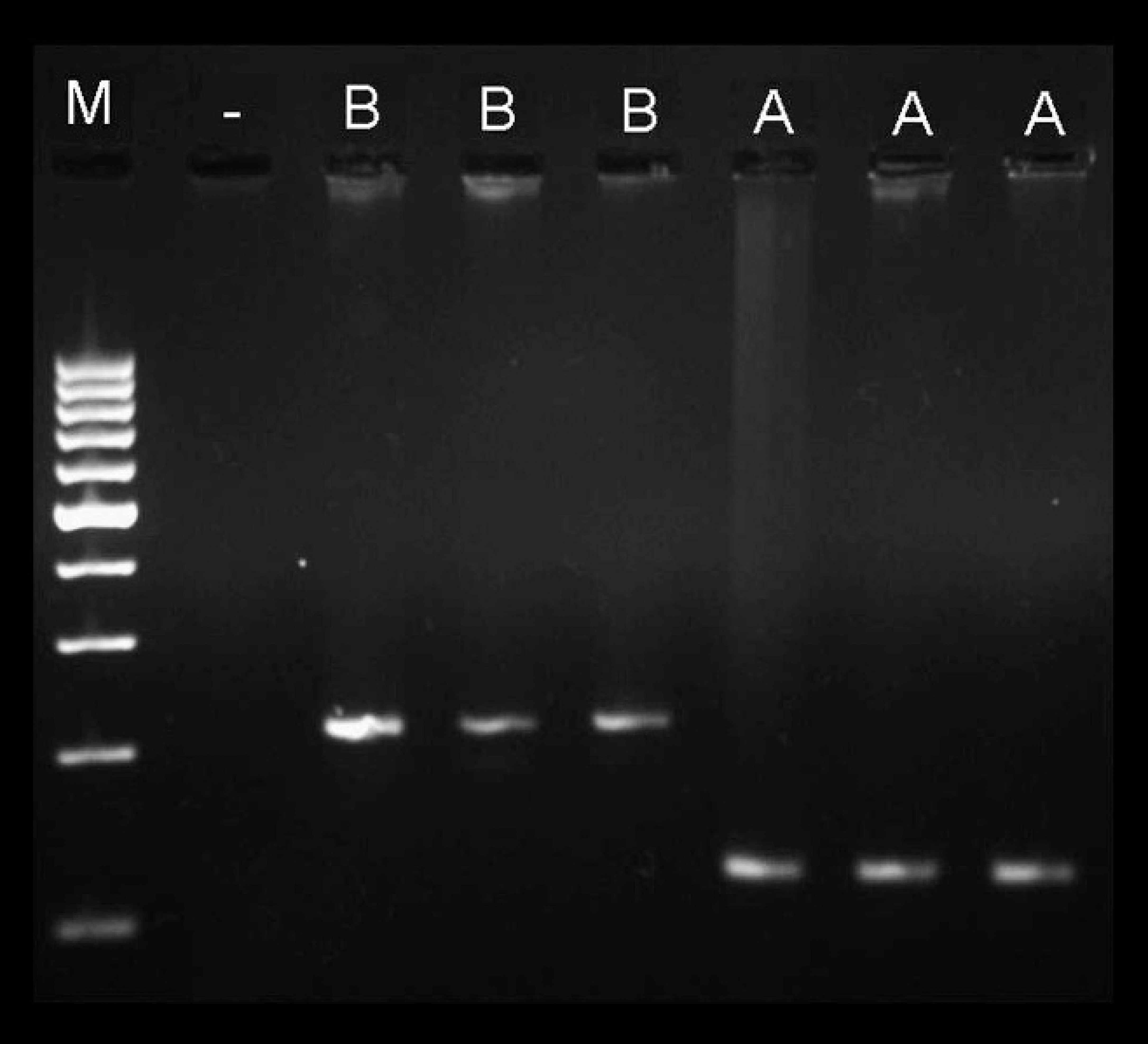
Amplification products from the allele-specific polymerase chain reaction assay. Three non-A:1 (lanes 1–3: P531, P358, and P329, respectively) and three A:1 (lanes 4–6: P131, P418, and P237, respectively) avian Pasteurella multocida strains represent the B and A genotypes of the ptfA gene. Lane M = 100–base pair molecular marker. f
In summary, an allele-specific PCR assay for the ptfA gene of avian P. multocida strains was developed in the present study. Such allele-specific PCR assays of this and other virulence genes could complement other strain characterization methods used in molecular epidemiological investigations of P. multocida isolates and may also have the potential to assess the pathogenic potential of individual strains.
Acknowledgements
The authors thank Dr. J. H. Morgan and Dr. A. J. Lax for the useful comments on the manuscript.
Footnotes
a.
Bio-Rad Laboratories Inc., Hercules, CA.
b.
DyNAzyme™ II, Finnzymes Oy, Espoo, Finland.
c.
Wizard® SV Gel and PCR Clean-Up System, BigDye® v 1.1; Promega Corp., Madison, WI.
d.
ABI Prism 3100, Applied Biosystems Inc., Foster City, CA.
e.
DNASTAR Inc., Madison, WI.
f.
DreamTaq™ DNA Polymerase, GeneRuler™ DNA Ladder; Fermentas Inc., Burlington, Canada.
g.
Biometra T1, Biometra GmbH, Göttingen, Germany.
