Abstract
Canine coronavirus (CCoV) is usually the cause of mild gastroenteritis in dogs and is known to have spread worldwide. However, to date, no CCoV cases have been confirmed in Greece. In the present work, the authors investigated an outbreak of enteritis in puppies from a Greek kennel for the presence of CCoV. Dogs were presented with clinical signs of diarrhea, anorexia, weakness, depression, dehydration, and 1 death. Canine coronavirus type II was detected by reverse transcription nested polymerase chain reaction in all 11 puppies, whereas 1 puppy presented dual infection with CCoV type II and canine parvovirus 2. Surprisingly, sequence analysis of the samples revealed higher similarity to the pantropic CCoV II strain CB/05 than to other reference strains, in the most variable region of the S gene.
Canine coronavirus (CCoV; formally known as
The disease has a low mortality rate and is considered to be extremely contagious, especially in places such as kennels and animal shelters. Seroprevalence is significantly higher in such places, with higher population density compared with dogs housed singly.
1,5
Moreover, CCoV can often be found in concurrent infections with bacteria, parasites, or other viruses, such as
In the present work, the findings of the laboratory analysis of samples received from an outbreak of gastroenteritis in young dogs are reported. The outbreak took place in a municipal dog kennel in central Greece during the spring of 2008. The kennel houses 40–50 stray dogs of different ages at all times.
The outbreak involved 11 puppies (1.5–3 months old) from 4 different litters that were introduced into the quarantine area (a small area separated from the rest of the facilities) at around the same time. Vaccinations a against all major infectious diseases (canine distemper, infectious hepatitis, parainfluenza, parvoviral enteritis, and leptospi-rosis), along with antiparasitic treatment (praziquantel and fenbendazole b ), were applied on the first day of their arrival at the kennel, 5 days before the outbreak.
The outbreak began with severe diarrhea in all the puppies (no other dogs living in the kennel at that time had any signs of disease). Most puppies presented, apart from the diarrhea, with weakness, with 3 puppies showing more severe clinical signs of depression, anorexia, and dehydration. From the onset of the clinical signs, the whole group was treated with systemic antibiotics per os and regulatory drugs of the intestinal track. The 3 puppies that presented more severe clinical signs were given fluids and antibiotics (intravenously at first and intramuscularly thereafter). Most of the puppies responded well to the treatment, and after approximately 10 days, the feces appeared normal or almost normal (on the floor were found feces, soft to normal). However, despite treatment, 1 of the 3 dogs died 4 days after the onset of disease.
Sampling took place 4 days after the onset of clinical signs. Stool samples were collected from all of the puppies using rectal swabs. Total RNA and DNA were extracted from the supernatant fluid of the homogenized and centrifuged fecal samples with commercial RNA and DNA extraction kits, c,d respectively. Viral DNA for CPV-2 polymerase chain reaction (PCR) testing was also extracted from the supernatant fluid by boiling for 10 min and chilling on ice to inactivate the PCR inhibitors. 20 DNA extracts were diluted 1:10 in distilled water to minimize residual inhibitors to an ineffective concentration. 9
For the CCoV RNA detection, a reverse transcription nested PCR (RT-nPCR) assay was carried out on the fecal samples, as previously described. 18 For CPV-2 detection and for CAdV type 1 and type 2 (CAdV-1, CAdV-2) detection and differentiation, PCR assays were per-formed. 9,15 In addition, a reverse transcription PCR (RT-PCR) assay with primers e p1 and p2 was used for CDV nucleoprotein RNA detection. 13
For CCoV types I and II detection, genotype-specific real-time RT-PCR assays were performed. 12 Moreover, minor groove binder (MGB) probe assays were used to characterize the CPV-2 variants and discriminate between the vaccines and the field strains. 8,10
According to the results, all puppies were positive for CCoV and specifically CCoV type II. Only 1 of them was positive for CPV-2, which was characterized as CPV-2a and was determined to be a field, rather than vaccine, strain. This puppy presented a dual infection because it was positive for CCoV type II. Finally, all the samples tested negative for CDV and CAdV types 1 and 2.
To determine the sequence of the CCoV type II strains, a 1.4-kb region of the first part of the S gene was amplified from 2 samples (56/08 and 59/08), using a commercial one-step RT-PCR kit f and primers El-Ins1 and S2. 11 The amplicons were sequenced by a commercial facility, g and sequence assembling and analysis were carried out using the BioEdit software package 14 and National Center for Biotechnology Information (NCBI; http://www.ncbi.nlm.nih.gov) and European Molecular Biology Laboratory (EMBL; http://www.ebi.ac.uk) analysis tools. The obtained sequences (GenBank accessions GQ121370 and GQ121371, respectively) were translated and the predicted amino acid sequences were used for phylogenetic and molecular evolutionary analysis using Mega3. 16 Phylogenetic trees, based on the first part of the Sprotein (500 amino acids) of the 2 strains, were constructed using both parsimony and neighbor-joining methods, supplying a statistical support with bootstrapping more than 1,000 replicates (Fig. 1).
Sequence analysis of the 2 strains, in the first 500 amino acids of the S protein, showed a 99.8% amino acid identity, and a genetic relatedness of 85.8–87.2% with 2 CCoV type II reference strains, BGF10 and Insavc1, and of 90.6–94.8% with 2 FCoV type II reference strains, 79-1683 and 79-1146. Specifically, the most amino acid identity (96.8–97%) is evident with the CCoV type II pantropic strain CB/05. According to the phylogenetic analysis result, the CCoV type II strains identified in the outbreak segregate with CCoV type II pantropic strain CB/05 in the same clade with the other CCoV type II and FCoV type II reference strains (Fig. 1). Porcine coronaviruses form a unique clade within the same cluster of CCoV type II and FCoV type II, which is separate from that of CCoV type I and FCoV type I.
Since the 1970s, CCoV has been recognized to have a worldwide distribution, with many reports indicating that the virus had been detected and/or isolated in United States, Europe, Asia, and Australia. 1,5 In Greece, to date, there has been no similar work regarding CCoV and no data, until the present study, about the presence of the virus in the country. This is the first report demonstrating that CCoV, and in particular CCoV type II, is present in Greece.
Moreover, in the present work, CPV-2 was detected in the feces of a single dog, revealing that the field strain was a CPV-2a variant. To date, there has been no similar molecular characterization of CPV-2 variants in Greece because the diagnosis of the disease in the field is based on enzyme-linked immunosorbent assay (ELISA) and immunochromatographic (IC) commercial tests.
Results of the present study indicate that only 1 out of the 11 puppies was positive for CPV-2, even though all 11 puppies resided together. This can be explained by the fact that the puppies were vaccinated 5 days before the onset of the symptoms. The puppies came from 4 different litters and were younger than 2 months old. It is well known that titers of the maternal antibodies against CPV-2, as well as the response to vaccination, vary among members of a litter. In the current case, it is possible that interference caused by these antibodies led to vaccination failure. 4,21 However, it seems that the presence of parvovirus may not be related to the clinical manifestation of the disease as it relates to the 10 other puppies.
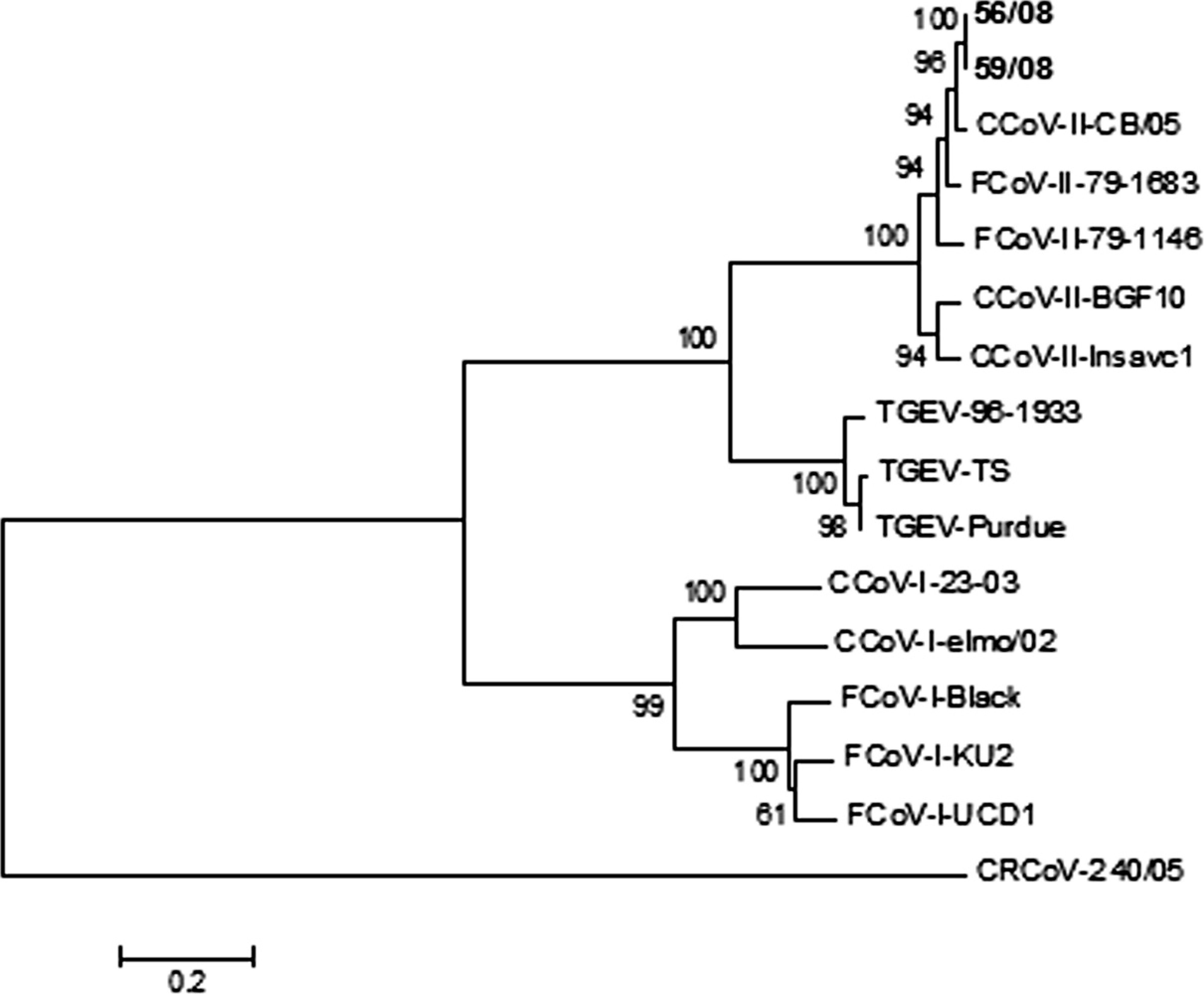
Phylogenetic tree of the Greek strains, based on the N terminus of the S protein, with reference strains CCoV-II-CB/05 (GenBank accession DQ112226), FCoV-II-79-1683 (X80799), FCoV-II-79-1146 (X06170), CCoV-II-BGF10 (AY342160), CCoV-II-Insavc1 (D13096), TGEV-96-1933 (AF104420), TGEV-TS (DQ201447), TGEV-Purdue (NC002306), CCoV-I-23-03 (AY307021), CCoV-I-elmo/02 (AY307020), FCoV-I-Black (EU186072), FCoV-I-KU2 (D32044), FCoV-I-UCD1 (AB088222), and CRCoV-240/05 (EU999954). The numbers represent the percentage of replicate trees based on 1,000 bootstrap replicates.
Furthermore, it should be noted that the puppy that presented dual infection with CCoV and CPV detected in feces, died. Canine coronavirus has been detected many times in dogs with other pathogens, causing more severe clinical symptoms, and dual infections with CCoV and CPV-2 have been reported. 1,2,5 The coexistence of the 2 pathogens in this puppy appears to be responsible for the observed mortality.
The 99.8% of amino acid identity between the 2 CCoV strains involved in the outbreak reveals that the puppies were infected from the same CCoV during the outbreak in the kennel. The high genetic similarity with the CCoV type II (85.8–97%) and FCoV type II (90.6–94.8%) reference strains confirms the characterization of this virus, by genotype-specific real-time RT-PCR, as CCoV type II. Remarkably, the identified strain has the most similarity, among all the CCoV type II reference strains, with the pantropic CCoV type II strain CB/05 in the most variable region of the S gene. Unfortunately, the deceased dog was not sent for necropsy and viral detection in the organs, so it was not possible to verify the presence of the virus in the biological samples.
Footnotes
a.
Vanguard® Plus5/L, Pfizer Animal Health S.A., Louveigne, Belgium.
b.
Caniquantel® Plus, Fendigo sa/nv, Brussels, Belgium.
c.
QIAamp® Viral RNA Mini Kit, Qiagen GmbH, Hilden, Germany.
d.
DNeasy® Tissue Kit, Qiagen GmbH, Hilden, Germany.
e.
Primers, Eurofins MWG GmbH, Ebersberg, Germany.
f.
SuperScriptTM One-Step RT-PCR for Long Templates, Invitrogen SRL, San Giuliano Milanese, Italy.
g.
Genome Express, Meylan, France.
